Team:Imperial College London/Lab Diaries/Vectors team
From 2010.igem.org
(→Week 6) |
|||
| (12 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{:Team:Imperial_College_London/Templates/Header}} | {{:Team:Imperial_College_London/Templates/Header}} | ||
{{:Team:Imperial_College_London/Templates/LabDiariesHeader}} | {{:Team:Imperial_College_London/Templates/LabDiariesHeader}} | ||
| - | |||
| - | |||
| - | |||
| - | |||
{| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
|style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Objectives | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Objectives | ||
|- | |- | ||
| | | | ||
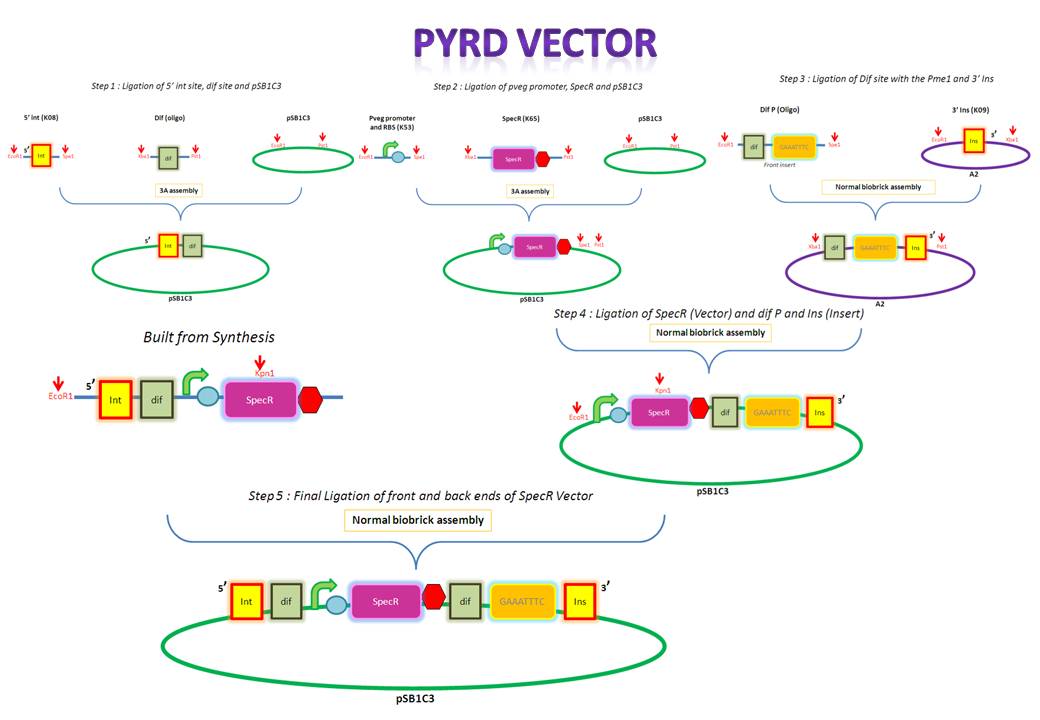
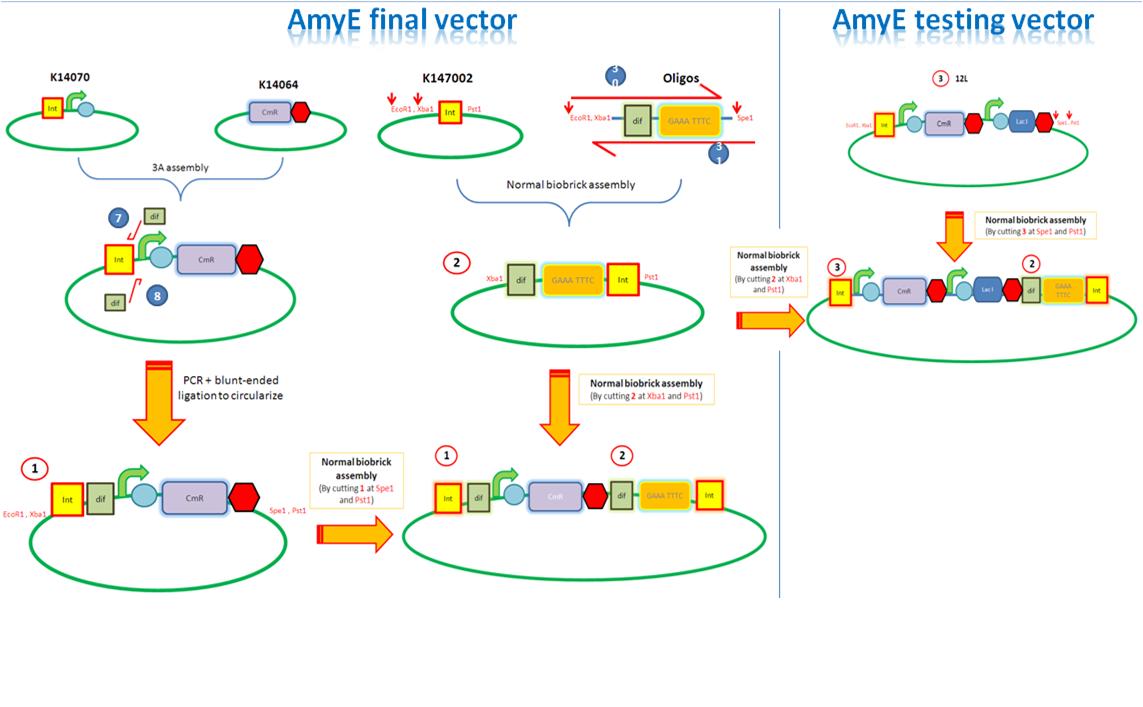
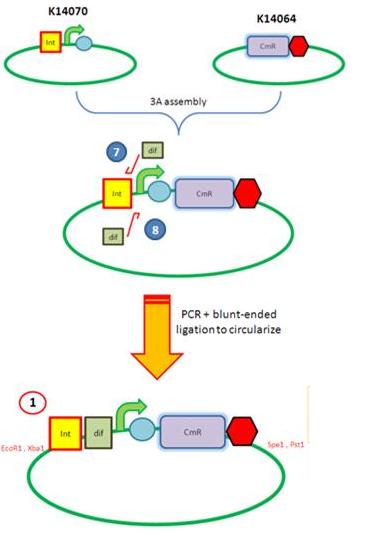
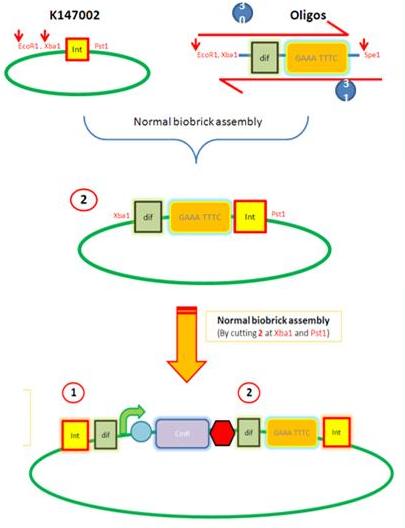
| - | We are assembling the AmyE and PyrD vectors in order to transform B. Subtilis with our parts. Once completed, these vectors will be reusable and can then be used to introduce any relevant piece of DNA directly into B.subtilis genome. The vectors will be used both for the final assembly and for testing constructs. | + | We are assembling the AmyE and PyrD vectors in order to transform ''B. Subtilis'' with our parts. Once completed, these vectors will be reusable and can then be used to introduce any relevant piece of DNA directly into ''B.subtilis'' genome. The vectors will be used both for the final assembly and for testing constructs. |
|} | |} | ||
{| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
| Line 19: | Line 15: | ||
|} | |} | ||
| - | === | + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" |
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Week 6 | ||
| + | |- | ||
| + | | | ||
| Line 57: | Line 56: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;color:#555555;height:100px;width:200px;text-align:left;"> |
<ul> | <ul> | ||
<li>Restriction digest of 5' dif XP with XbaI and PstI </li> | <li>Restriction digest of 5' dif XP with XbaI and PstI </li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;color:#555555;height:100px;width:200px;text-align:left;"> |
</td> | </td> | ||
</tr> | </tr> | ||
| Line 68: | Line 67: | ||
<td style="background-color:#FFCC66;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Afternoon</b> | <td style="background-color:#FFCC66;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Afternoon</b> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:center;"> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:center;"> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:center;"> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<b>Start assembly of PyrD vector</b> | <b>Start assembly of PyrD vector</b> | ||
| Line 86: | Line 85: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of resultant products from 5' dif XP digest</li> | <li>Gel analysis of resultant products from 5' dif XP digest</li> | ||
| Line 97: | Line 96: | ||
</html> | </html> | ||
| - | + | '''Thursday, August 12''' | |
* Annealing the forward and reverse strands of the '''dif XP''' oligo | * Annealing the forward and reverse strands of the '''dif XP''' oligo | ||
| - | The forward and reverse strands of the 5' dif site with XbaI and PstI restriction sites on either side have been synthesized separately. The synthesized fragments arrive in solid powder form. These were immediately diluted in ddH2O to obtain a stock concentration of 1 ng/ | + | The forward and reverse strands of the 5' dif site with XbaI and PstI restriction sites on either side have been synthesized separately. The synthesized fragments arrive in solid powder form. These were immediately diluted in ddH2O to obtain a stock concentration of 1 ng/μl. They were then allowed to anneal together by first heating them to 95°C for denaturation and allowing them to cool down and anneal overnight. |
| - | + | '''Friday, August 13''' | |
* Restriction digest of '''dif XP''' | * Restriction digest of '''dif XP''' | ||
| Line 109: | Line 108: | ||
The cut oligo was PCR purified in order to get rid of any contaminants. PCR purification gets rid of short pieces of DNA which are less than about 40 base pairs. | The cut oligo was PCR purified in order to get rid of any contaminants. PCR purification gets rid of short pieces of DNA which are less than about 40 base pairs. | ||
| - | * Gel Analysis of | + | * Gel Analysis of 'dif XP' |
| + | |} | ||
| - | === | + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" |
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Week 7 | ||
| + | |- | ||
| + | | | ||
<html> | <html> | ||
| Line 134: | Line 137: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color#555555"> |
<ul> | <ul> | ||
| - | <li>Restriction digestion of 5' ins [K143008] (using | + | <li>Restriction digestion of 5' ins [K143008] (using EcoI and SpeI)</li> |
<li>PCR amplification of vector backbone pSB1C3</li> | <li>PCR amplification of vector backbone pSB1C3</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis and extraction of 5' ins</li> | <li>Gel analysis and extraction of 5' ins</li> | ||
<li>Gel purification of 5' ins</li> | <li>Gel purification of 5' ins</li> | ||
| - | <li>Restriction digestion of pSB1C3 (using | + | <li>Restriction digestion of pSB1C3 (using EcoRI and PstI)</li> |
<li>PCR purification of pSB1C3 after digestion</li> | <li>PCR purification of pSB1C3 after digestion</li> | ||
<li>Gel analysis of 5' ins, dif and pSB1C3 to work out ratios for ligation</li> | <li>Gel analysis of 5' ins, dif and pSB1C3 to work out ratios for ligation</li> | ||
| - | <li>Restriction digestion of | + | <li>Restriction digestion of pVeg promoter and RBS [K143053] (using EcoRI and SpeI)</li> |
| - | <li>Restriction | + | <li>Restriction digestion of SpecR-T [K143065] (using XbaI and PstI)</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Replica plating and colony PCR of transformed colonies (containing 5' dif in pSB1C3)</li> | <li>Replica plating and colony PCR of transformed colonies (containing 5' dif in pSB1C3)</li> | ||
| - | <li>Gel extraction and PCR purification of | + | <li>Gel extraction and PCR purification of pVeg and SpecR-T in preparation for ligations this afternoon</li> |
| - | <li>Gel analysis of | + | <li>Gel analysis of pVeg , Spec-T and pSB1C3 in order to work out ratios for the ligation</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Transformation of overnight ligations: 5' ins, dif with pSB1C3 and | + | <li>Transformation of overnight ligations: 5' ins, dif with pSB1C3 and pVeg, SpecR-T and pSB1C3 </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Replica plating and colony PCR of: 5' ins, dif with pSB1C3 and | + | <li>Replica plating and colony PCR of: 5' ins, dif with pSB1C3 and pVeg, SpecR-T and pSB1C3 </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 178: | Line 181: | ||
<tr> | <tr> | ||
| - | <td style="background-color:#FFCC66;width:100px;text-align: | + | <td style="background-color:#FFCC66;width:100px;text-align:center;color:#555555;"><b>Afternoon</b> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>PCR purification of PSB1C3 vector </li> | <li>PCR purification of PSB1C3 vector </li> | ||
| Line 187: | Line 190: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Ligation of 5' ins and dif with pSB1C3 - a bench ligation (1 hour) and an overnight ligation were set up </li> | <li>Ligation of 5' ins and dif with pSB1C3 - a bench ligation (1 hour) and an overnight ligation were set up </li> | ||
