Team:Lethbridge/Results
From 2010.igem.org
Liszabruder (Talk | contribs) |
Liszabruder (Talk | contribs) |
||
| Line 127: | Line 127: | ||
<BLOCKQUOTE> | <BLOCKQUOTE> | ||
| + | |||
| + | =<font color="white">Catechol Degradation= | ||
| + | |||
| + | The focus of our <html><a href="https://2010.igem.org/Team:Lethbridge/Project"><font color="#00DC00">project</font></a></html> is to decrease the toxicity of tailing pond water through bioremediation. We are specifically interested in the <html><a href="https://2010.igem.org/Team:Lethbridge/Project/Catechol_Degradation"><font color="#00DC00">degradation of the toxic molecule catechol</font></a></html> into 2-hydroxymuconate semialdyhyde (2-HMS); a bright yellow substrate that can be metabolized by the cell. This conversion is accomplished by catechol 2,3-dioxygenase (xylE). | ||
| + | |||
| + | ===<font color="white">Characterized Parts</font>=== | ||
| + | <hr> | ||
| + | <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_118021" target="new"><font color="#00DC00" size="+1">BBa_118021</font></a></html> | ||
| + | <br><br> | ||
| + | |||
| + | ===<font color="white">Hypothesis</font>=== | ||
| + | <hr> | ||
| + | The addition of the catechol to a solution of xylE will result in the production of 2-HMS. | ||
| + | <br><br> | ||
| + | |||
| + | ===<font color="white">Method</font>=== | ||
| + | <hr> | ||
| + | <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_118021" target="new"><font color="#00DC00">BBa_118021</font></a></html> was transformed into <i>Escherichia coli</i> DH5α (E. coli) cells using our transformation protocol. We confirmed that the cells hosted the plasmid containing the BBa_118021 part and we begun experiments to characterize catechol degredation. | ||
| + | <br><br> | ||
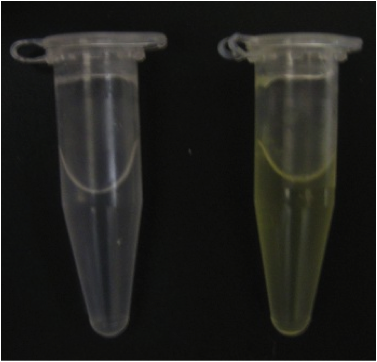
| + | In our first experiment we grew a 5 mL culture of our engineered <i>E. coli cells</i> in M9 minimal media overnight. The cells were spun down at 14000 rfc for 2 minutes. We raised the catechol concentration of this solution to 100 mM, the solution immediately turned bright yellow. | ||
| + | <br><br> | ||
| + | [[image:UofLcatecholfigure1.jpg]] | ||
| + | <html> | ||
