Project/Lac1/AraC Promoter/
From 2010.igem.org
| (16 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | + | {{Template:WITS-South_Africa_Main_Menu_19_07}} | |
| + | {{Template:WITS-South_Africa_Main_Menu_0910}} | ||
| + | == Lac/Ara-1 IPTG-Inducible Promoter - [http://partsregistry.org/Part:BBa_K354000 BBa_K354000] == | ||
=== Background === | === Background === | ||
| + | While the string of open reading frames lie mute within Lacto-sense, the trigger that errs it into motion upon sighting IPTG, the chosen proxy for HPV presence, is the Lac/Ara-1 promoter. This fusion promoter recruits elements from promoters belonging to both the Lactose and Arabinose operons to create a synthetic IPTG (or appropriate Arabinose isomer) -inducible promoter. The promoter was synthesized as described by Lutz and Bujard [1] via annealed primers (that share complementary sequences of overlap) and their subsequent elongation via PCR growth. | ||
<br /> | <br /> | ||
| - | === Sequence and Annotation === | + | <br /> |
| + | Derived from the naturally-occurring L-arabinose operon in '''E. coli'', the AraC homodimer posses a specific DNA-binding domain that binds the araI sequence within the promoter [2]. AraC adhesion to this sequence contorts the promoter into a loop structure thus preventing the RNA polymerase from binding and initiating transcription. In the presence of arabinose, or a suitably-similar isomer, AraC undergoes a conformational shift thus altering its DNA binding domain and allowing RNA polymerase to bind the promoter. However, in the case of our synthetic inducible promoter, the araI sequence has been strategically placed adjacently upstream of the -33 hexamer that has been flagged as a RNA polymerase binding site. In the absence of arabinose, AraC-araI adhesion will impede RNA polymerase binding thus inhibiting transcription. The conformational alteration in the presence of arabinose, or an isomer, will ‘free’ AraC from its sequence and allow transcription to occur. | ||
| + | <br /> | ||
| + | <br /> | ||
| + | IPTG - isopropyl-β-D-1-thiogalactopyranoside - is an isomer of β-galactosidase that is structurally proximal enough to arabinose so as to induce the required conformational shift in AraC and hence induce transcription. Furthermore, due to the presence of the sulfur atom on the ring structure, IPTG is unable to be catabolised by the cell and thus remains at a constant concentration [3]. Due to this inherent latency of IPTG, the positive pressure that it exerts on transcription remains constant. Thus, addition of exogenous Arabinose or IPTG will induce polymerase engagement with the promoter and subsequent transcription of the downstream open reading frames. Due to the duality in activation of both Arabinose and IPTG, promoter activity can be activated to differing degrees by the selective addition of either or both inducers and hence activity can be finely regulated via appropriate dose administration. | ||
| + | <br /> | ||
| + | |||
| + | === Sequence Developement and Annotation === | ||
| + | <div class="heading"> | ||
| + | '''Promoter Assembly''' | ||
| + | </div> | ||
| + | Specific forward and reverse primers were ordered that shared a 17bp region overlap so as to ensure appropriate annealing to one another. | ||
| + | <br /> | ||
| + | [[Image:Primers _annealing.JPG]] | ||
| + | <br /> | ||
| + | Once annealed to one another, the primers are extended by 10 rounds of PCR so as to yield the full double-stranded promoter sequence. | ||
| + | <br /> | ||
| + | [[Image:Primer_extension.JPG]] | ||
| + | <br /> | ||
| + | Due to the activity of Taq polymerase, each of the (now complete) primers posses a single adenine overhang on their 3’ end. The pTZ57R Fermentas TA cloning vector possesses a complementary single thymine overhang in its multiple cloning site. This facilitates the insertion and ligation of the promoter fragment into the vector. | ||
| + | <br /> | ||
| + | [[Image:Extended_primers.JPG]] | ||
| + | <br /> | ||
| + | [[Image:PTZ_MCS.JPG]] | ||
| + | <br /> | ||
| + | Once ligated, the plasmid is transformed into competent bacteria and cloned. | ||
| + | <br /> | ||
| + | [[Image:Ligated_Promoter.JPG]] | ||
| + | <br /> | ||
| + | |||
| + | <div class="heading"> | ||
