Team:Imperial College London/Modules/Signaling
From 2010.igem.org
| Detection Module | Signaling Module | Fast Response Module |
| We decided to design a new mechanism for parasite detection - by using the proteases they release. A novel protein bound to the cell surface, with a signaling peptide attached via a protease cleavage site. When the protease comes along, the signal peptide is released, allowing it to activate our signaling module. | To transduce the signal we used a quorum sensing system of a gram positive bacterium. The two component signal transduction system taken from S. pneumoniae transfers our peptide signal into the cell, activating the fast response module. | Our fast response mechanism is based around using two enzymatic amplification steps involving a transcripted enzyme, a deactivated enzyme and a presynthesised substrate. This greatly reduces the time required for producing a recognisable output, enabling useful field testing kits. |
| Signalling Module |
|
Using quorum sensing systems to detect proteases Having decided that we wanted to detect the parasite protease, we realised that a possible method might be to have it cleave a protein, which would then result in a downstream signalling pathway. A well-known signalling system using peptides is that of quorum sensing in gram positive bacteria. The signals, or quorum sensing molecules, are known as autoinducing peptides (AIPs) and are transported out of a cell on activation of certain pathways.
AIPs can remain as linear peptides, or they can be post-translationally modified to become circular. We needed to find a system which signals via a linear AIP, and one such example is the ComCDE system in Streptococcus pneumoniae which signals via a two component signal transduction system. ComC codes for the competent-stimulating peptide-1 (CSP-1) which is transported out of cells and subsequently detected by a the sensory histidine kinase ComD. ComD then autophosphorylates and then activates the response regulator ComE by phosphorylating it. ComE then binds to two 9bp imperfect direct repeats to induce transcription of specific target genes.
In order for us to get a strong enough readout, it was really important to establish the concentration of CSP-1 needed to activate the system. From reading 'An unmodified heptadecapeptide pheromone induces competence for genetic transformation in Streptococcus pneumoniae' by Havarstein, Coomaraswamy & Morrison, we found out that in previous studies only 10ng/ul of CSP-1 was needed to activate the two component signal transduction system. They also comment on the stability of CSP-1 and outlined other parameters affecting response to CSP-1, such as pH. This information was used by the modelling team to calculate how strong our response would be, and whether there was any possibility of getting a false positive.
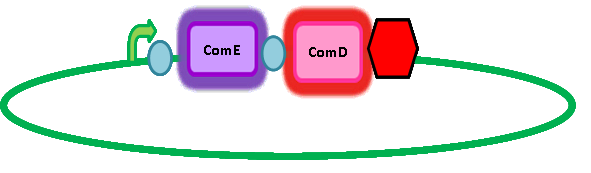
In order to get transcription of TEV protease upon activation of ComE, we had to add a ComE binding site upstream of the TEV protease gene. We read 'Two Separate Quorum-Sensing Systems Upregulate Transcription of the Same ABC Transporter in Streptococcus pneumoniae' by Knutsen, Ween & Håvarstein to determine the ComE binding site. They also assume that ComE binds DNA as a homodimer. The ComE binding sites are described in detail by Halfmann, Hakenbeck & Brückner. They took the ComA and ComC promoters, which each contained a ComE binding site, out of context and used it to initiate transcription of various target genes. However, they integrated this system into S. pneumoniae so we cannot be sure that it will work in B. subtilis. Similarly, there is a possibility that there is a similar transcriptional regulator inherent in B. subtilis which might result in false positives. However, having done homology searches in BLAST, there seems to be no homologs and so we have assumed that there is no risk of false positives arising from a homologous response regulator in B. subtilis. Here's a picture of the final construct: |
 "
"