Team:Imperial College London/Modelling/Protein Display
From 2010.igem.org
m |
(Having added menu and the paramters and constants) |
||
| Line 1: | Line 1: | ||
{{:Team:Imperial_College_London/Templates/Header}} | {{:Team:Imperial_College_London/Templates/Header}} | ||
| - | {| style="width:900px;background:#f5f5f5;text-align: | + | {| style="width:900px;background:#f5f5f5;text-align:justify;font-family: helvetica, arial, sans-serif;color:#555555;margin-top:25px;" cellspacing="20" |
| - | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"| | + | |style="font-family: helvetica, arial, sans-serif;font-size:2em;color:#ea8828;"|Output Amplification Model |
|- | |- | ||
|<html> | |<html> | ||
| + | <head> | ||
| + | <script type="text/javascript" src="http://ajax.googleapis.com/ajax/libs/jquery/1.4.2/jquery.min.js"></"></script> | ||
| + | <script type="text/javascript"> | ||
| + | $(document).ready(function() { | ||
| + | |||
| + | $('div.accordionButton').click(function() { | ||
| + | |||
| + | $(this).next().slideToggle('normal'); | ||
| + | }); | ||
| + | |||
| + | $("div.accordionContent").hide(); | ||
| + | |||
| + | }); | ||
| + | |||
| + | </script> | ||
| + | <style type="text/css"> | ||
| + | #wrapper { | ||
| + | width: 800px; | ||
| + | margin-left: auto; | ||
| + | margin-right: auto; | ||
| + | } | ||
| + | |||
| + | .accordionButton { | ||
| + | width: 400px; | ||
| + | margin-left: 175px; | ||
| + | height: 20px; | ||
| + | float: left; | ||
| + | text-align: center; | ||
| + | display: block; | ||
| + | color: #ffffff; | ||
| + | background: #Ea8828; | ||
| + | cursor: pointer; | ||
| + | padding: 10px; | ||
| + | } | ||
| + | |||
| + | .accordionContent { | ||
| + | width: 775px; | ||
| + | float: left; | ||
| + | text-align: justify; | ||
| + | background: #f5f5f5; | ||
| + | display: none; | ||
| + | padding:10px; | ||
| + | } | ||
| + | |||
| + | .accordionButton:hover { | ||
| + | color: #555555; | ||
| + | border-bottom: 1px solid #555555; | ||
| + | } | ||
| + | |||
| + | </style> | ||
| + | </head> | ||
| + | <body style="background-color:FFFFCC"> | ||
| + | <div id="wrapper"> | ||
| + | <div class="accordionButton">Details</div> | ||
| + | <div class="accordionContent"> | ||
| + | |||
The aim of this model is to determine the concentration of Schistosoma elastase or TEV protease that should be added to the bacteria to trigger the response. It is also attempted to determine how long it will take for the protease or elastase to cleave enough peptides. | The aim of this model is to determine the concentration of Schistosoma elastase or TEV protease that should be added to the bacteria to trigger the response. It is also attempted to determine how long it will take for the protease or elastase to cleave enough peptides. | ||
<br /><br /> | <br /><br /> | ||
| Line 217: | Line 273: | ||
<li>Knutsen, E., Ween, O. & Havarstein, L. (2003) Two Separate Quorum-Sensing Systems Upregulate Transcription of the Same ABC Transporter in Streptococcus pneumoniae. Journal of Bacteriology. [Online] 186(10), 3078-3085. Available from: http://jb.asm.org/cgi/reprint/186/10/3078 [Accessed 1st September 2010]</li> | <li>Knutsen, E., Ween, O. & Havarstein, L. (2003) Two Separate Quorum-Sensing Systems Upregulate Transcription of the Same ABC Transporter in Streptococcus pneumoniae. Journal of Bacteriology. [Online] 186(10), 3078-3085. Available from: http://jb.asm.org/cgi/reprint/186/10/3078 [Accessed 1st September 2010]</li> | ||
</ol> | </ol> | ||
| + | |||
| + | </div> | ||
| + | |||
| + | <div class="accordionButton">Parameters & Constants</div> | ||