| + | '''Promoter Annotation''' | ||
| + | </div > | ||
| + | Furthermore, in accordance with the Standard Assembly protocols set about by the Registry, each primer, forward and reverse, was designed to include the sequences recognised by the restriction endonucleases EcoR1, Xba1, Spe1 and Pst1. | ||
| + | <br /> | ||
| + | The forward Lac/Ara-1 promoter sequence is: | ||
| + | <br /> | ||
| + | [[Image:Lac_forward_primer_1.JPG]] | ||
| + | <br /> | ||
| + | Where the sequence in red denotes the EcoR1 site, blue the Xba1 site and purple being the region of overlap shared with the reverse primer. | ||
| + | <br /> | ||
| + | <br /> | ||
| + | The reverse Lac/Ara-1 promoter sequence is: | ||
| + | <br /> | ||
| + | [[Image:Lac_reverse_primer_1.JPG]] | ||
| + | <br /> | ||
| + | Where the sequence in green denotes the Pst1 site, orange the Spe1 site and purple the region of overlap. | ||
| + | <br /> | ||
| + | <br /> | ||
| + | The complete, synthetic Lac/Ara-1 promoter sequence (without flanking restriction sites) is: <br /> | ||
| + | [[Image:Lac_promoter_full.JPG]] | ||
| + | <br /> | ||
| + | The sequence in red indicates the region where the AraC repressor protein binds. The sequences in blue represent the hexamers recognised, and subsequently bound, by RNA polymerase in the absence of AraC and the region in green is the operator sequence derived from the lac operon. | ||
| + | |||
| + | |||
| + | <div class="heading"> | ||
| + | '''The package''' | ||
| + | </div> | ||
| + | While in the process of development, the Lac/Ara-1 promoter was flipped in and out of vectors at a rate second to none. However, what is of use to you, the budding synthetic biologist, is the format in which you will receive our BioBrick. So here it is. In all it's glory, in the Standard Assembly PSB1C3 plasmid backbone (exhibiting chloramphenicol resistance) with appropriate annotations of Standard Assembly restriction sites as well as key plasmid features. Illustration courtesy Dr Marco Weinberg [4]. | ||
| + | |||
| + | |||
| + | [[Image:LacAraC_in_PSB1C3.JPG]] | ||
<br /> | <br /> | ||
=== Testing and Validation === | === Testing and Validation === | ||
| + | Once ligated into PSB1C3, the construct was digested with EcoR1 and Pst1 so as to confirm. The following screen was performed on a 2% (m/v) agarose gel. <br /> | ||
| + | [[Image:Lac_confirmation_gel.JPG]] <br /> <br /> | ||
| + | The above gel shows 3 clones chosen and screened, of which only 1 (that in lane 1) is positive. The PSB1C3 linearised plasmid is shown (2072bp) as well as the Lac/Ara-1 insert (154bp). | ||
| + | |||
| + | <br /> | ||
| + | === References === | ||
| + | [1] Lutz R. & Bujard H. (1997), Independent and tight regulation of transcriptional units in Escherchia coli via the LacR/O, the TetR/O and AraC/I_1- I_2 regulatory elements. Nucleic Acids Research 25: 1203-1210. | ||
| + | <br /> | ||
| + | [2] Schleif R. (2000), Regulation of the L-arabinose operon in Escherichia coli. Trends in Genetics 16: 559-565. | ||
| + | <br /> | ||
| + | [3] Hansen L.S., Knudsen S. & Sorensen S.J. (1998), The Effect of the lacY Gene on the Induction of IPTG Inducible Promoters, Studies in Escherichia coli and Pseudomonas fluorescens. Current Microbiology 36: 341-347. | ||
<br /> | <br /> | ||
| + | [4] Marco Weinberg, PhD. Antiviral Gene Therapy Research Unit, Department of Molecular Medicine and Haematology, University of the Witwatersrand Medical School, 7 York Rd. Parktown 2193 SOUTH AFRICA, marc.weinberg@wits.ac.za. | ||
Latest revision as of 20:06, 25 October 2010

Contents |
Lac/Ara-1 IPTG-Inducible Promoter - [http://partsregistry.org/Part:BBa_K354000 BBa_K354000]
Background
While the string of open reading frames lie mute within Lacto-sense, the trigger that errs it into motion upon sighting IPTG, the chosen proxy for HPV presence, is the Lac/Ara-1 promoter. This fusion promoter recruits elements from promoters belonging to both the Lactose and Arabinose operons to create a synthetic IPTG (or appropriate Arabinose isomer) -inducible promoter. The promoter was synthesized as described by Lutz and Bujard [1] via annealed primers (that share complementary sequences of overlap) and their subsequent elongation via PCR growth.
