Team:Heidelberg/Notebook/miRNA Kit/September
From 2010.igem.org
(→29/09/2010) |
|||
| (19 intermediate revisions not shown) | |||
| Line 68: | Line 68: | ||
===01/09/2010=== | ===01/09/2010=== | ||
| - | [[Image:20100901_gel1bb.png|thumb|350 px| | + | [[Image:20100901_gel1bb.png|thumb|350 px|left|Gel 100901-1]] |
| - | [[Image:20100901_gel2bb.png|thumb|350 px| | + | [[Image:20100901_gel2bb.png|thumb|350 px|left|Gel 100901-2]] |
<br /> | <br /> | ||
* 8 colonies of each plate (see [[Igem2010/Main/synthetic miR Kit/August#31/08/2010 | previous day]]) were picked and colony-PCR was performed. Furthermore, the colonies were used for inocculation of LB miniprep cultures. The numbers behind the construct name refer to vector type (1 = not SAP treated, 2 = dephosphorylated using SAP; see previous day). For hRluc (1300 bp band)), Luc2 (2300 bp band), CMV_TetO (1000 bp band) and the FRT (500 bp band) site, positive colonies could be observed (gel 100901-1 and 100901-2). | * 8 colonies of each plate (see [[Igem2010/Main/synthetic miR Kit/August#31/08/2010 | previous day]]) were picked and colony-PCR was performed. Furthermore, the colonies were used for inocculation of LB miniprep cultures. The numbers behind the construct name refer to vector type (1 = not SAP treated, 2 = dephosphorylated using SAP; see previous day). For hRluc (1300 bp band)), Luc2 (2300 bp band), CMV_TetO (1000 bp band) and the FRT (500 bp band) site, positive colonies could be observed (gel 100901-1 and 100901-2). | ||
| Line 77: | Line 77: | ||
<br /> | <br /> | ||
===02/09/2010=== | ===02/09/2010=== | ||
| - | [[Image:20100802_gel1.png|thumb|350 px| | + | [[Image:20100802_gel1.png|thumb|350 px|left|Gel 100902-1]] |
| + | [[Image:20100902_gel2.png|thumb|350 px|right|Gel 100902-2]] | ||
| + | [[Image:20100902_gel3.png|thumb|350 px|right|Gel 100902-3]] | ||
<br /> | <br /> | ||
* 8 colonies were picked from plates 1 and 4 colonies from plates 2 from the previous days cloning and analyzed via colony PCR performed according to the [https://2010.igem.org/Team:Heidelberg/Notebook/Methods#Colony_PCR PCR standard protocol]. The results of the colony PCR were analyzed on a 1 % agarose gel run for 1 h @ 100 V (gel 100902-1 and 100902-2). The following colonies were afterwards inocculated (LB miniprep cultures): | * 8 colonies were picked from plates 1 and 4 colonies from plates 2 from the previous days cloning and analyzed via colony PCR performed according to the [https://2010.igem.org/Team:Heidelberg/Notebook/Methods#Colony_PCR PCR standard protocol]. The results of the colony PCR were analyzed on a 1 % agarose gel run for 1 h @ 100 V (gel 100902-1 and 100902-2). The following colonies were afterwards inocculated (LB miniprep cultures): | ||
| Line 86: | Line 88: | ||
:* shRNA9 (1.2) | :* shRNA9 (1.2) | ||
<br /> | <br /> | ||
| - | + | ||
| - | + | ||
* The cultures inocculated the day before were mini prepped by applying a Qiagen Miniprep Kit. Afterwards, a test digestion with NotI was performed and the result was analyzed on a 1 % agarose gel, run for 35 min @ 100 V (gel 100902-3). According to the test digestion result, the following samples were send for sequencing @ GATC: | * The cultures inocculated the day before were mini prepped by applying a Qiagen Miniprep Kit. Afterwards, a test digestion with NotI was performed and the result was analyzed on a 1 % agarose gel, run for 35 min @ 100 V (gel 100902-3). According to the test digestion result, the following samples were send for sequencing @ GATC: | ||
:* CMV_TetO2 (.4, 1.6) | :* CMV_TetO2 (.4, 1.6) | ||
| Line 94: | Line 95: | ||
:* hRluc (1.1, 1.3, 1.4) | :* hRluc (1.1, 1.3, 1.4) | ||
<br /> | <br /> | ||
| - | + | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> | |
| Line 108: | Line 109: | ||
* cloning of shRNAs into pcDNA5/FRT/TO | * cloning of shRNAs into pcDNA5/FRT/TO | ||
| - | <br /><br /><br /><br /><br /><br /><br /><br /><br /> | + | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> |
| - | + | <br /><br /><br /> | |
===04/09/2010=== | ===04/09/2010=== | ||
| Line 174: | Line 175: | ||
* ligation of both inserts for the tuning construct, extraction of the right insert (3000bp) and ligation into the backbone | * ligation of both inserts for the tuning construct, extraction of the right insert (3000bp) and ligation into the backbone | ||
| - | <br /><br /><br /> | + | <br /><br /><br /><br /><br /><br /> |
===09/09/2010=== | ===09/09/2010=== | ||
| Line 186: | Line 187: | ||
<br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> | ||
