Team:Lethbridge/Results
From 2010.igem.org
Adam.smith4 (Talk | contribs) (→Method) |
Liszabruder (Talk | contribs) |
||
| Line 178: | Line 178: | ||
<br> | <br> | ||
<br> | <br> | ||
| + | ==<font color="white">Characterization of Cyan Fluorescent Protein</font>== | ||
| + | ===<font color="white">Characterized Parts</font>=== | ||
| + | <hr> | ||
| + | <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331033" target="new"><font color="#00DC00" size="+1">BBa_K331033</font></a></html> | ||
| + | <br><br> | ||
| + | <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_E0020"target="new"><font color="#00DC00" size="+1">BBa_E0020</font></a></html> | ||
| + | <br><br> | ||
| + | <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331025"target="new"><font color="#00DC00" size="+1">BBa_K331025</font></a></html> | ||
| + | <br><br> | ||
| + | <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_”R0040"target="new"><font color="#00DC00" size="+1">BBa_R0040</font></a></html> | ||
| + | <br><br> | ||
| + | <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_B0034"target="new"><font color="#00DC00" size="+1">BBa_B0034</font></a></html> | ||
| + | <br><br> | ||
| + | <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_"target="new"><font color="#00DC00" size="+1">BBa_</font></a></html> | ||
| + | <br><br> | ||
| + | ===<font color="white">Hypothesis</font>=== | ||
| + | <hr> | ||
| + | Light will be observed at 476nm when cells containing <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331033" target="new"><font color="#00DC00">BBa_K331033</font></a></html> are illuminated with light at 439nm as a result of the excitation and subsequent fluorescence of Cyan Fluorescent Protein (CFP). | ||
| + | <br><br> | ||
| + | |||
| + | ===<font color="white">Introduction</font>=== | ||
| + | <hr> | ||
| + | Prior to introducing a oligoarginine tail onto the catechol 2,3 dioxygenase protein with the intention of directing the fusion protein into a lumazine synthase microcompartment, there must be some confirmation that oligoarginine tail is indeed causing the protein to which it is attached to localize within the microcompartment. We chose to utilize a technique known as fluorescent resonance energy transfer1 (FRET) to verify localization. In FRET, when a dye pair comes into close proximity to one another, one dye (the donor) will transfer some of its energy to the second dye (the acceptor). This will cause the acceptor to fluoresce without direct excitation. The dye pair we have chose is CFP as the donor and yellow fluorescent protein (YFP) as the acceptor. | ||
| + | <br><br> | ||
| + | Here we investigate the ability of <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331033" target="new"><font color="#00DC00">BBa_K331033</font></a></html> to produce CFP and emit light at its characteristic emission wavelength. | ||
| + | <br><br> | ||
| + | |||
| + | ===<font color="white">Method</font>=== | ||
| + | <hr> | ||
| + | In order to characterize the tetracycline repressible CFP (BioBrick <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331033" target="new"><font color="#00DC00">BBa_K331033</font></a></html>), cyan fluorescent protein (CFP-<html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_E0020"target="new"><font color="#00DC00"">BBa_E0020</font></a></html>) with an oligoarginine tag fused to the C-terminus (<html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331025"target="new"><font color="#00DC00">BBa_K331025</font></a></html>) (and preceded by a ribosomal binding site – <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_B0034"target="new"><font color="#00DC00">BBa_B0034</font></a></html>) was synthesized. We used our Red/White 3-Antibiotic assembly method to add a tetracycline repressible promoter (<html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_”R0040"target="new"><font color="#00DC00">BBa_R0040</font></a></html>) for constitutive expression of the fusion protein. This construction yielded BioBrick <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331033" target="new"><font color="#00DC00">BBa_K331033</font></a></html>. | ||