Derived from the naturally-occurring L-arabinose operon in 'E. coli, the AraC homodimer posses a specific DNA-binding domain that binds the araI sequence within the promoter [2]. AraC adhesion to this sequence contorts the promoter into a loop structure thus preventing the RNA polymerase from binding and initiating transcription. In the presence of arabinose, or a suitably-similar isomer, AraC undergoes a conformational shift thus altering its DNA binding domain and allowing RNA polymerase to bind the promoter. However, in the case of our synthetic inducible promoter, the araI sequence has been strategically placed adjacently upstream of the -33 hexamer that has been flagged as a RNA polymerase binding site. In the absence of arabinose, AraC-araI adhesion will impede RNA polymerase binding thus inhibiting transcription. The conformational alteration in the presence of arabinose, or an isomer, will ‘free’ AraC from its sequence and allow transcription to occur.
IPTG - isopropyl-β-D-1-thiogalactopyranoside - is an isomer of β-galactosidase that is structurally proximal enough to arabinose so as to induce the required conformational shift in AraC and hence induce transcription. Furthermore, due to the presence of the sulfur atom on the ring structure, IPTG is unable to be catabolised by the cell and thus remains at a constant concentration [3]. Due to this inherent latency of IPTG, the positive pressure that it exerts on transcription remains constant. Thus, addition of exogenous Arabinose or IPTG will induce polymerase engagement with the promoter and subsequent transcription of the downstream open reading frames. Due to the duality in activation of both Arabinose and IPTG, promoter activity can be activated to differing degrees by the selective addition of either or both inducers and hence activity can be finely regulated via appropriate dose administration.
Sequence Developement and Annotation
Promoter Assembly
Specific forward and reverse primers were ordered that shared a 17bp region overlap so as to ensure appropriate annealing to one another.
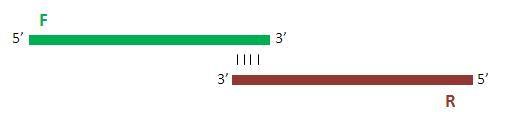
Once annealed to one another, the primers are extended by 10 rounds of PCR so as to yield the full double-stranded promoter sequence.
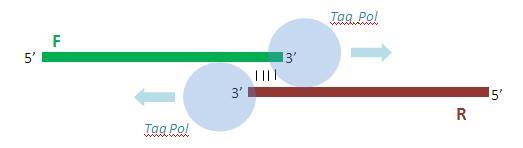
Due to the activity of Taq polymerase, each of the (now complete) primers posses a single adenine overhang on their 3’ end. The pTZ57R Fermentas TA cloning vector possesses a complementary single thymine overhang in its multiple cloning site. This facilitates the insertion and ligation of the promoter fragment into the vector.
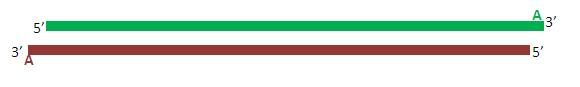

Once ligated, the plasmid is transformed into competent bacteria and cloned.