<br /><br /> | <br /><br /> | ||
| - | <br /><br /> | + | <br /><br /><br /><br /><br /><br /><br /><br /> |
| - | <br /><br /> | + | <br /><br /><br /><br /><br /> |
<br /><br /> | <br /><br /> | ||
| Line 210: | Line 211: | ||
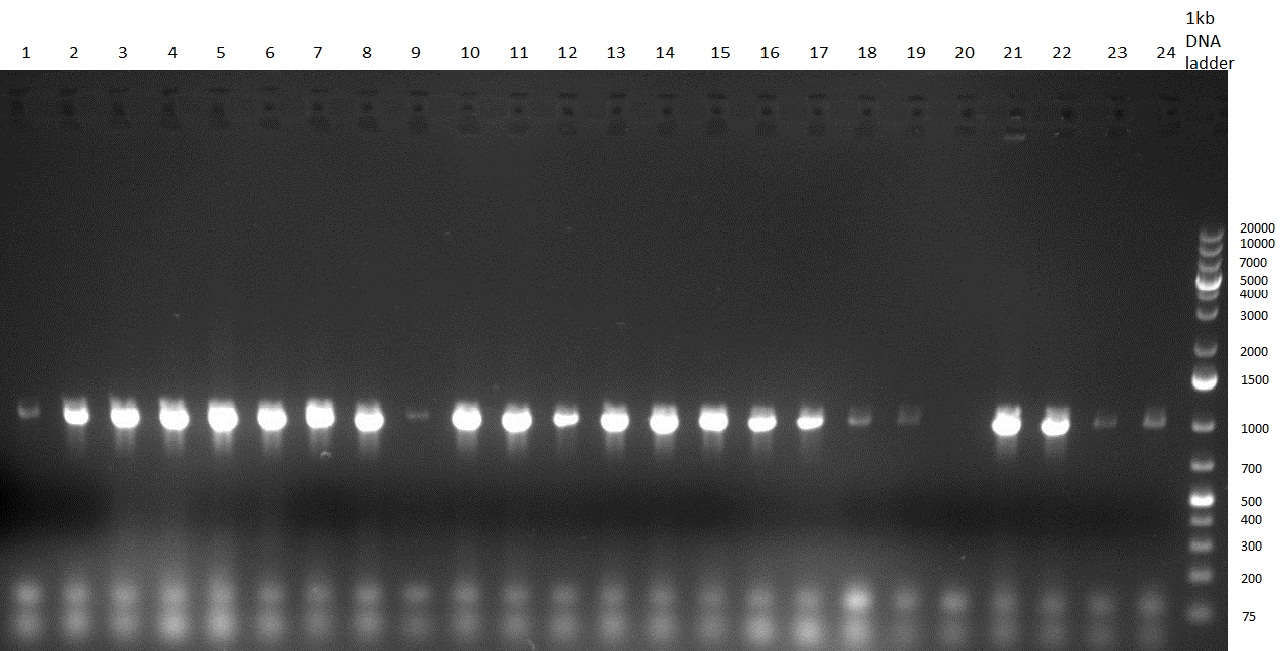
** PCR of CMV shRNA which leads to a size of 1200bp | ** PCR of CMV shRNA which leads to a size of 1200bp | ||
| - | <br /><br /><br /> | + | <br /><br /><br /><br /><br /><br /><br /><br /><br /> |
===11/09/2010=== | ===11/09/2010=== | ||
| Line 223: | Line 224: | ||
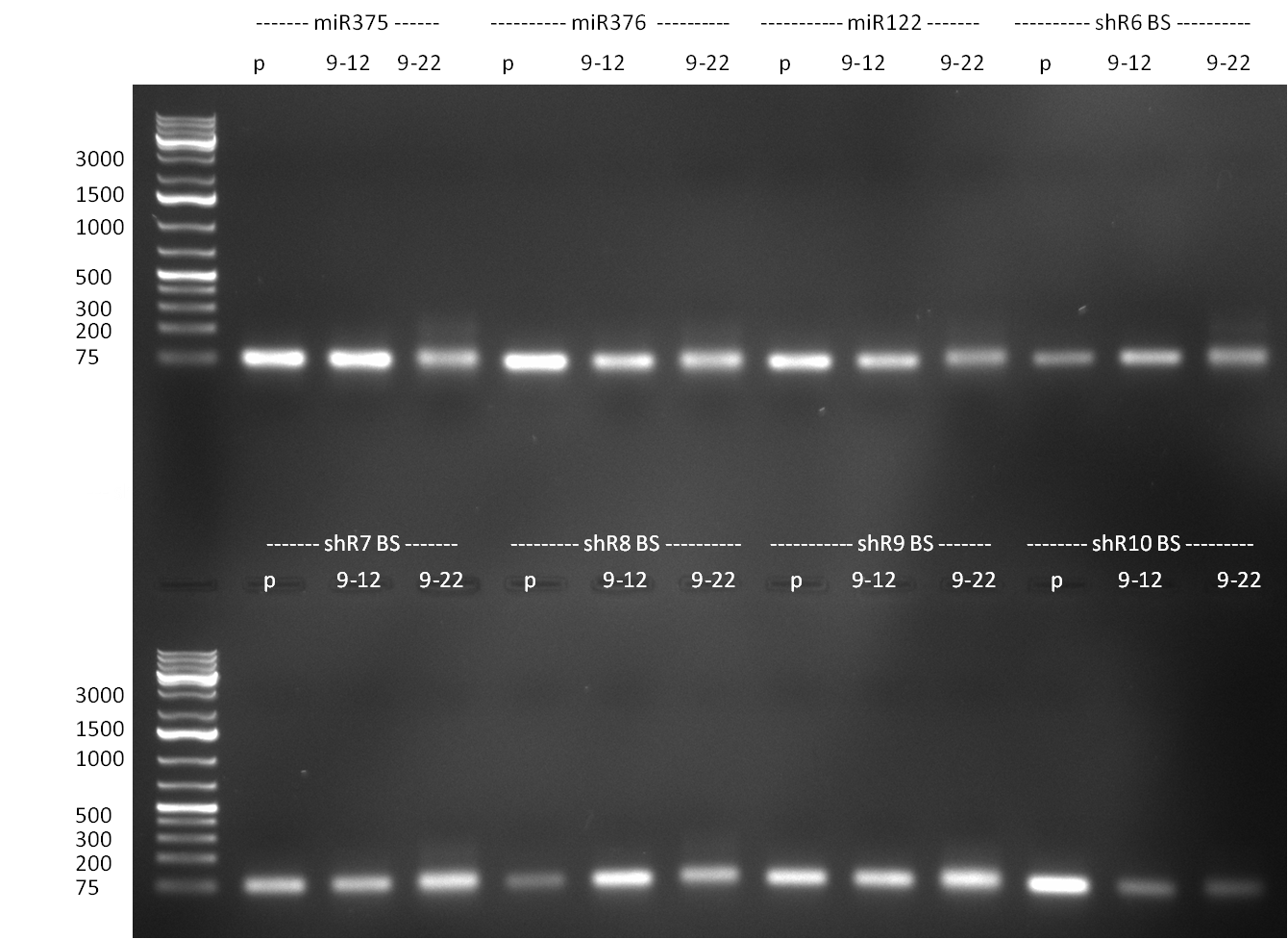
[[Image:Rb100911_testdigestshRNA.jpg|thumb|350px|right|test digest]] | [[Image:Rb100911_testdigestshRNA.jpg|thumb|350px|right|test digest]] | ||
| - | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> | + | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> |
===12/09/2010=== | ===12/09/2010=== | ||
| Line 233: | Line 234: | ||
* miniprep of shRNA constructs and Tuning constructs | * miniprep of shRNA constructs and Tuning constructs | ||
| - | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> | + | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> |
===13/09/2010=== | ===13/09/2010=== | ||
| Line 260: | Line 261: | ||
* Colony- PCR for the cloning products done on the previous day (gels 100914-1 and 100914-2 on the right) | * Colony- PCR for the cloning products done on the previous day (gels 100914-1 and 100914-2 on the right) | ||
* positive clones were Mini-Prepped and test digested | * positive clones were Mini-Prepped and test digested | ||
| - | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> | + | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> |
===15/09/2010=== | ===15/09/2010=== | ||
| Line 344: | Line 345: | ||
<br /> | <br /> | ||
* Selection of colonies from the plates (see previous days' cloning) via colony-PCR using the standard sequencing primers; a ~5.5 kb fragment should be visible for the assembled construct K1-K8, if colony-PCR was positive (gel 100917-1 and 100917-2). No sample shows a 5.5 kb band, but that's most likely due to the very difficult amplification of such a long fragment with Fermentas 2x PCR MasterMix (Taq-based). Therefor 3 miniprep cultures were inocculated for each construct (K1-K8). The TetR oligo insertion