| - | <li>Transformation of E.Coli with the bench ligated products</li> | + | <li>Transformation of ''E.Coli'' with the bench ligated products</li> |
| - | <li>Gel analysis and extraction of | + | <li>Gel analysis and extraction of pVeg and SpecR-T</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Overnight ligation of | + | <li>Overnight ligation of pVeg and SpecR-T with pSB1C3 </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of colony PCR products from the transformations</li> | <li>Gel analysis of colony PCR products from the transformations</li> | ||
| - | <li> Overnight annealing of | + | <li> Overnight annealing of dif PES (Synthesized oligo)</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 216: | Line 219: | ||
| - | + | '''Monday, August 16''' | |
*Restriction digest of '''5' ins [K143008]''' | *Restriction digest of '''5' ins [K143008]''' | ||
| Line 227: | Line 230: | ||
The PCR amplified vector was purified in order to get rid of any contaminants. For example, short pieces of DNA like the primers. | The PCR amplified vector was purified in order to get rid of any contaminants. For example, short pieces of DNA like the primers. | ||
| - | + | '''Tuesday, August 17''' | |
*Gel extraction and purification of '''5' ins ES''' | *Gel extraction and purification of '''5' ins ES''' | ||
| Line 238: | Line 241: | ||
The digested pSB1C3 was re-purified in order to get rid of any contaminant DNA that arose during the digestion. | The digested pSB1C3 was re-purified in order to get rid of any contaminant DNA that arose during the digestion. | ||
| - | *Restriction digests of ''' | + | *Restriction digests of '''pVeg and Spec-T''' |
| - | + | pVeg (promoter and RBS) and Spec-T (Spectinomycin with a terminator) were digested in preparation for 3A assembly. | |
*Ligation of '''5' ins (ES) and dif (XP) with pSB1C3 (EP)''' | *Ligation of '''5' ins (ES) and dif (XP) with pSB1C3 (EP)''' | ||
The digested 5' ins and dif (the front inserts) were ligated overnight with pSB1C3 (the vector). A bench ligation and an overnight ligation were set up. | The digested 5' ins and dif (the front inserts) were ligated overnight with pSB1C3 (the vector). A bench ligation and an overnight ligation were set up. | ||
| - | *Transformation of E.Coli with bench ligate | + | *Transformation of ''E.Coli'' with bench ligate |
| - | E.Coli was transformed via the chemical method using the bench ligate. | + | ''E.Coli'' was transformed via the chemical method using the bench ligate. |
| - | *Gel extraction and purification of ''' | + | *Gel extraction and purification of '''pVeg and Spec-T''' |
Since these are both inserts they were gel extracted and purified. PCR purification is not carried out for inserts since they are small and would therefore be lost during the process. | Since these are both inserts they were gel extracted and purified. PCR purification is not carried out for inserts since they are small and would therefore be lost during the process. | ||
| - | + | '''Wednesday, August 18''' | |
* Replica plating and colony PCR of 5' ins and dif in pSB1C3 (bench ligation) | * Replica plating and colony PCR of 5' ins and dif in pSB1C3 (bench ligation) | ||
| - | * Ligation of | + | * Ligation of pVeg (ES) and SpecR-T (XP) with pSB1C3 (EP) |
| - | + | pVeg and SpecR-T (the front inserts) were ligated overnight with pSB1C3 (the vector). | |
| - | + | '''Thursday, August 19''' | |
| - | *Transformation of E.Coli with the overnights ligates | + | *Transformation of ''E.Coli'' with the overnights ligates |
#''' 5' ins and dif in pSB1C3''' | #''' 5' ins and dif in pSB1C3''' | ||
| - | #''' | + | #'''pVeg and SpecR-T in pSB1C3''' |
| - | + | '''Friday, August 20''' | |
*Replica plating and colony PCR of both transformations from yesterday. | *Replica plating and colony PCR of both transformations from yesterday. | ||
*Annealing the forward and reverse strands of the dif P ES oligo | *Annealing the forward and reverse strands of the dif P ES oligo | ||
| - | The forward and reverse strands of the 3' dif Pme1 sites with XbaI and PstI restriction sites on either side have been synthesized separately. The synthesized fragments arrive in solid powder form. These were immediately diluted in ddH2O to obtain a stock concentration of 1 ng/ | + | The forward and reverse strands of the 3' dif Pme1 sites with XbaI and PstI restriction sites on either side have been synthesized separately. The synthesized fragments arrive in solid powder form. These were immediately diluted in ddH2O to obtain a stock concentration of 1 ng/μl. They were then allowed to anneal together by heating them to 95°C for denaturation and allowing them to cool down and anneal overnight. |
| + | |} | ||
| - | === | + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" |
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Week 8 | ||
| + | |- | ||
| + | | | ||
<html> | <html> | ||
<table width="850px" border="0"> | <table width="850px" border="0"> | ||
| Line 285: | Line 292: | ||
</td> | </td> | ||
<td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Friday</b> | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Friday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Saturday</b> | ||
</td> | </td> | ||
</tr> | </tr> | ||
| Line 291: | Line 300: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Set up overnight ligations for standard assembly (BBA) and 3A cloning (3A) of dif P</li> | <li>Set up overnight ligations for standard assembly (BBA) and 3A cloning (3A) of dif P</li> | ||
<li>Restriction digestion of 3' ins in the pSB1AK3 vector ( Using Eco and Xba) for BBA </li> | <li>Restriction digestion of 3' ins in the pSB1AK3 vector ( Using Eco and Xba) for BBA </li> | ||
<li>PCR purification of 3' ins in AK3 ( Using Eco and Xba) for BBA </li> | <li>PCR purification of 3' ins in AK3 ( Using Eco and Xba) for BBA </li> | ||
| - | <li>Set up 5 ml culture of | + | <li>Set up 5 ml culture of pVeg, SpecR-T in pSB1C3 from colony 1 of replica plate and kept in shaking incubator at 37°C </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Restriction digestion of 3' ins in the AK3 vector ( Using Xba and Pst) for 3A | <li>Restriction digestion of 3' ins in the AK3 vector ( Using Xba and Pst) for 3A | ||
| Line 307: | Line 316: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<ul> | <ul> | ||
| Line 315: | Line 324: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Replica plating of transformed colonies having dif P with 3' ins in AK3 -45 sigle colonies were plated</li> | <li>Replica plating of transformed colonies having dif P with 3' ins in AK3 -45 sigle colonies were plated</li> | ||
| Line 322: | Line 331: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of colony PCR from yesterday (first 15 replica plated colonies</li> | <li>Gel analysis of colony PCR from yesterday (first 15 replica plated colonies</li> | ||
| Line 329: | Line 338: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li> | + | <li>Mini prep of 4 overnight cultures - diff P with 3' ins in AK3; colonies 4,5,7 & 9</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 337: | Line 346: | ||
<tr> | <tr> | ||
| - | <td style="background-color:#FFCC66;width:100px;text-align: | + | <td style="background-color:#FFCC66;width:100px;text-align:center;color:#555555;"><b>Afternoon</b> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of PCR purified 3' ins in AK3 and gel purified 3' diff P to work out ratios for the liagation </li> | <li>Gel analysis of PCR purified 3' ins in AK3 and gel purified 3' diff P to work out ratios for the liagation </li> | ||