| + | <br><br> | ||
| + | The BioBrick containing plasmid was transformed into Escherichia coli DH5α cells. These cells were grown to an OD600 of approximately 0.7 in LB media, and diluted 1:10 with MilliQ H2O immediately prior to analysis by fluorescent spectroscopy. | ||
| + | <br><br> | ||
| + | This dilution of cells was excited at 439 nm, and the emission spectra was read from 444nm to 650nm. | ||
| + | <br><br> | ||
| + | |||
| + | ===<font color="white">Results</font>=== | ||
| + | <hr> | ||
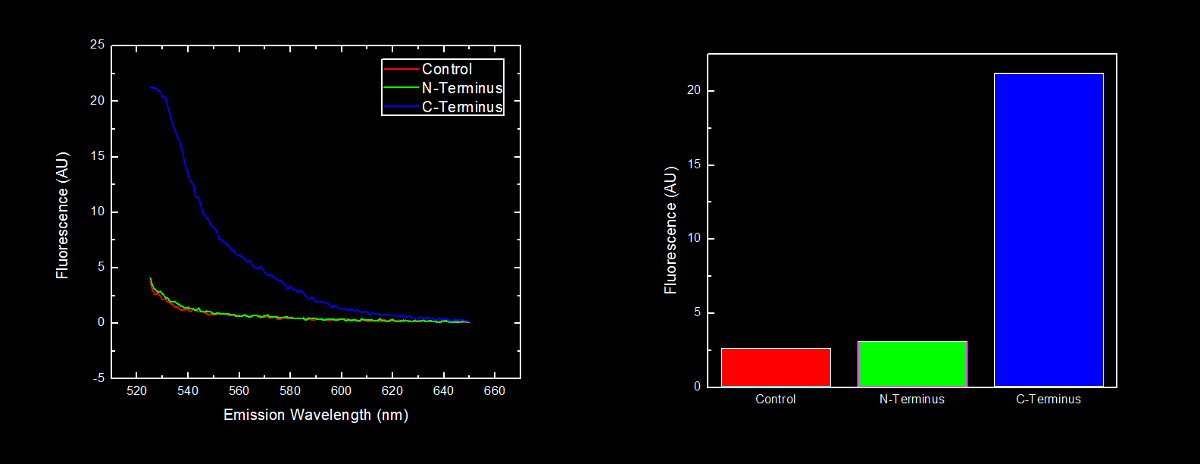
| + | Fluorescence at 476nm was observed. This peak, along with a shoulder occurring at approximately 510nm is consistent with results obtained by McRae1 et al in their rapid purification of ECFP. | ||
| + | [[image:UofL.cfpfigureblackjpg|900px]] | ||
| + | <br><br> | ||
| + | ===<font color="white">Conclusion</font>=== | ||
| + | <hr> | ||
| + | The BioBrick <html><a href="http://partsregistry.org/wiki/index.php/Part:BBa_K331033" target="new"><font color="#00DC00">BBa_K331033</font></a></html> that we constructed generates CFP in the absence of tetracycline, as expected. | ||
| + | <br><br> | ||
| + | |||
| + | ===<font color="white">Future Directions=== | ||
| + | We will assembly this into a single biobrick which can differentially express CFP/YFP and lumazine synthase for confirmation (through FRET) that the oligoarginine tail can be utilized to localize proteins within a modified lumazine synthase microcompartment. | ||
| + | <br><br> | ||
| + | |||
| + | ===<font color="white">Reference</font>=== | ||
| + | <hr> | ||
| + | <sup>1</sup>Förster T. (1948), Zwischenmolekulare Energiewanderung und Fluoreszenz. <i>Annalen der Physik</i>, 437: 55-75. | ||
| + | <br><br> | ||
| + | <sup>2</sup>McRae S.R., Brown C.L., Bushell G.R. (2005), Rapid Purification of EGFP, EYFP, and ECFP with high yield and purity. <i>Protein Expression and Purification</i> 41. 1 121-127. | ||
| + | <br><br> | ||
| + | =<font color="white">Magnetic Nanoparticles Parts= | ||
| + | One of the sub-projects for the bioremediation of the tailings ponds is to reduce heavy metals to create magnetic nanoparticles that can then be removed from the pond. To do this we need to be able to produce Mms6 (the iron reducing protein) and show that it can successfully produce the nanoparticles within the <i>Escherichia coli</i> cell. He is the work we have accomplished so far to characterize Mms6. | ||
| + | <br><br> | ||
| + | ==<font color="white">Characterization of Mms6== | ||
| + | ===<font color=“white”>Hypothesis=== | ||
| + | <hr> | ||
| + | Induction of Mms6 protein overexpression in <i>Escherichia coli</i> DH5α cells will cause a change in growth. | ||
| + | <br><br> | ||
| + | ===<font color="white">Introduction=== | ||
| + | <hr> | ||
| + | A parallel project we are undertaking in <html><a href="https://2010.igem.org/Team:Lethbridge/Project"><font color="#00DC00">oilsands remediation</font></a></html> is in the clean-up of heavy metal contaminants utilizing the Mms6 protein expressed by <i>Magnetiospirilum magneticum</i><sup>1</sup>. The Mms6 protein is capable of reducing aqueous iron into <html><a href="https://2010.igem.org/Team:Lethbridge/Project/Magnetic_Nanoparticles"><font color="#00DC00">magnetite nanoparticles</font></a></html>. The magnetite nanoparticles can be removed with much greater ease than aqueous iron. | ||
