Team:Northwestern/Project/Modeling
From 2010.igem.org
| Home | Brainstorm | Team | Acknowledgements | Project | Human Practices | Parts | Notebook | Calendar | Protocol | Safety | Links | References | Media | Contact |
|---|
|
| |||||||||||
ObjectiveThe main purpose of modeling was to characterize the topography of the kinetics behind foreign protein/product production in recombinant E.Coli biofilm with a diffusing inducer, and the effect on the various species involved by primarily the following factors:
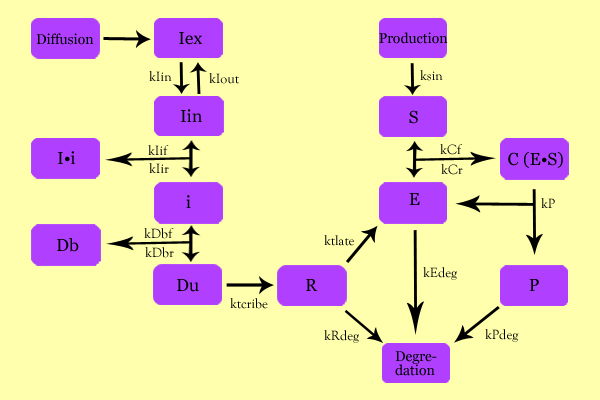
ModelingOverall ModelUsing enzyme kinetics equations, we elected to mathematically simulate the following model: VariablesIex: External Inducer, determined by diffusion through Fick's law (IPTG in our experiment) Iin: Internal Inducer (IPTG) Ii: Inducer bound to Repressor (IPTG bound to lacI) i: Repressor (lacI) Db: Repressor-bound DNA (lacI-bound DNA(CHS3) region in plasmid) Dunb: transcribe-able or Repressor-unbound DNA (lacI-unbound DNA(CHS3)) Re: mRNA for Enzyme (CHS3 mRNA) E: Enzyme (CHS3) S: Substrate (N-Acetyl Glucosamine) C: Enzyme Substrate Complex (CHS3-(N-Acetyl-Glucosamine)-Chitin or (NAG)n Complex) P: Protein Product (Chitin or (NAG)n+1) EquationsThe differential of the variables were found as follows:
IPTG DiffusionFirst, Fick's Law of Diffusion was modeled through MATLAB. The diffusion constant used was 220um^2/s.[4] IPTG is sprayed at the top of the colony, which then diffuses as according to Fick's law. The spatial local concentration will then differentially induce downstream processes. Semi-Empirical DeterminationFitted ModelMatlab was used to generate a theoretical model where IPTG would diffuse down the biofilm as according to Fick's Law of Diffusion and initiate the process. The extracellular substrate concentration was assumed to be much greater than the uptake/use, and so would diffuse in at a constant rate. This model was fitted with empirical data using cp-lacpi-gfp to estimate the rate constant, and therefore the effects of varying cp, lacpi, and rbs on enzyme and final product production. References1. A novel structured kinetic modeling approach for the analysis of plasmid instability in recombinant bacterial cultures William E. Bentley, Dhinakar S. Kompala Article first published online: 18 FEB 2004 DOI: 10.1002/bit.260330108 http://onlinelibrary.wiley.com/doi/10.1002/bit.260330108/pdf
Jongdae Lee, W. Fred Ramirez Article first published online: 19 FEB 2004 DOI: 10.1002/bit.260390608 http://onlinelibrary.wiley.com/doi/10.1002/bit.260390608/pdf
DOMINIQUE MENGIN-LECREULX, BERNARD FLOURET, AND JEAN VAN HEIJENOORT* E.R. 245 du C.N.R.S., Institut de Biochimie, Universit' Paris-Sud, Orsay, 91405, France Received 9 February 1983/Accepted 15 March 1983 http://www.ncbi.nlm.nih.gov/pmc/articles/PMC217602/pdf/jbacter00247-0262.pdf
Philip S. Stewart Center for Biofilm Engineering and Department of Chemical Engineering, Montana State University–Bozeman, Bozeman, Montana, 59717-3980 http://www.ncbi.nlm.nih.gov/pmc/articles/PMC148055/pdf/0965.pdf
PATRICIA L. EDELMANN' AND GORDON EDLIN Department of Genetics, University of California, Davis, California 95616 Received for publication 21 March 1974 http://www.ncbi.nlm.nih.gov/pmc/articles/PMC245824/pdf/jbacter00335-0105.pdf
MATLAB file provided upon request. | |||||||||||
 "
"