<li> Dephosphorylation of digested AK3 vector with 3' ins </li> | <li> Dephosphorylation of digested AK3 vector with 3' ins </li> | ||
<li>Overnight ligation of dif P with 3' ins in AK3 (insert and vector) and AK3 containing 3' ins vector only </li> | <li>Overnight ligation of dif P with 3' ins in AK3 (insert and vector) and AK3 containing 3' ins vector only </li> | ||
| - | <li> Set up 100 ml culture of | + | <li> Set up 100 ml culture of pVeg and Spec-T in pSB1C3 by transferring 5 ml culture set up in the morning and kept in shaking incubator at 37°C </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Transformation of E.Coli with ligates from yesterday in AmpR;dif P with 3' ins in AK3 (insert and vector) and AK3 containing 3' ins vector only</li> | + | <li>Transformation of ''E.Coli'' with ligates from yesterday in AmpR; dif P with 3' ins in AK3 (insert and vector) and AK3 containing 3' ins vector only</li> |
<li>Gel extraction of 3' ins (now our insert) described this morning for 3A </li> | <li>Gel extraction of 3' ins (now our insert) described this morning for 3A </li> | ||
<li>Gel analysis of gel extracted dif P and 3' ins (inserts) and pSB1C3 (vector) to work out ratios for the ligation </li> | <li>Gel analysis of gel extracted dif P and 3' ins (inserts) and pSB1C3 (vector) to work out ratios for the ligation </li> | ||
| - | <li> | + | <li>Midi prep of pVeg, SpecR-T in pSB1C3 from colony </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of gel extracted dif P and 3' ins (inserts) and pSB1C3 (vector) to work out ratios for the ligation </li> | <li>Gel analysis of gel extracted dif P and 3' ins (inserts) and pSB1C3 (vector) to work out ratios for the ligation </li> | ||
| - | <li>pveg, SpecR-T in pSB1C3 | + | <li>pveg, SpecR-T in pSB1C3 midi prep sent for sequencing </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Set up overnight 5 ml cultures containing dif P with 3' ins in AK3 for | + | <li>Set up overnight 5 ml cultures containing dif P with 3' ins in AK3 for mini prep tomorrow - 4 cultures were set up by looking at the gel this morning; 2 positive looking (4 & 7), 1 negative (5) and one containing nothing (9</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Diagnostic digests of | + | <li>Diagnostic digests of mini preps containing dif P with 3' ins in AK3 - Two digests : One with SpeI & PstI and other with XbaI & SpeI </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 386: | Line 395: | ||
</html> | </html> | ||
| - | === | + | |} |
| + | |||
| + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Week 9 | ||
| + | |- | ||
| + | | | ||
<html> | <html> | ||
<table width="850px" border="0"> | <table width="850px" border="0"> | ||
| Line 408: | Line 422: | ||
</td> | </td> | ||
| - | <td style="background-color:# | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of Mini preps and diagnostic digests from Saturday - Colony 4 looked best</li> | <li>Gel analysis of Mini preps and diagnostic digests from Saturday - Colony 4 looked best</li> | ||
| Line 414: | Line 428: | ||
</td> | </td> | ||
| - | <td style="background-color:# | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li> | + | <li>Midi prep of Colony 4 - concentration 110 ng/μl </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:# | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<ul> | <ul> | ||
| - | <li>Gel purification of insert (35 | + | <li>Gel purification of insert (35 μl) </li> |
<li>Gel analysis of vector and insert to work out ratios for ligations </li> | <li>Gel analysis of vector and insert to work out ratios for ligations </li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:# | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Transformation with overnight ligations </li> | <li>Transformation with overnight ligations </li> | ||
| Line 434: | Line 448: | ||
</td> | </td> | ||
| - | <td style="background-color:# | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>The transformations were highly successful!! The Vector only plate showed no colonies and the Insert & Vector plate showed many individual colonies </li> | <li>The transformations were highly successful!! The Vector only plate showed no colonies and the Insert & Vector plate showed many individual colonies </li> | ||
| Line 443: | Line 457: | ||
</tr> | </tr> | ||
| - | <td style="background-color:#FFCC66;width:100px;text-align: | + | <td style="background-color:#FFCC66;width:100px;text-align:center;color:#555555;"><b>Afternoon</b> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Set up 200 ml overnight culture of colony 4 (containing Dif P and K09) for midiprep tomorrow </li> | <li>Set up 200 ml overnight culture of colony 4 (containing Dif P and K09) for midiprep tomorrow </li> | ||
| Line 452: | Line 466: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Digests set up for Vector (SpecR) with | + | <li>Digests set up for Vector (SpecR) with SpeI & PstI and Insert (dif P & K09)with XbaI & PstI </li> |
| - | <li>PCR purification of vetor (35 | + | <li>PCR purification of vetor (35 μl)</li> |
<li>Gel extraction of insert after gel analysis (35 ul)</li> | <li>Gel extraction of insert after gel analysis (35 ul)</li> | ||
</ul> | </ul> | ||
| Line 461: | Line 475: | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Dephosphorylation of vector </li> | <li>Dephosphorylation of vector </li> | ||
| Line 468: | Line 482: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Both ligations (Insert & Vector and Vector only) were plated in CmR and incubated overnight </li> | <li>Both ligations (Insert & Vector and Vector only) were plated in CmR and incubated overnight </li> | ||
| Line 474: | Line 488: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of colony PCR products </li> | <li>Gel analysis of colony PCR products </li> | ||
| Line 482: | Line 496: | ||
</table> | </table> | ||
</html> | </html> | ||
| + | |} | ||
| - | === | + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" |
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Week 10 | ||
| + | |- | ||
| + | | | ||
<html> | <html> | ||
| Line 506: | Line 524: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Colony PCR of 10 transformed C-Spec colonies</li> | <li>Colony PCR of 10 transformed C-Spec colonies</li> | ||
| Line 513: | Line 531: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li> | + | <li>Mini prep of C-Spec colonies 2, 5 & 9 in SpecR and CmR</li> |
| - | <li>Digest of | + | <li>Digest of mini preps; double digest (EcoRI and SpeI) and triple digest (PmeI, EcoRI and SpeI)</li> |
| - | <li>Gel analysis of | + | <li>Gel analysis of midi prepped undigested and digested C-Spec</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<ul> | <ul> | ||
| - | <li> | + | <li>Mini prep of C-Spec colonies 2, 5 & 9 in SpecR only and CmR only</li> |
| - | <li>Digest of | + | <li>Digest of mini preps; double digest (EcoRI and SpeI) and triple digest (PmeI, EcoRI and SpeI) </li> |
| - | <li>Gel analysis of | + | <li>Gel analysis of midi prepped undigested and digested C-Spec</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Start | + | <li>Start midi prep of C-Spec from colony 2 (isopropanol added elute was refrigerated for 4 hrs)</li> |