| + | <br><br> | ||
| + | We chose to investigate how the presence of the Mms6 protein affected <i>Escherichia coli</i> DH5α, its bacterial host cell. | ||
| + | <br><br> | ||
| + | ===<font color="white">Method=== | ||
| + | <hr> | ||
| + | In order to characterize the effect of Mms6 production on <i>Escherichia coli</i> DH5α cells, we utilized BioBrick <html><a href=" http://partsregistry.org/Part:BBa_K249019" target="new"><font color="#00DC00">BBa_K249019</font></a></html>, a part submitted by the <html><a href="http://partsregistry.org/Part:" target="new"><font color="#00DC00">2009 Lethbridge iGEM</font></a></html> team. <html><a href="http://partsregistry.org/Part: BBa_K249019" target="new"><font color="#00DC00">BBa_K249019</font></a></html> is the Mms6 coding region (<html><a href="http://partsregistry.org/Part: BBa_K249016" target="new"><font color="#00DC00">BBa_K249016</font></a></html>) with a ribosomal binding side (<html><a href="http://partsregistry.org/Part: BBa_B0030" target="new"><font color="#00DC00">BBa_B0030</font></a></html>) and a double terminator (<html><a href="http://partsregistry.org/Part: BBa_B0015" target="new"><font color="#00DC00">BBa_B0015</font></a></html>) under the control of a lactose inducible promoter (BBa_R0010</font></a></html>). | ||
| + | <br><br> | ||
| + | We inoculated two 500 mL cultures of LB media with <i>Escherichia coli</i> DH5α cells containing <html><a href="http://partsregistry.org/Part: BBa_K249019" target="new"><font color="#00DC00">BBa_K249019</font></a></html> and grew the cells until they reached approximately 0.6 OD<sub>600</sub> units. At this point, expression of the Mms6 protein was induced in one of the 500 mL cultures by the addition of 1 mM Isopropyl β-D-1-thiogalactopyranoside (IPTG). One milliliter of cells from each sample was removed at one hour intervals and the optical density at 600 nm was recorded and plotted. | ||
| + | <br><br> | ||
| + | ===<font color="white">Results=== | ||
| + | <hr> | ||
| + | The addition of IPTG to cells containing the <html><a href="http://partsregistry.org/Part: BBa_K249019" target="new"><font color="#00DC00">BBa_K249019</font></a></html> BioBrick caused a slowdown of cell growth after 2 hours, and a subsequent plateau at approximately one-half of the cells density that uninduced cells reached after three hours post-induction. | ||
| + | <br><br> | ||
| + | ===<font color="white">Conclusion=== | ||
| + | <hr> | ||
| + | The slowdown in cell growth as a result of production of Mms6 protein could be attributed to a number of factors. | ||
| + | <br><br> | ||
| + | One potential explanation could be a significant diversion of cellular resources to the production of Mms6 protein. This is unlikely, as subsequent analysis of cell fractions with SDS-PAGE showed no increase in banding at the expected molecular weight. | ||
| + | <br><br> | ||
| + | Another explanation could be that the Mms6 protein is toxic to the cells, and causes them to die. This possibility is being explored in collaboration with the <html><a href=" https://2010.igem.org/Team:Calgary" target="new"><font color="#00DC00">Calgary team</font></a></html> by using their troubleshooting kit. | ||
| + | <br><br> | ||
| + | ===<font color="white">References=== | ||
| + | <hr> | ||
| + | <sup>1</sup>Arakaki A., Masuda F., Amemiya Y., Tanaka T., Matsunaga T. (2010). Control of the morphology and size of magnetite particles with peptides mimicking the Mms6 protein from magnetotactic bacteria. <i>Journal of Colloid and Interface Science</i>. 343:65-60. | ||
| + | <br><br> | ||
 "
"