and the Kozag_hRluc_BGH cloning into pSB1C3 gave the right bands on the gel. | * Selection of colonies from the plates (see previous days' cloning) via colony-PCR using the standard sequencing primers; a ~5.5 kb fragment should be visible for the assembled construct K1-K8, if colony-PCR was positive (gel 100917-1 and 100917-2). No sample shows a 5.5 kb band, but that's most likely due to the very difficult amplification of such a long fragment with Fermentas 2x PCR MasterMix (Taq-based). Therefor 3 miniprep cultures were inocculated for each construct (K1-K8). The TetR oligo insertion and the Kozag_hRluc_BGH cloning into pSB1C3 gave the right bands on the gel. | ||
| - | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> | + | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> |
===19/09/2010=== | ===19/09/2010=== | ||
| Line 365: | Line 366: | ||
[https://2010.igem.org/Team:Heidelberg/Notebook/Methods#Dual_Luciferase_Assay Dual Luciferase Assay]<br/> | [https://2010.igem.org/Team:Heidelberg/Notebook/Methods#Dual_Luciferase_Assay Dual Luciferase Assay]<br/> | ||
In order to test for promoter efficiency and to check whether the miRNA kit assembly works fine | In order to test for promoter efficiency and to check whether the miRNA kit assembly works fine | ||
| - | 50ng of each construct with different promoter set-ups were transfected into HEK 293 T-REx cells and other cell lines HEK, HeLa, Huh7 in 96-well plate format using FuGENE transfection reagent. As every construct is expressing firefly luciferase (luc2) and renilla luciferase (hRluc) at the same time the setup allows for standardization of transfection efficiency as only luc2 is tagged with binding sites. Each sample was transfected and measured by Dual luciferase assay in 8 replicates. As by this time no shRNA has been cloned into plasmid no knock-down of luc2 is expected and the different expression efficiencies allow for characterization of the different promoters. [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337032 BBa_K337032] leads to a relative luciferase unit (RLU) of luc2 to hRluc expression of 6 RLU. [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337035 BBa_K337035] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337035 BBa_K337035] are showing a