| - | <li>Measure concentration of | + | <li>Measure concentration of midi prepped N-Spec (732 ng/ul)</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Dilution of N-Spec and C-Spec (4x)</li> | <li>Dilution of N-Spec and C-Spec (4x)</li> | ||
| - | <li>Double digest of N-Spec and C-Spec ( | + | <li>Double digest of N-Spec and C-Spec (EcoRI and KpnI) </li> |
<li>Gel analysis of diluted undigested and double digested Specs</li> | <li>Gel analysis of diluted undigested and double digested Specs</li> | ||
| Line 549: | Line 567: | ||
</tr> | </tr> | ||
| - | <td style="background-color:#FFCC66;width:100px;text-align: | + | <td style="background-color:#FFCC66;width:100px;text-align:center;color:#555555;"><b>Afternoon</b> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of colony PCR</li> | <li>Gel analysis of colony PCR</li> | ||
| Line 560: | Line 578: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Start | + | <li>Start midi prep of N-Spec (isopropanol added elute refrigerated overnight)</li> |
<li>Set up 5 ml cultures for C-Spec from colonies 2,5 & 9 (in SpecR only and CmR only)</li> | <li>Set up 5 ml cultures for C-Spec from colonies 2,5 & 9 (in SpecR only and CmR only)</li> | ||
</ul> | </ul> | ||
| Line 568: | Line 586: | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Continue | + | <li>Continue midi prep of N-Spec (by ethanol preciptation)</li> |
| - | <li>Set up a 500 ml overnight culture (with SpecR) of C-Spec from colony 2 for a | + | <li>Set up a 500 ml overnight culture (with SpecR) of C-Spec from colony 2 for a midi prep tomorrow</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Continue | + | <li>Continue midi prep of C-Spec (by ethanol precipitation)</li> |
| - | <li>Digest of | + | <li>Digest of midi prepped N-Spec and C-Spec (EcoRI and SpeI)</li> |
| - | <li>Gel analysis of undigested and digested ( | + | <li>Gel analysis of undigested and digested (EcoRI and KpnI) Specs</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Single digest of diluted C-Spec ( | + | <li>Single digest of diluted C-Spec (EcoRI only)</li> |
| - | <li>Gel analysis of diluted undigested, single digested ( | + | <li>Gel analysis of diluted undigested, single digested (EcoRI only) and double digested (EcoRI and KpnI) </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 595: | Line 613: | ||
| - | === | + | |} |
| + | |||
| + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Week 11 | ||
| + | |- | ||
| + | | | ||
<html> | <html> | ||
| Line 618: | Line 641: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Set up digests of N-Spec and C-Spec ( | + | <li>Set up digests of N-Spec and C-Spec (EcoRI and KpnI)- (CD 1)</li> |
<li>Gel analysis and extraction of both N and C Specs</li> | <li>Gel analysis and extraction of both N and C Specs</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel purification of N and C Specs from CD 2</li> | <li>Gel purification of N and C Specs from CD 2</li> | ||
| Line 632: | Line 655: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<ul> | <ul> | ||
| - | <li>Backbone PCR of pSB1C3 vector using | + | <li>Backbone PCR of pSB1C3 vector using Pfu ultra buffer</li> |
<li>PCR purification of pSB1C3</li> | <li>PCR purification of pSB1C3</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Backbone PCR of pSB1C3 using Barns buffer</li> | <li>Backbone PCR of pSB1C3 using Barns buffer</li> | ||
| Line 647: | Line 670: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel purification of pSB1C3</li> | <li>Gel purification of pSB1C3</li> | ||
| - | <li> | + | <li>Mini prep of 5 ml overnight cultures of colonies 1, 2 and 3</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 656: | Line 679: | ||
</tr> | </tr> | ||
| - | <td style="background-color:#FFCC66;width:100px;text-align: | + | <td style="background-color:#FFCC66;width:100px;text-align:center;color:#555555;"><b>Afternoon</b> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Set up digests of N-Spec and C-Spec again, however, with a higher dilution of N-Spec - (CD 2)</li> | <li>Set up digests of N-Spec and C-Spec again, however, with a higher dilution of N-Spec - (CD 2)</li> | ||
| Line 667: | Line 690: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Dephosphorylate vector (C-Spec) from CD 2</li> | <li>Dephosphorylate vector (C-Spec) from CD 2</li> | ||
| Line 675: | Line 698: | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Transformation of E.Coli with the two Spec final ligates - C-Spec (vector only) and N and C Specs (vector and insert)</li> | + | <li>Transformation of ''E.Coli'' with the two Spec final ligates - C-Spec (vector only) and N and C Specs (vector and insert)</li> |
<li>Gel analysis of purified pSB1C3</li> | <li>Gel analysis of purified pSB1C3</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis and extraction of pSB1C3</li> | <li>Gel analysis and extraction of pSB1C3</li> | ||
| Line 688: | Line 711: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Test digest of final spec | + | <li>Test digest of final spec mini preps with EcoI and Kpn1 and EcoRI and SpeI</li> |
| - | <li>Cloning digest of pSB1C3 with | + | <li>Cloning digest of pSB1C3 with EcoRI and PstI</li> |
| - | <li>Gel analysis of undigested and digested | + | <li>Gel analysis of undigested and digested mini preps and digested pSB1C3</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 698: | Line 721: | ||
</table> | </table> | ||
</html> | </html> | ||
| + | |} | ||
| - | === | + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" |
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Week 12 | ||
| + | |- | ||
| + | | | ||
<html> | <html> | ||
<table width="850px" border="0"> | <table width="850px" border="0"> | ||
| Line 715: | Line 742: | ||
</td> | </td> | ||
<td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Friday</b> | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Friday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Saturday</b> | ||
</td> | </td> | ||
</tr> | </tr> | ||
| Line 721: | Line 750: | ||
</td> | </td> | ||
| - | <td style="background-color:# | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Set up 5 ml culture for | + | <li>Set up 5 ml culture for mini prep of C-Spec of colnies 1 and 2 from synthesis in AmpR</li> |
<li>Set up sequencing mixes for 3' dif P Ins in AK3, C-Spec minis from cultures 5 & 9 and final spec mini</li> | <li>Set up sequencing mixes for 3' dif P Ins in AK3, C-Spec minis from cultures 5 & 9 and final spec mini</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Start | + | <li>Start midi prep of C-Spec</li> |
| - | <li> | + | <li>Mini prep of C-Spec cultures from yesterday</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
| - | <li>Start | + | <li>Start midi prep of C-Spec colonies again</li> |
<ul> | <ul> | ||
<ul> | <ul> | ||
| Line 742: | Line 771: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Cloning digests of C-Spec with Kpn1 and | + | <li>Cloning digests of C-Spec with Kpn1 and PstI and N-Spec with EcoRI and Kpn1</li> |
<li>Gel purification of the inserts</li> | <li>Gel purification of the inserts</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
</ul> | </ul> | ||
| Line 761: | Line 790: | ||
</tr> | </tr> | ||
| - | <td style="background-color:#FFCC66;width:100px;text-align: | + | <td style="background-color:#FFCC66;width:100px;text-align:center;color:#555555;"><b>Afternoon</b> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li> | + | <li>Mini prep cultures of C-Spec have not grown well enough therefore they will be left overnight</li> |
