Team:Imperial College London/Modelling/Output/Detailed Description
From 2010.igem.org
| Line 40: | Line 40: | ||
[[Image:IC_1-step_amplification.JPG|300px|thumb|center|alt=A|1-step amplification]] | [[Image:IC_1-step_amplification.JPG|300px|thumb|center|alt=A|1-step amplification]] | ||
| - | |||
This model consists of the basic enzymatic reaction: | This model consists of the basic enzymatic reaction: | ||
Revision as of 19:33, 17 October 2010
| Temporary sub-menu: Objectives; Detailed Description; Parameters & Constants; Results & Conclusion;Download MatLab Files; |
| Output Amplification Model |
Detailed Description1. Model based on Law of Mass ActionDuring a meeting with our advisors, it was noted that our initial models (which had assumed that our system obeyed Michaelis Menten kinetics) were wrong. This is due to the fact that Michaelis Menten kinetics does not apply to our system. Click here to learn more about our models based on Michaelis Menten kinetics. In order to be able to use Michaelis Menten kinetics, there a lot of assumptions that have to be made. A few of these assumptions were not met by our system:
Since we are continuously producing enzyme, Vmax will change. Therefore the conservation E0 = E + ES does not hold for our system.
We are producing both substrate and enzyme, so we have approximately the same amount of substrate and enzyme.
Therefore, the model above is not representative of the enzymatic reaction. As we cannot use the Michaelis-Menten model we will have to solve from first principle (which involves writing down all of the biochemical equations and solving for these in Matlab). Change of output During our literature research, we came across a better output, so we abandoned the idea of using GFP as an output. Instead, we are using catechol. An enzyme, dioxygenase, will be acting on the catechol, which will then result in a coloured output. Catechol will be added to the bacteria manually (i.e. the bacteria will not produce catechol). Hence, in our models dioxygenase will be treated as an output as this enzyme is the only activator of catechol in our system. This means that the change of catechol into its colourful form is dependent on the dioxygenase concentration.
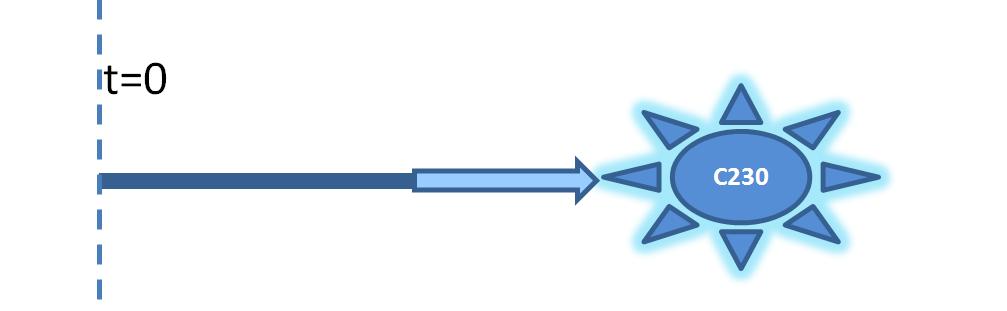
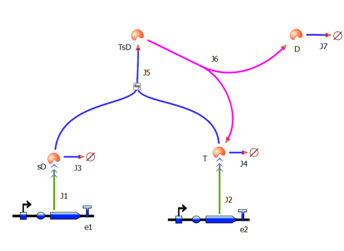
2. Model preA: Simple production of dioxygenaseThis model includes transcription and translation of the dioxygenase. It does not involve any amplification steps. It is our control model against which we will be comparing the results of other models. 3. Model A: Activation of Dioxygenase by TEV enzymeThis model consists of the basic enzymatic reaction: This is a simple enzymatic reaction, where TEV is the enzyme, Dioxygenase the product and split Dioxygenase the substrate. Choosing k1, k2, k3 as reaction constants, the reaction can be rewritten in these four sub-equations: These four equations were implemented in Matlab, using a built-in function (ode45) which solves ordinary differential equations. Implementation in TinkerCell Another approach to model the amplification module would be to implement it in a program such as TinkerCell (or CellDesigner). This would be useful to check whether the Matlab model works.
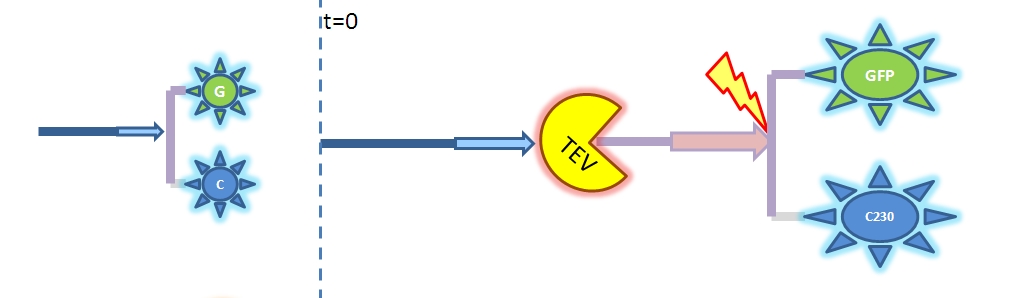
4. Model B: Activation of Dioxygenase by TEV or activated split TEV enzymeThis version includes the following features:
5. Model C: Further improvementsThis model has not been implemented because of the conclusions that we reached from Models A and B. It would include the following features:
|
 "
"