comparable expression of 12 - 13 RLU, which is in line with the knowledge that both luciferases are driven by the CMV promoter. Hek 293 T-Rex cells stably express the Tet repressor thus allows us to observe very efficient repression of Firefly luciferse expression if a CMV-TetO2 promoter is driving luc2 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K337038 BBa_K337038] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337046 BBa_K337046]). [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337040 BBa_K337040] transfection into Hek T-Rex cells results in an expression of 15 RLU. [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337042 BBa_K337042] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337044 BBa_K337044] are constructed in a way that luc2 is driven by the CMV promoter and hRluc is driven by the RSV promoter and show a comparable expression of 17-20 RLU. This leads to the conclusion that the CMV promoter | + | 50ng of each construct with different promoter set-ups were transfected into HEK 293 T-REx cells and other cell lines HEK, HeLa, Huh7 in 96-well plate format using FuGENE transfection reagent. As every construct is expressing firefly luciferase (luc2) and renilla luciferase (hRluc) at the same time the setup allows for standardization of transfection efficiency as only luc2 is tagged with binding sites. Each sample was transfected and measured by Dual luciferase assay in 8 replicates. As by this time no shRNA has been cloned into plasmid no knock-down of luc2 is expected and the different expression efficiencies allow for characterization of the different promoters. [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337032 BBa_K337032] leads to a relative luciferase unit (RLU) of luc2 to hRluc expression of 6 RLU. [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337035 BBa_K337035] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337035 BBa_K337035] are showing a comparable expression of 12 - 13 RLU, which is in line with the knowledge that both luciferases are driven by the CMV promoter. Hek 293 T-Rex cells stably express the Tet repressor thus allows us to observe very efficient repression of Firefly luciferse expression if a CMV-TetO2 promoter is driving luc2 ([http://partsregistry.org/wiki/index.php?title=Part:BBa_K337038 