| - | <li>Set up 100 ml culture for | + | <li>Set up 100 ml culture for midi prepping tomorrow</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Test digest C-Spec mini with | + | <li>Test digest C-Spec mini with EcoRI and KpnI</li> |
| - | <li>Run test digest on gel, both minis from colonies 1 & 2 look good, therefore proceed to | + | <li>Run test digest on gel, both minis from colonies 1 & 2 look good, therefore proceed to midi prep</li> |
| - | <li>Continue with | + | <li>Continue with midi prep but no pellet after 45 minute spin</li> |
| - | <li>Set up 100 ml cultures of colonies 1 & 2 for | + | <li>Set up 100 ml cultures of colonies 1 & 2 for midi prep tomorrow</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Continue | + | <li>Continue Midi prep of C-Spec, pellets obtained</li> |
| - | <li>Transformation of E.Coli with the two Spec final ligates - C3 (vector only) and N and C Specs in C3 (vector and insert)</li> | + | <li>Transformation of ''E.Coli'' with the two Spec final ligates - C3 (vector only) and N and C Specs in C3 (vector and insert)</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Ligation gel with inserts; N-Spec ( | + | <li>Ligation gel with inserts; N-Spec (EcoRI & Kpn1) and C-Spec (Kpn1 & PstI) and Vector; pSB1C3 (EcoRI & PstI)</li> |
<li>Dephosphorylation of Vector and overnight ligation of vector alone and vector with inserts</li> | <li>Dephosphorylation of Vector and overnight ligation of vector alone and vector with inserts</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Transformation of E.Coli with the two Spec final ligates - C3 (vector only) and N and C Specs in C3 (vector and insert)</li> | + | <li>Transformation of ''E.Coli'' with the two Spec final ligates - C3 (vector only) and N and C Specs in C3 (vector and insert)</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Transformations failed, re-try next week!</li> | <li>Transformations failed, re-try next week!</li> | ||
| Line 810: | Line 839: | ||
</html> | </html> | ||
| - | === | + | |} |
| + | |||
| + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Week 13 | ||
| + | |- | ||
| + | | | ||
<html> | <html> | ||
<table width="850px" border="0"> | <table width="850px" border="0"> | ||
| Line 826: | Line 860: | ||
</td> | </td> | ||
<td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Friday</b> | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Friday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Saturday</b> | ||
</td> | </td> | ||
</tr> | </tr> | ||
| Line 832: | Line 868: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Set up 5 ml culture for | + | <li>Set up 5 ml culture for mini prep of C-Spec of colnies 1 and 2 from synthesis in AmpR</li> |
<li>Set up sequencing mixes for 3' dif P Ins in AK3, C-Spec minis from cultures 5 & 9 and final spec mini</li> | <li>Set up sequencing mixes for 3' dif P Ins in AK3, C-Spec minis from cultures 5 & 9 and final spec mini</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Start | + | <li>Start midi prep of C-Spec</li> |
| - | <li> | + | <li>Mini prep of C-Spec cultures from yesterday</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
| - | <li>Start | + | <li>Start midi prep of C-Spec colonies again</li> |
<ul> | <ul> | ||
<ul> | <ul> | ||
| Line 853: | Line 889: | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Cloning digests of C-Spec with Kpn1 and | + | <li>Cloning digests of C-Spec with Kpn1 and PstI and N-Spec with EcoRI and Kpn1</li> |
<li>Gel purification of the inserts</li> | <li>Gel purification of the inserts</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
</ul> | </ul> | ||
| Line 872: | Line 908: | ||
</tr> | </tr> | ||
| - | <td style="background-color:#FFCC66;width:100px;text-align: | + | <td style="background-color:#FFCC66;width:100px;text-align:center;color:#555555;"><b>Afternoon</b> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li> | + | <li>Mini prep cultures of C-Spec have not grown well enough therefore they will be left overnight</li> |
| - | <li>Set up 100 ml culture for | + | <li>Set up 100 ml culture for midi prepping tomorrow</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Test digest C-Spec mini with | + | <li>Test digest C-Spec mini with EcoRI and KpnI</li> |
| - | <li>Run test digest on gel, both minis from colonies 1 & 2 look good, therefore proceed to | + | <li>Run test digest on gel, both minis from colonies 1 & 2 look good, therefore proceed to midi prep</li> |
| - | <li>Continue with | + | <li>Continue with midi prep but no pellet after 45 minute spin</li> |
| - | <li>Set up 100 ml cultures of colonies 1 & 2 for | + | <li>Set up 100 ml cultures of colonies 1 & 2 for midi prep tomorrow</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Continue | + | <li>Continue Midi prep of C-Spec, pellets obtained</li> |
| - | <li>Transformation of E.Coli with the two Spec final ligates - C3 (vector only) and N and C Specs in C3 (vector and insert)</li> | + | <li>Transformation of ''E.Coli'' with the two Spec final ligates - C3 (vector only) and N and C Specs in C3 (vector and insert)</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Ligation gel with inserts; N-Spec ( | + | <li>Ligation gel with inserts; N-Spec (EcoRI & Kpn1) and C-Spec (Kpn1 & PstI) and Vector; pSB1C3 (EcoRI & PstI)</li> |
<li>Dephosphorylation of Vector and overnight ligation of vector alone and vector with inserts</li> | <li>Dephosphorylation of Vector and overnight ligation of vector alone and vector with inserts</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Transformation of E.Coli with the two Spec final ligates - C3 (vector only) and N and C Specs in C3 (vector and insert)</li> | + | <li>Transformation of ''E.Coli'' with the two Spec final ligates - C3 (vector only) and N and C Specs in C3 (vector and insert)</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> |
<ul> | <ul> | ||
<li>Transformations failed, re-try next week!</li> | <li>Transformations failed, re-try next week!</li> | ||
| Line 920: | Line 956: | ||
</table> | </table> | ||
</html> | </html> | ||
| + | |} | ||
| + | |||
| + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
| + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Week 14 | ||
| + | |- | ||
| + | | | ||
| + | <html> | ||
| + | <table width="850px" border="0"> | ||
| + | <tr> | ||
| + | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;; font-family: helvetica, arial, sans-serif;color:#555555;"> | ||
| + | <b>Day</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Monday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Tuesday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Wednesday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Thursday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Friday</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Saturday</b> | ||
| + | </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td style="background-color:#FFCC66;width:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Morning</b> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | <li>Digest J23101 XylE C3 with EcoRI and PstI</li> | ||
| + | <li>Gel extract digested vector : C3 EP</li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | <li>Set up gel with N-Spec (EcoRI & Kpn1), C-Spec (Kpn1 & PstI) and C3 (EcoRI & PstI) to work out ligation ratio</li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | <li>Transformations successful!!!! Final Spec vector is available!</li> | ||