Promoter Annotation
Furthermore, in accordance with the Standard Assembly protocols set about by the Registry, each primer, forward and reverse, was designed to include the sequences recognised by the restriction endonucleases EcoR1, Xba1, Spe1 and Pst1.
The forward Lac/Ara-1 promoter sequence is:
Where the sequence in red denotes the EcoR1 site, blue the Xba1 site and purple being the region of overlap shared with the reverse primer.
The reverse Lac/Ara-1 promoter sequence is:
Where the sequence in green denotes the Pst1 site, orange the Spe1 site and purple the region of overlap.
The complete, synthetic Lac/Ara-1 promoter sequence (without flanking restriction sites) is:
The sequence in red indicates the region where the AraC repressor protein binds. The sequences in blue represent the hexamers recognised, and subsequently bound, by RNA polymerase in the absence of AraC and the region in green is the operator sequence derived from the lac operon.
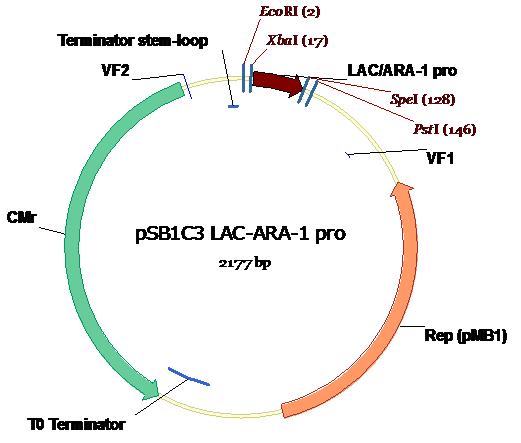
The package
While in the process of development, the Lac/Ara-1 promoter was flipped in and out of vectors at a rate second to none. However, what is of use to you, the budding synthetic biologist, is the format in which you will receive our BioBrick. So here it is. In all it's glory, in the Standard Assembly PSB1C3 plasmid backbone (exhibiting chloramphenicol resistance) with appropriate annotations of Standard Assembly restriction sites as well as key plasmid features. Illustration courtesy Dr Marco Weinberg [4].
Testing and Validation
Once ligated into PSB1C3, the construct was digested with EcoR1 and Pst1 so as to confirm. The following screen was performed on a 2% (m/v) agarose gel.
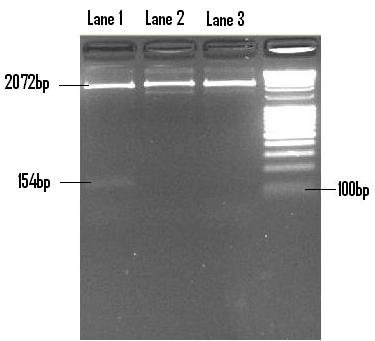
The above gel shows 3 clones chosen and screened, of which only 1 (that in lane 1) is positive. The PSB1C3 linearised plasmid is shown (2072bp) as well as the Lac/Ara-1 insert (154bp).
References
[1] Lutz R. & Bujard H. (1997), Independent and tight regulation of transcriptional units in Escherchia coli via the LacR/O, the TetR/O and AraC/I_1- I_2 regulatory elements. Nucleic Acids Research 25: 1203-1210.
[2] Schleif R. (2000), Regulation of the L-arabinose operon in Escherichia coli. Trends in Genetics 16: 559-565.
[3] Hansen L.S., Knudsen S. & Sorensen S.J. (1998), The Effect of the lacY Gene on the Induction of IPTG Inducible Promoters, Studies in Escherichia coli and Pseudomonas fluorescens. Current Microbiology 36: 341-347.
[4] Marco Weinberg, PhD. Antiviral Gene Therapy Research Unit, Department of Molecular Medicine and Haematology, University of the Witwatersrand Medical School, 7 York Rd. Parktown 2193 SOUTH AFRICA, marc.weinberg@wits.ac.za.
 "
"