BBa_K337038] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337046 BBa_K337046]). [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337040 BBa_K337040] transfection into Hek T-Rex cells results in an expression of 15 RLU. [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337042 BBa_K337042] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337044 BBa_K337044] are constructed in a way that luc2 is driven by the CMV promoter and hRluc is driven by the RSV promoter and show a comparable expression of 17-20 RLU. This leads to the conclusion that the CMV promoter shows comparable expression to the RSV promoter in Hek T-Rex cell lines. If transfecting [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337035 BBa_K337035] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337036 BBa_K337036] are transfected into different cell lines it is obvious that Hek293T cells are the easiest to transfect with both constructs an expression of 17-22 RLU is to be measured. Hek T-Rex cells are showing and expression level of 12 RLU of both constructs. Hela cells are also showing constant expression levels of 8 RLU with both constructs. A rather high standard deviation in the luciferase expression and also differences between the 2 constructs is to be seen in Huh7 cells. This might be due to low transfection efficiency of this cell line in general. Alltogether it is to say that [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337035 BBa_K337035] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337036 BBa_K337036] show comparable expression. |
| Line 389: | Line 390: | ||
|- | |- | ||
|} | |} | ||
| + | <br /><br /><br /> | ||
| + | [[Image:Promoter_test_220910_hd2010.jpg| thumb | 600px | centre | Promoter strength characterization in HEK 293 T-REx cell line]] | ||
| + | <br /><br /><br /> | ||
| + | [[Image:23092010 DLA promoters cell lines.jpg | thumb | 600px | centre| Promoter strength characterization in different cell lines]] | ||
| - | + | <br /> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
===23/09/2010=== | ===23/09/2010=== | ||
| Line 407: | Line 407: | ||