| + | <li>Replica plated 3 colonies and set up minis for testing</li> | ||
| + | |||
| + | <ul> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | <li>Min iprep and test digest with EcoRI and Pme1</li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | <li>Midi prep of final Spec vector!! YAY!</li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | </tr> | ||
| + | |||
| + | <td style="background-color:#FFCC66;width:100px;text-align:center;color:#555555;"><b>Afternoon</b> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | <li>Dephosphorylation of Vector and overnight ligation of vector alone and vector with inserts</li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | <li>Transformation of ''E.Coli'' with the two Spec final ligates - C3 (vector only) and N and C Specs in C3 (vector and insert)</li> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | |||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | </ul> | ||
| + | </td> | ||
| + | |||
| + | <td style="background-color:#e7e7e7;height:100px;width:200px;text-align:left;color:#555555;"> | ||
| + | <ul> | ||
| + | </ul> | ||
| + | </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | </html> | ||
| + | |} | ||
| + | |||
| + | * The C3 vector amplified using the C3 template from the registry and subsequently cut with EcoRI and SpeI proved to be inefficient. This could be because the enzymes do not cut the vector properly. Therefore, we obtained the vector from a previous registry part by cutting, extracting and purifying. The ligation of the final spec vector was then successful! | ||
| + | |||
{| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:5px;" cellspacing="20" | ||
| Line 953: | Line 1,102: | ||
# Ligated two single stranded oligos together to produce Dif and insertion site | # Ligated two single stranded oligos together to produce Dif and insertion site | ||
# Standard biobrick assembly of oligos to K14002 | # Standard biobrick assembly of oligos to K14002 | ||
| - | # Ligation and transformation into E.coli competent cells (strain) | + | # Ligation and transformation into ''E.coli'' competent cells (strain) |
# Screen plated colonies for correct insertion | # Screen plated colonies for correct insertion | ||
Next Steps: | Next Steps: | ||
| - | # Purify the correct insert out of E.coli | + | # Purify the correct insert out of ''E.coli'' |
# Next step assembly - LacI test vector and Final assembly vector | # Next step assembly - LacI test vector and Final assembly vector | ||
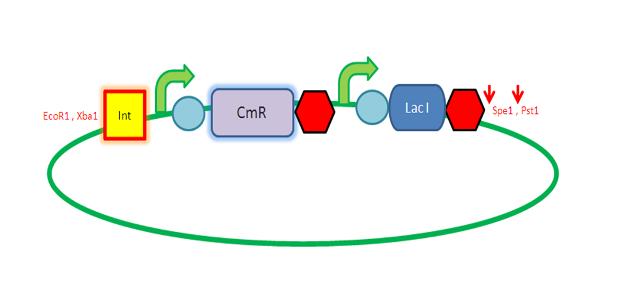
'''LacI Testing vector''' | '''LacI Testing vector''' | ||
| - | Currently waiting for Part B step 2 Midi | + | Currently waiting for Part B step 2 Midi prep results to start this step. |
[[Image:IC_LacI_Test_Vector.JPG |250px|right|alt=A| LacI Testing Vector]] | [[Image:IC_LacI_Test_Vector.JPG |250px|right|alt=A| LacI Testing Vector]] | ||
| Line 997: | Line 1,146: | ||
<td style="background-color:#FFCC66;width:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Morning</b> | <td style="background-color:#FFCC66;width:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Morning</b> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Starting assembly of AmyE vector: | <li>Starting assembly of AmyE vector: | ||
| - | <li>Restriction digestion of BB k14070 for the AmyE vector (using | + | <li>Restriction digestion of BB k14070 for the AmyE vector (using EcoRI and SpeI) |
<li>Restriction digestion of BB k14064 for the AmyE vector (using Eco and Xba) | <li>Restriction digestion of BB k14064 for the AmyE vector (using Eco and Xba) | ||
| - | <li>Restriction digestion of 5' integration site (k08) for the PyrD vector (using | + | <li>Restriction digestion of 5' integration site (k08) for the PyrD vector (using EcoRI and SpeI) |
| - | <li>PCR amplification of vector backbone | + | <li>PCR amplification of vector backbone pSB1C3 for the PyrD vector</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis and extraction of 5' int site for PyrD vector (repetition of step due to absence of DNA during gel analysis yesterday) | <li>Gel analysis and extraction of 5' int site for PyrD vector (repetition of step due to absence of DNA during gel analysis yesterday) | ||
| Line 1,012: | Line 1,161: | ||
<li>Gel purification of k08 | <li>Gel purification of k08 | ||
<li>Re-analysis of k70, k64 (for AmyE) on gel to work out ratios for ligation set up | <li>Re-analysis of k70, k64 (for AmyE) on gel to work out ratios for ligation set up | ||
| - | <li>Re-analysis of k08, | + | <li>Re-analysis of k08, PmeI oligos and pSB1C3 (for PyrD) on gel to work out ratios for ligation set up |
| - | <li>Restriction digestion of | + | <li>Restriction digestion of pVeg promoter (k53) (using EcoRI and SpeI) and spec cassette (k65) (using XbaI and PstI) for PyrD vector |
| - | <li>Restriction digestion of pSB1C3 for PyrD (using | + | <li>Restriction digestion of pSB1C3 for PyrD (using EcoRI and PstI) </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Check for transformed colonies (colonies that have taken up the vector with the 5'diff) and prepare for colony PCR | <li>Check for transformed colonies (colonies that have taken up the vector with the 5'diff) and prepare for colony PCR | ||
| Line 1,024: | Line 1,173: | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Transformation of overnight ligations of: k64 and k70, k70 only, Spec and 5' PyrD diff</li> | <li>Transformation of overnight ligations of: k64 and k70, k70 only, Spec and 5' PyrD diff</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Replica plating and colony PCR of: Spec and 5' PyrD diff | <li>Replica plating and colony PCR of: Spec and 5' PyrD diff | ||
| Line 1,039: | Line 1,188: | ||
<td style="background-color:#FFCC66;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Afternoon</b> | <td style="background-color:#FFCC66;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Afternoon</b> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>PCR purification of PSB1C3 vector | <li>PCR purification of PSB1C3 vector | ||
| Line 1,047: | Line 1,196: | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Restriction digestion of k64 and subsequent PCR purification (repetition of step due to absence of DNA during gel analysis) | <li>Restriction digestion of k64 and subsequent PCR purification (repetition of step due to absence of DNA during gel analysis) | ||
| Line 1,053: | Line 1,202: | ||
<li>Gel analysis and extraction of k53 and k65 | <li>Gel analysis and extraction of k53 and k65 | ||
<li>Bench (1 hour) and overnight ligation of 5'diff with the pSB1C3 vector | <li>Bench (1 hour) and overnight ligation of 5'diff with the pSB1C3 vector | ||
| - | <li>Transformation of E.Coli with the bench ligated vector </li> | + | <li>Transformation of ''E.Coli'' with the bench ligated vector </li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Dephosphorylation of k64 and set up of overnight ligations for k64 and k70 (vector and insert) and k64 (vector)only (for negative control; check of background) | <li>Dephosphorylation of k64 and set up of overnight ligations for k64 and k70 (vector and insert) and k64 (vector)only (for negative control; check of background) | ||