| + | <body> | ||
| + | <center> | ||
| + | <table border="0" cellpadding="8" width="28%" style="background-color:#000000"> | ||
| + | |||
| + | <tr> | ||
| + | <th> | ||
| + | <img src=""/> | ||
| + | </th> | ||
| + | </tr> | ||
| + | |||
| + | </tr> | ||
| + | <tr><th colspan="1" align="left"> | ||
| + | <p> | ||
| + | <font color="white">Figure 1. Left: M9 media spiked with 100 mM catechol. The solution contains <i>E. coli</i> cells hosting the pUC19 plasmid. Right: M9 media spiked with 100 mM catechol. The solution contains <i>E. coli</i> cells hosting part BBa_118021. The yellow colour suggests the production of 2-HMS. | ||
| + | </th> | ||
| + | </tr> | ||
| + | |||
| + | </table> | ||
| + | </center> | ||
| + | </body> | ||
| + | </html> | ||
| + | |||
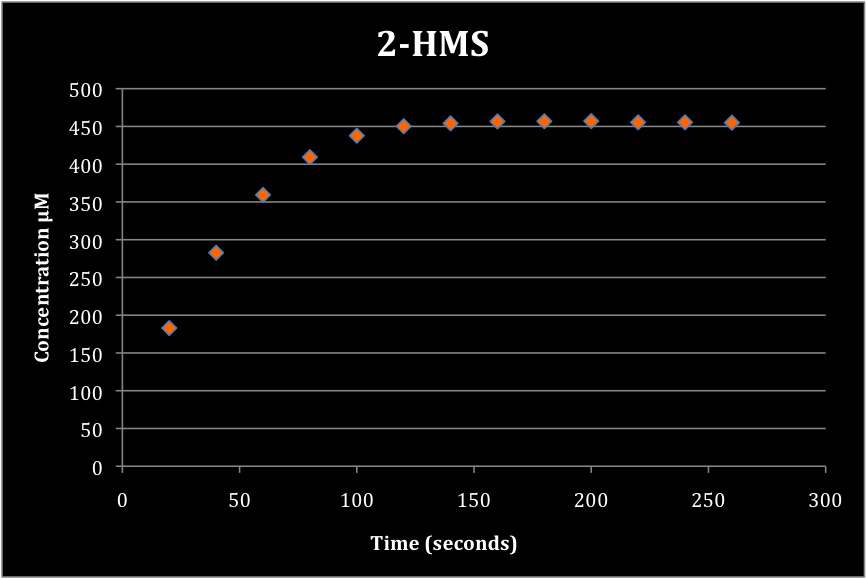
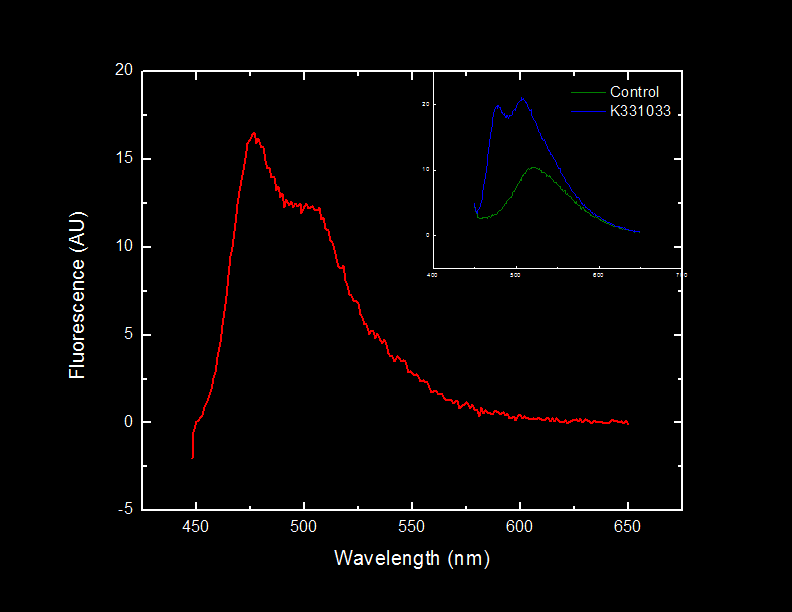
| + | In our second experiment we wanted to measure the absorbance of 2-hydroxymuconate semialdehyde over time. We grew a 5 mL culture of our engineered <i>E. coli</i> cells in M9 media overnight. 1 mL of the cell solution was removed for analysis. The cell solution was raised to 100 mM catachol. Immediately the formation of 2-HMS was tracked. The formation of 2-HMS can easily be tracked, as it absorbs light at 375 nm. | ||
| + | <br><br> | ||
| + | [[image:UofLcatecholfigure2.jpg]] | ||
| + | <html> | ||
| + | <body> | ||
| + | <center> | ||
| + | <table border="0" cellpadding="8" width="28%" style="background-color:#000000"> | ||
| + | |||
| + | <tr> | ||
| + | <th> | ||
| + | <img src=""/> | ||
| + | </th> | ||
| + | </tr> | ||
| + | |||
| + | </tr> | ||
| + | <tr><th colspan="1" align="left"> | ||
| + | <p> | ||
| + | <font color="white">Figure 2. Production of 2-HMS over time. | ||
| + | </th> | ||
| + | </tr> | ||
| + | |||
| + | </table> | ||
| + | </center> | ||
| + | </body> | ||
| + | </html> | ||
| + | |||
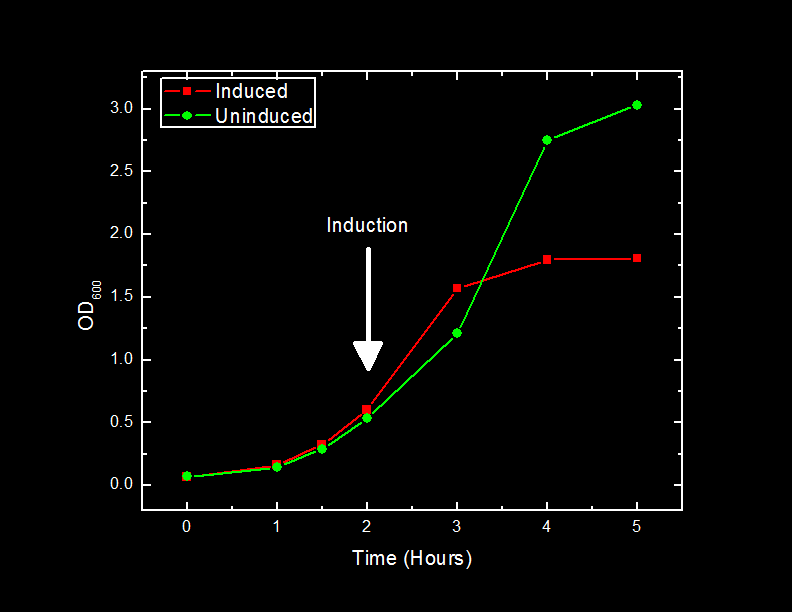
| + | xylE contains an iron molecule in its active site. Our hypothesis was that the iron molecule is oxidized after converting a single catechol molecule to 2-HMS; rendering the xylE inactive. We also want to test how xylE would behave in vitro rather than <i>in vivo</i>. | ||