[https://2010.igem.org/Team:Heidelberg/Notebook/Methods#Dual_Luciferase_Assay Dual Luciferase Assay] was performed as on [[https://2010.igem.org/Team:Heidelberg/Notebook/miRNA_Kit/September#22.2F09.2F2010 previous day]]. Cells were not harvested after 20h, but 48h. | [https://2010.igem.org/Team:Heidelberg/Notebook/Methods#Dual_Luciferase_Assay Dual Luciferase Assay] was performed as on [[https://2010.igem.org/Team:Heidelberg/Notebook/miRNA_Kit/September#22.2F09.2F2010 previous day]]. Cells were not harvested after 20h, but 48h. | ||
| - | + | <br /><br /><br /><br /><br /><br /> | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
===24/09/2010=== | ===24/09/2010=== | ||
| Line 449: | Line 444: | ||
** ligate 150ng and 250ng of insert into 30ng [http://partsregistry.org/Part:pSB1A3 pSB1A3] 1h at RT and transform ligation | ** ligate 150ng and 250ng of insert into 30ng [http://partsregistry.org/Part:pSB1A3 pSB1A3] 1h at RT and transform ligation | ||
<br><br><br><br><br><br> | <br><br><br><br><br><br> | ||
| - | <br /><br /><br /><br /><br /><br /><br /><br /><br /> | + | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> |
===27/09/2010=== | ===27/09/2010=== | ||
| Line 499: | Line 494: | ||
<br><br><br><br><br><br><br><br> | <br><br><br><br><br><br><br><br> | ||
| - | <br><br><br><br><br><br><br><br><br><br> | + | <br><br><br><br><br><br><br><br><br><br><br /><br /><br /><br /> |
| - | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> | + | <br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> |
===29/09/2010=== | ===29/09/2010=== | ||
| Line 538: | Line 533: | ||
* ligation of P56 (14) with R8 and R11 and SAPed backbone and control ligation with backbone only, transformation | * ligation of P56 (14) with R8 and R11 and SAPed backbone and control ligation with backbone only, transformation | ||
<br/><br/><br/><br/><br/><br/> | <br/><br/><br/><br/><br/><br/> | ||
| - | <br/><br/><br/><br/><br/><br/> | + | <br/><br/><br/><br/><br/><br/><br /><br /><br /> |
| - | <br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/> | + | <br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /><br /> |
<br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/> | <br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/> | ||
Latest revision as of 05:57, 27 October 2010

01/09/2010
02/09/2010
03/09/2010
04/09/2010
05/09/2010
06/09/2010
07/09/2010
