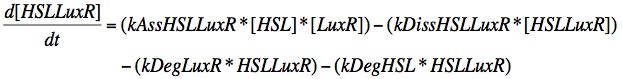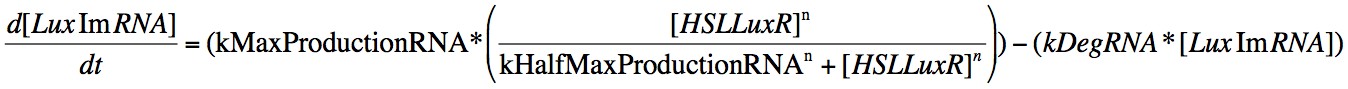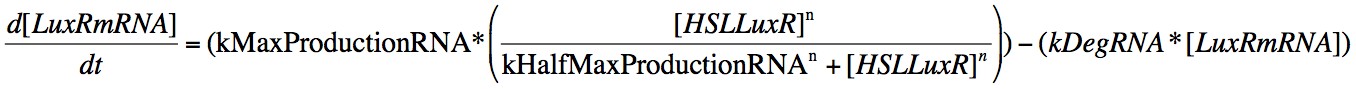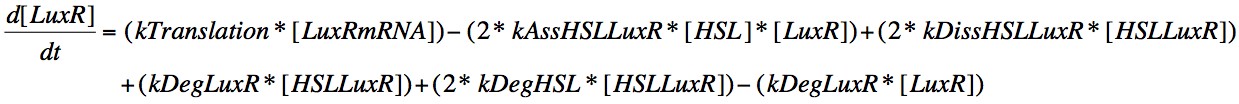Team:St Andrews/project/modelling/models
Modelling Methodology
Differential Equations
In order to capture this behaviour mathematically, we developed a system of 8 differential equations which together encapsulate the main behaviour of the LuxR circuit. Each equation represents the change in concentration of either a molecule or piece of mRNA with time, and all are first order.
Eqn. 1: The HSL-LuxR complex is broken apart when either HSL or LuxR degrades, or it may naturally dissociate at a rate kDissHSLLuxR. It is prodcued by the association of HSL and LuxR at a rate kAssHSLLuxR.
Eqn. 3: The production of LuxRmRNA is promoted by the HSL-LuxR complex. We model this behaviour using a Hill function controlled by the parameters kMaxProductionRNA and kHalfMaxProductionRNA.
Eqn. 5 : LuxI is translated from the mRNA at a reate kTranslation and degrades at a rate kDegRNA
Eqn. 6: LuxR is translated from its mRNA at a constant rate, kTranslation. It can also increase due to the dissociation of the HSL-LuxR complex or the degradation of the HSL or LuxR within the complex, which will result in those molecules which didn't degrade being included in the concentration present in the cell.
Eqn. 7: The GFP concentration is our primary indicator of activation of the quorum circuit. Similarly to LuxI it is produced from its respective mRNA at the rate kTranslation and degrades at a rate kDegGFP.
The above equations form the backbone of our model and are the framework on which all of our further work is based.
Our next issue was then to develop a method of simulating a change from low to high cell density in order to see if our model displayed the bistability that was theoreticaly predicted.
The Models
Over the course of the project, our model has evolved through various stages of operability and complexity. These can be split into four key stages of development, the culmination of which being our finalised model of the LuxR quorum sensing network. Each model has its own set of assumptions and theoretical foundation which was either reaffirmed or dispelled as our understanding of the biology behind the system became more complete. However we present our methodology and our results for all of our models in order to allow the interested party to better understand the reasoning behind our work and follow the logic that brought us to our final conclusions.
The four models are labelled:
Model 2: Single Cell With Loop
Model 3: Psuedo Multi Cell One Dimension
[[Team:St_Andrews/project/modelling/model_4 |Model 4: Psuedo Multi Cell Two Dimensions]
 "
"