| Line 1,062: | Line 1,211: | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of colony PCR products of Spec and 5' PyrD diff | <li>Gel analysis of colony PCR products of Spec and 5' PyrD diff | ||
| Line 1,102: | Line 1,251: | ||
<td style="background-color:#FFCC66;width:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Morning</b> | <td style="background-color:#FFCC66;width:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Morning</b> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Set up overnight ligations for standard assembly (BBA) and 3A cloning (3A) of dif P | <li>Set up overnight ligations for standard assembly (BBA) and 3A cloning (3A) of dif P | ||
| - | <li>Restriction digestion of 3' integration site (K02) in the A2 vector ( Using | + | <li>Restriction digestion of 3' integration site (K02) in the A2 vector ( Using EcoRI and XbaI) for BBA |
| - | <li>Restriction digestion of 3' integration site (K09) in the AK3 vector ( Using | + | <li>Restriction digestion of 3' integration site (K09) in the AK3 vector ( Using EcoRI and XbaI) for BBA |
| - | <li>PCR purification of 3' integration site (K02) in the A2 vector ( Using | + | <li>PCR purification of 3' integration site (K02) in the A2 vector ( Using EcoRI and XbaI) for BBA |
| - | <li>PCR purification of 3' integration site (K09) in the AK3 vector ( Using | + | <li>PCR purification of 3' integration site (K09) in the AK3 vector ( Using EcoRI and XbaI) for BBA |
| - | <li>Set up 5 ml culture of spec from colony 1 of replica plate in shaking incubator @ | + | <li>Set up 5 ml culture of spec from colony 1 of replica plate in shaking incubator @ 37°C</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Restriction digestion of 3' integration site (K02) in the A2 vector ( Using | + | <li>Restriction digestion of 3' integration site (K02) in the A2 vector ( Using XbaI and PstI) for 3A |
| - | <li>Restriction digestion of 3' integration site (K09) in the AK3 vector ( Using | + | <li>Restriction digestion of 3' integration site (K09) in the AK3 vector ( Using XbaI and PstI) for 3A</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Transformations using the overnight ligations showed a lot of background. Therefore we set up ligations for K02 and K09 using 3A cloning. The results will show if this method is preferable due to less background. | <li>Transformations using the overnight ligations showed a lot of background. Therefore we set up ligations for K02 and K09 using 3A cloning. The results will show if this method is preferable due to less background. | ||
| - | <li> | + | <li>Midi prep of CmR vector and test digest using EcoRI and SpeI |
| - | <li>Backbone PCR of | + | <li>Backbone PCR of pSB1C3 vector (1st attempt) for common use</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Replica plating of transformed colonies for k09 from the plate with the insert (diff P) - 45 sigle colonies were plated | <li>Replica plating of transformed colonies for k09 from the plate with the insert (diff P) - 45 sigle colonies were plated | ||
| Line 1,131: | Line 1,280: | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of colony PCR from yesterday (first 15 replica plated colonies | <li>Gel analysis of colony PCR from yesterday (first 15 replica plated colonies | ||
| Line 1,140: | Line 1,289: | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
| - | <li> | + | <li>Mini prep of 4 overnight cultures (dif P and 3' Insert for PyrD) - colonies 4,5,7 & 9 |
<li>Run Colony PCR results on a Gel - pick promissing candidates for mini prep.</li> | <li>Run Colony PCR results on a Gel - pick promissing candidates for mini prep.</li> | ||
</ul> | </ul> | ||
| Line 1,150: | Line 1,299: | ||
<td style="background-color:#FFCC66;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Afternoon</b> | <td style="background-color:#FFCC66;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Afternoon</b> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of PCR purified K02 and K09 with the insert (diff P) in between to work out ratios for the liagation | <li>Gel analysis of PCR purified K02 and K09 with the insert (diff P) in between to work out ratios for the liagation | ||
| Line 1,158: | Line 1,307: | ||
<li>Set up overnight ligation of AK3 vector with 3' integration site (K09) with diff P (insert) and AK3 vector only | <li>Set up overnight ligation of AK3 vector with 3' integration site (K09) with diff P (insert) and AK3 vector only | ||
<li>Colony PCR and gel analysis of plated culture (from plate wash on Friday) with K64 and K70 | <li>Colony PCR and gel analysis of plated culture (from plate wash on Friday) with K64 and K70 | ||
| - | <li>Overnight 100 ml culture of spec @ | + | <li>Overnight 100 ml culture of spec @ 14°C</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Electroporation of the 4 overnight ligations described on Thursday afternoon | <li>Electroporation of the 4 overnight ligations described on Thursday afternoon | ||
| Line 1,170: | Line 1,319: | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>PCR purification of PSB1C3 PCR amplified vector | <li>PCR purification of PSB1C3 PCR amplified vector | ||
<li>Check concentration of midi prepped CmR | <li>Check concentration of midi prepped CmR | ||
| - | <li>Run a gel to visualise the results - Gel contained pSB1C3 (PCR purified) , CmR ( | + | <li>Run a gel to visualise the results - Gel contained pSB1C3 (PCR purified) , CmR (Midi prepped) and CmR digested (Midi prepped)</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Backbone PCR of pSB1C3 (2nd attempt), PCR purified and then digested with | + | <li>Backbone PCR of pSB1C3 (2nd attempt), PCR purified and then digested with EcoRI and PstI |
| - | <li> | + | <li>Midi preps sent for sequencing ( Spec and CmR)</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Backbone PCR of pSB1C3 using | + | <li>Backbone PCR of pSB1C3 using Pfu (3rd attempt) |
| - | <li>Set up overnight 5 ml cultures for | + | <li>Set up overnight 5 ml cultures for mini prepping tomorrow - 4 cultures were set up by looking at the gel this morning; 2 positive looking (4 & 7), 1 negative (5) and one containing nothing (9) </li> |
</td> | </td> | ||
</ul> | </ul> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Diagnostic digests of | + | <li>Diagnostic digests of mini preps - Two digests : One with SpeI & PstI and other with XbaI & SpeI</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| Line 1,223: | Line 1,372: | ||
<td style="background-color:#FFCC66;width:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Morning</b> | <td style="background-color:#FFCC66;width:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Morning</b> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of Mini preps and diagnostic digests from Saturday - Colony 4 looked best | <li>Gel analysis of Mini preps and diagnostic digests from Saturday - Colony 4 looked best | ||
| Line 1,229: | Line 1,378: | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Midiprep of Colony 4 - concentration 110 ng/ | + | <li>Midiprep of Colony 4 - concentration 110 ng/μl |
<li>Restriction digest of K02 from registrty for 3A assembly</li> | <li>Restriction digest of K02 from registrty for 3A assembly</li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel purification of insert (35 ul) | <li>Gel purification of insert (35 ul) | ||
| Line 1,241: | Line 1,390: | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li> Transformation with overnight ligations | <li> Transformation with overnight ligations | ||