All Ligations were controlled by loading 10 ul of ligation reactiong onto a 1 % agarose gel (100907-1), run for 35 min @ 100 V
08/09/2010
09/09/2010
10/09/2010
Subsequent Transformation into DH5alpha cells
11/09/2010
12/09/2010
13/09/2010
14/09/2010
15/09/2010
Each PCR was performed in two replicates. The result of the PCRs was analyzed on a 1 % agarose gel (100915-1), run for 1 h @ 100 V. The PCR nr. 2 was subsequently PCR purified by applying a Qiagen PCR purification KIT and used for the cloning 16/09/2010
17/09/2010
19/09/2010
20/09/2010
22/09/2010
23/09/2010
24/09/2010
25/09/2010
26/09/2010
27/09/2010
28/09/2010tuning construct:
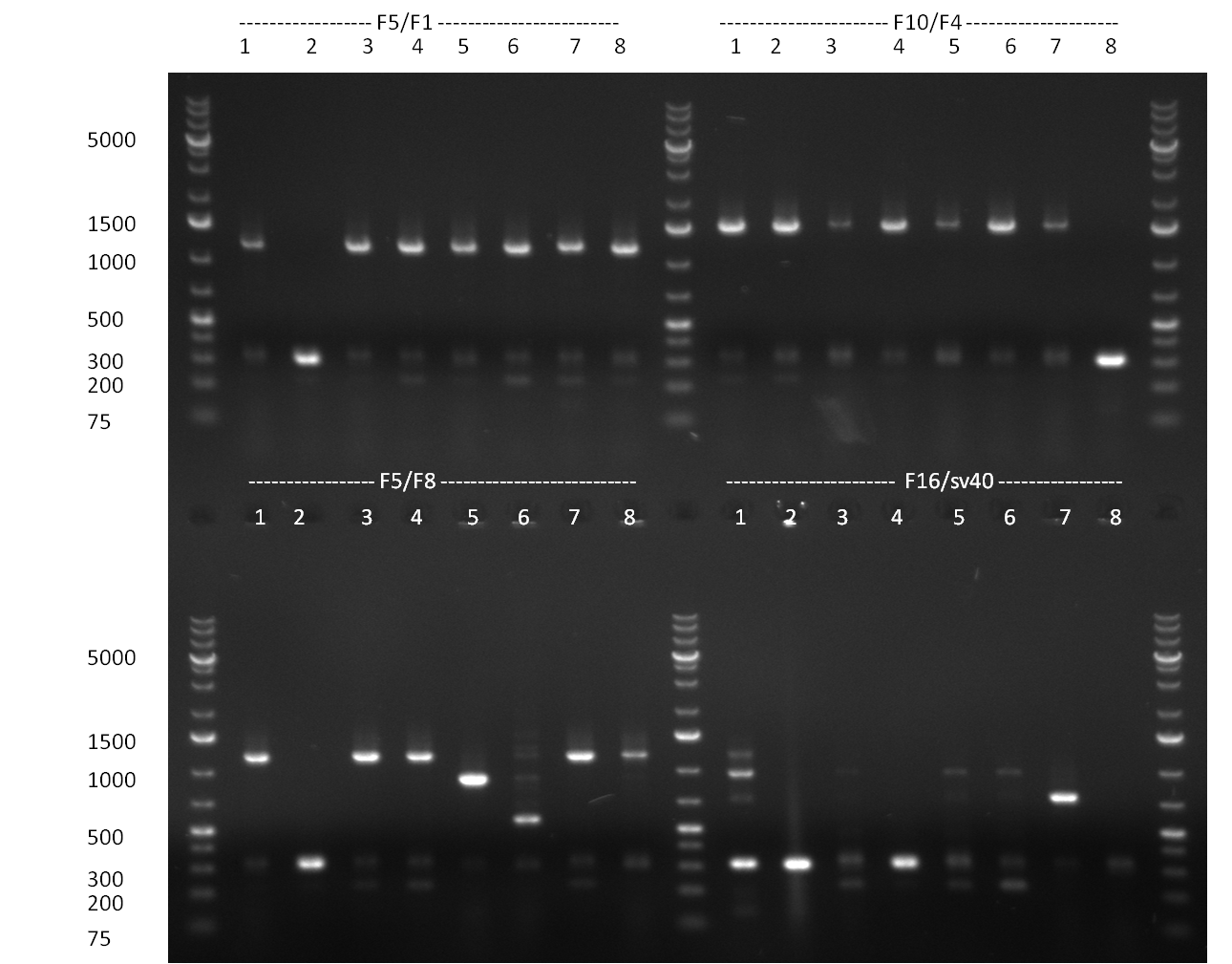
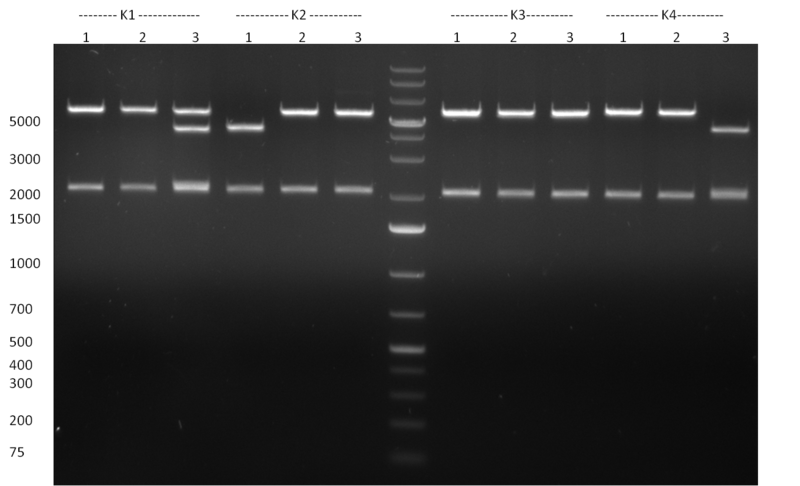
29/09/2010 Gel 100929-1 first gel: double digestion of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337032 K1] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337036 K3] with PstI and EcoRI: [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337032 K1] (lane 1-4), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337036 K3] (lane 5-8), second gel: digestion of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337032 K1] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337036 K3] with BamHI: [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337032 K1] (lane 1-4), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337036 K3] (lane 5-8)  Gel 1000929-2 digestion of [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337032 K1] and [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337036 K3] with HindIII and AflII: [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337032 K1] (lane 1&2), [http://partsregistry.org/wiki/index.php?title=Part:BBa_K337036 K3](lane 3-12) tuning construct:
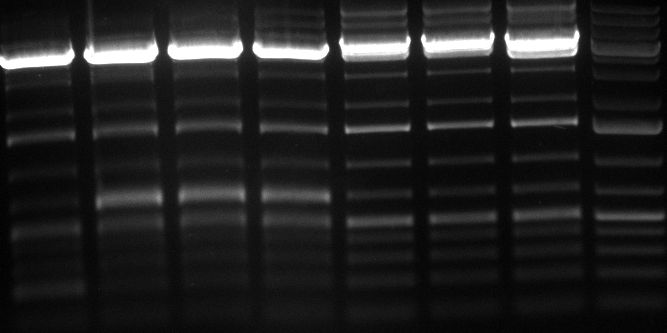
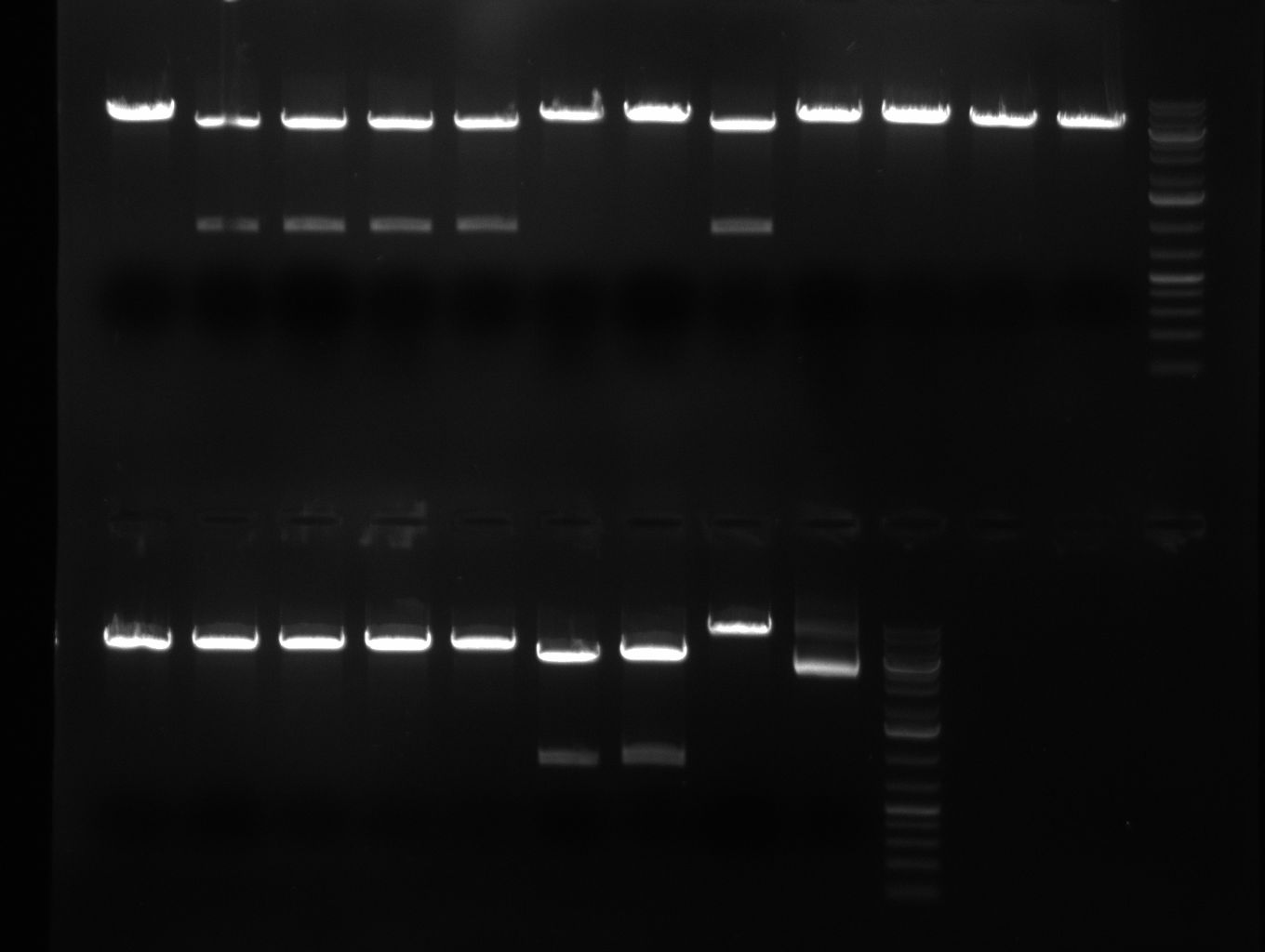
 Gel 100929 r-5: test digestion of miniprep number 14 withe EcoRI and NheI shows expected band at 1100bp. Digestions of R8 and R11 were also positive, DNA cut out with SpeI and PstI runs at 800bp. Lane 1=miniprep 14 undigested, lane2=miniprep 14 digested, lane3=miniprep 23 digested, lane4=R8 undigested, lane5= R8 digested, lane6=R11 undigested, lane7=R11 digested repressor construct:
30/09/2010tuning construct:
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"