| + | <br><br> | ||
| + | To test this hypothesis we grew <i>E. coli</i> containing BBa_K118021 in 500 mL of Lysogeny broth (LB) media with ampicillin. <i>E.coli<i> containing the pUC19 plasmid were also grown in 500 mL of Lysogeny broth (LB) media with ampicillin to act as a negative control through out the course experiment. The cells were grown to a final optical density of 4.55 AU (measured at 600 nm). The cells were first spun down at 3800 rcf for 5 minutes. The supernatant was decanted and the cells were re-suspended in 40 mL M9 minimal media and incubated with 10 mg of lysozyme for 10 minutes. This cell suspension was spun down at 10000 rcf for 30 minutes. The supernatant was taken off and spun at 30000 rcf (S30) for 1 hour. The supernatant of the S30 samples was divided. Half the samples were flash frozen in liquid nitrogen and stored at -80<sup>o</sup>C. The second half was spun at 100000 rcf for 45 minutes. | ||
| + | <br><br> | ||
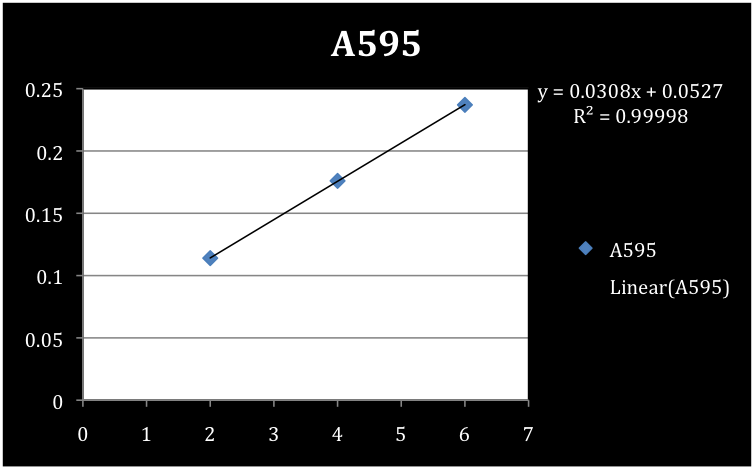
| + | In order to measure the 2-HMS production we needed to determine the concentration of protein per volume of sedimentation material. To determine the concentration of the S30 and S100 xylE sedimentation a Bradford assay was conducted using a standard curve of BSA. Concentrations of the negative controls (pUC 19) for each of the S30 and S100 samples were also determined. Absorbance readings were taken at a 595 nm wavelength and concentration reported in µg/mL. | ||
| + | <br><br> | ||
| + | |||
| + | ===<font color="white">Results of the Bradford Assay=== | ||
| + | [[image:UofLcatecholfigure3.jpg]] | ||
| + | <html> | ||
| + | <body> | ||
| + | <center> | ||
| + | <table border="0" cellpadding="8" width="28%" style="background-color:#000000"> | ||
| + | |||
| + | <tr> | ||
| + | <th> | ||
| + | <img src=""/> | ||
| + | </th> | ||
| + | </tr> | ||
| + | |||
| + | </tr> | ||
| + | <tr><th colspan="1" align="left"> | ||
| + | <p> | ||
| + | <font color="white">Figure 3. Standard curve of BSA. | ||
| + | </th> | ||
| + | </tr> | ||
| + | |||
| + | </table> | ||
| + | </center> | ||
| + | </body> | ||
| + | </html> | ||
| + | |||