| Line 1,248: | Line 1,397: | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align: | + | <td style="background-color:#e7e7e7;height:100px;width:150px;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>The transformations were highly successful!! The '''Vector only''' plate showed no colonies and the '''Insert & Vector''' plate showed many individual colonies | <li>The transformations were highly successful!! The '''Vector only''' plate showed no colonies and the '''Insert & Vector''' plate showed many individual colonies | ||
| Line 1,259: | Line 1,408: | ||
<td style="background-color:#FFCC66;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Afternoon</b> | <td style="background-color:#FFCC66;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Afternoon</b> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
| - | <li>Set up 200 ml overnight culture of colony 4 (containing Dif P and K09) for | + | <li>Set up 200 ml overnight culture of colony 4 (containing Dif P and K09) for midi prep tomorrow |
| - | <li>Second Test digest SpeI | + | <li>Second Test digest SpeI & PmeI - no positive results. Decided to repeat the step using 3A assembly method to reduce background from vector.</li> |
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
| - | <li> Digests set up for Vector (SpecR) with | + | <li> Digests set up for Vector (SpecR) with SpeI & PstI and Insert (dif P & K09) with XbaI & PstI |
<li> PCR purification of vetor (35 ul) | <li> PCR purification of vetor (35 ul) | ||
<li> Gel extraction of insert after gel analysis (35 ul) | <li> Gel extraction of insert after gel analysis (35 ul) | ||
| Line 1,273: | Line 1,422: | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Dephosphorylation of vector | <li>Dephosphorylation of vector | ||
| Line 1,279: | Line 1,428: | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Both ligations ('Insert & Vector' and 'Vector only') were plated in CmR and incubated overnight </li> | <li>Both ligations ('Insert & Vector' and 'Vector only') were plated in CmR and incubated overnight </li> | ||
</ul> | </ul> | ||
</td> | </td> | ||
| - | <td style="background-color:#e7e7e7;text-align: | + | <td style="background-color:#e7e7e7;text-align:left;font-family: helvetica, arial, sans-serif;color:#555555;"> |
<ul> | <ul> | ||
<li>Gel analysis of colony PCR products | <li>Gel analysis of colony PCR products | ||
<li>Set up ligation reaction K02+dif using 3A method for Transformation on Monday.</li> | <li>Set up ligation reaction K02+dif using 3A method for Transformation on Monday.</li> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
</ul> | </ul> | ||
</td> | </td> | ||
Latest revision as of 03:54, 28 October 2010
| Lab Diaries | Overview | Surface Protein Team | XylE Team | Vectors Team | Modelling Team |
| Here are the technical diaries for our project. We've split them up into three lab teams and the modelling team. We think it's really important that absolutely anyone can find out what we've been doing. For a really detailed look at what we did, and when, you've come to the right place! | |
| Objectives |
|
We are assembling the AmyE and PyrD vectors in order to transform B. Subtilis with our parts. Once completed, these vectors will be reusable and can then be used to introduce any relevant piece of DNA directly into B.subtilis genome. The vectors will be used both for the final assembly and for testing constructs. |
| PyrD Vector |
| Week 6 | |||||||||||||||||||||
|
Thursday, August 12
The forward and reverse strands of the 5' dif site with XbaI and PstI restriction sites on either side have been synthesized separately. The synthesized fragments arrive in solid powder form. These were immediately diluted in ddH2O to obtain a stock concentration of 1 ng/μl. They were then allowed to anneal together by first heating them to 95°C for denaturation and allowing them to cool down and anneal overnight. Friday, August 13
After annealing the two strands, the oligo was cut with XbaI and PstI to obtain overhangs that would later ligate with the compatible overhangs of a cut vector.
The cut oligo was PCR purified in order to get rid of any contaminants. PCR purification gets rid of short pieces of DNA which are less than about 40 base pairs.
|
| Week 7 | ||||||||||||||||||
|
K143008 is the 5' integration site for the PyrD vector. This will be used as a front insert together with the dif XP for the pSB1C3 vector.
The pSB1C3 vector backbone from the registry was amplified with the use of SB3 and SB2a primers. Submission of parts to the registry requires them to be in a pSB1C3 vector therefore any parts to be submitted will be inserted into this vector.
The PCR amplified vector was purified in order to get rid of any contaminants. For example, short pieces of DNA like the primers. Tuesday, August 17
The digested 5' ins was first analyzed on the gel to verify it's size and then extracted for purification. The 5' ins was gel purified in order to extract only the relevant piece of DNA.
The pSB1C3 vector was digested so that it would contain compatible overhangs for ligation with inserts.
The digested pSB1C3 was re-purified in order to get rid of any contaminant DNA that arose during the digestion.
pVeg (promoter and RBS) and Spec-T (Spectinomycin with a terminator) were digested in preparation for 3A assembly.
The digested 5' ins and dif (the front inserts) were ligated overnight with pSB1C3 (the vector). A bench ligation and an overnight ligation were set up.
E.Coli was transformed via the chemical method using the bench ligate.
Since these are both inserts they were gel extracted and purified. PCR purification is not carried out for inserts since they are small and would therefore be lost during the process. Wednesday, August 18
pVeg and SpecR-T (the front inserts) were ligated overnight with pSB1C3 (the vector). Thursday, August 19
Friday, August 20
The forward and reverse strands of the 3' dif Pme1 sites with XbaI and PstI restriction sites on either side have been synthesized separately. The synthesized fragments arrive in solid powder form. These were immediately diluted in ddH2O to obtain a stock concentration of 1 ng/μl. They were then allowed to anneal together by heating them to 95°C for denaturation and allowing them to cool down and anneal overnight. |
| Week 8 | |||||||||||||||||||||
|
|
| Week 9 | ||||||||||||||||||
|
|
| Week 10 | ||||||||||||||||||
|
|
| Week 11 | ||||||||||||||||||
|
|
| Week 12 | |||||||||||||||||||||
|
|
| Week 13 | |||||||||||||||||||||
|
|
| Week 14 | |||||||||||||||||||||
|
|
- The C3 vector amplified using the C3 template from the registry and subsequently cut with EcoRI and SpeI proved to be inefficient. This could be because the enzymes do not cut the vector properly. Therefore, we obtained the vector from a previous registry part by cutting, extracting and purifying. The ligation of the final spec vector was then successful!
| AmyE Vector |
|
Starting from the top, we are assembling the first two fragments (K14070 and K14064) and K147002 with our oligos to add in a Dif site. Two dif sites on either side of resistance cassettes can be used to later excise antibiotic resistance from our final constructs.
Next Steps:
K14002 and oligos
Next Steps:
|
| Schedule & Lab Notes |
| Week 7 | ||||||||||||||||||
|
|
| Week 8 | |||||||||||||||||||||
|
|
| Week 9 | ||||||||||||||||||
|
|
 "
"