| + | We determined the concentrations of S30, S30 negative control, S100, and S100 negative control to be 779 µg/mL, 104.5 µg/mL, 519.3 µg/mL, and 100.1 µg/mL respectively. | ||
| + | <br><br> | ||
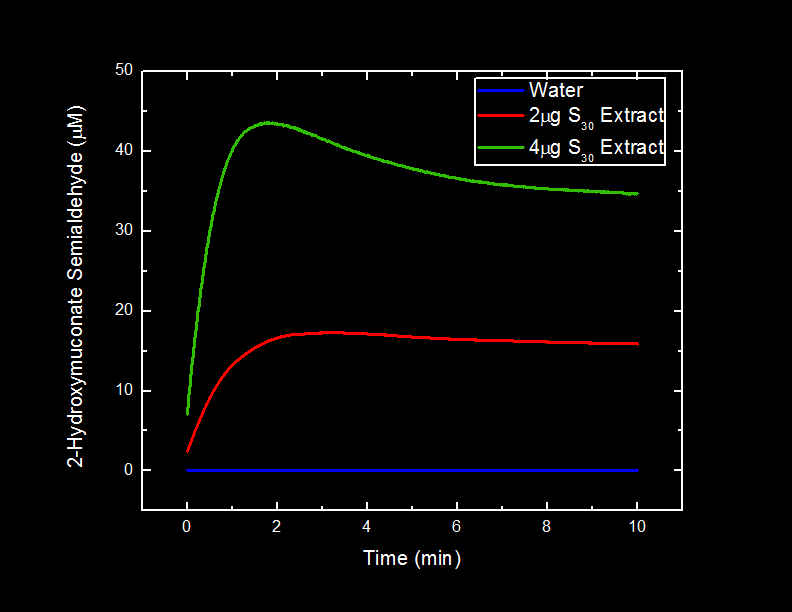
| + | The different volumes of S30 and S100 extracts of xylE cells and pUC19 cells were incubated in a 0.05 mM catechol solution and the production of 2-HMS was observed over time. This observation was measured by recording absorbance readings at 375 nm over a ten minute period immediately after the introduction of catechol. To measure the absorbance of 2-HMS the S30 and S100 samples had to be diluted. The samples were diluted with 20 mM Tris pH 8.0 and also with d<sub>2</sub>H<sub>2</sub>O. | ||
| + | <br><br> | ||
| + | [[image:UofLcatecholfigure3.jpg]] | ||
| + | <html> | ||
| + | <body> | ||
| + | <center> | ||
| + | <table border="0" cellpadding="8" width="28%" style="background-color:#000000"> | ||
| + | |||
| + | <tr> | ||
| + | <th> | ||
| + | <img src=""/> | ||
| + | </th> | ||
| + | </tr> | ||
| + | |||
| + | </tr> | ||
| + | <tr><th colspan="1" align="left"> | ||
| + | <p> | ||
| + | <font color="white">Figure 3. Standard curve of BSA. | ||
| + | </th> | ||
| + | </tr> | ||
| + | |||
| + | </table> | ||
| + | </center> | ||
| + | </body> | ||
| + | </html> | ||
| + | |||
| + | ===<font color="white">Results=== | ||
| + | |||
| + | We did not observe a difference in the catechol production in samples containing Tris vs. samples containing d<sub>2</sub>H<sub>2</sub>O. | ||
| + | |||
| + | [[image:UofLcatecholfigure4.jpg]] | ||
| + | <html> | ||
| + | <body> | ||
| + | <center> | ||
| + | <table border="0" cellpadding="8" width="28%" style="background-color:#000000"> | ||
| + | |||
| + | <tr> | ||
| + | <th> | ||
| + | <img src=""/> | ||
| + | </th> | ||
| + | </tr> | ||
| + | |||
| + | </tr> | ||
| + | <tr><th colspan="1" align="left"> | ||
| + | <p> | ||
| + | <font color="white">Figure 4. | ||
| + | </th> | ||
| + | </tr> | ||
| + | |||
| + | </table> | ||
| + | </center> | ||
| + | </body> | ||
| + | </html> | ||
=<font color="white">Compartmentalization Parts= | =<font color="white">Compartmentalization Parts= | ||
 "
"