| + | <div class="accordionContent" style="font-family: helvetica, arial, sans-serif;color:#555555;"> | ||
| + | <html> | ||
| + | <h1>Constants for the Protein Display Model</h1> | ||
| + | </html> | ||
| + | |||
| + | <html> | ||
| + | <table width="775px" border="0"> | ||
| + | |||
| + | <tr> | ||
| + | <td style="background-color:#FFFF66;height:50px;width:75;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;"><b>Type of Constant</b> | ||
| + | </td> | ||
| + | <td style="background-color:#FFFF99;height:50px;width:700;text-align:center"><b>Derivation of Value</b> | ||
| + | </td> | ||
| + | </tr> | ||
| + | |||
| + | <tr> | ||
| + | <td style="background-color:#FFCC66;height:100px;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>TEV Enzyme Dynamics</b> | ||
| + | </td> | ||
| + | <td style="background-color:#e7e7e7;height:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;">Enzymatic Reaction: E+S <var>↔</var> ES <var>→</var> E+P | ||
| + | <br /> | ||
| + | The derivation of these values is made in <a href="http://www.openwetware.org/wiki/IGEM:Imperial/2010/Variables1">Variables for Amplification Module Section</a>. | ||
| + | <ul> | ||
| + | <li>k<sub>1</sub> = rate constant for E + S <var>→</var> ES = 10<sup>8</sup> M<sup>-1</sup>s<sup>-1</sup> | ||
| + | <li>k<sub>2</sub> = rate constant for E + S <var>←</var> ES = 10<sup>3</sup> s<sup>-1</sup> | ||
| + | <li>k<sub>cat</sub> = rate constant for ES <var>→</var> E + P = 0.16 <var>±</var> 0.01 s<sup>-1</sup> | ||
| + | </ul> | ||
| + | <br /> | ||
| + | We are assuming the same cleaving rates of TEV as on other substrates. However, we are planning to measure them to gain more confidence in the model. | ||
| + | |||
| + | </td> | ||
| + | </tr> | ||
| + | |||
| + | <tr> | ||
| + | |||
| + | <td style="background-color:#FFCC66;height:100px;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Production Rate of Surface Proteins</b> | ||
| + | </td> | ||
| + | <td style="background-color:#e7e7e7;height:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;">It was found that each cell displays 2.4x10<sup>5</sup> peptides <a href="http://onlinelibrary.wiley.com/doi/10.1111/j.1574-6968.2000.tb09188.x/pdf">[1]</a>. | ||
| + | Hence, we adjusted our simple production of display protein model to converge to that value. As production rate was the constant that we could not obtain, that value was manipulated. | ||
| + | <br /> | ||
| + | The result 4.13x10<sup>-8</sup>mol/dm<sup>3</sup>/s seemed to be of reasonable order of magnitude. | ||
| + | Ideally, we would like to get this value measured as it is resulting from a very vague estimate. | ||
| + | </td> | ||
| + | </tr> | ||
| + | |||
| + | <tr> | ||
| + | <td style="background-color:#FFCC66;height:100px;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Degradation Rate of Surface Proteins (common for all)</b> | ||
| + | </td> | ||
| + | <td style="background-color:#e7e7e7;height:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;">Assumption: To be approximated by cell division (dilution of media) as none of the proteins are involved in any active degradation pathways. | ||
| + | <br /> | ||
| + | Derived in <a href="http://www.openwetware.org/wiki/IGEM:Imperial/2010/Variables1">Variables for Amplification Module Section</a>: | ||
| + | <br /> | ||
| + | k<sub>deg</sub>= 0.000289s<sup>-1</sup> | ||
| + | <br /> | ||
| + | For all proteins that are outside of cells or the timescale that is short enough to neglect cell division effect: k<sub>deg</sub>=0 | ||
| + | |||
| + | </td> | ||
| + | </tr> | ||
| + | |||
| + | <tr> | ||
| + | <td style="background-color:#FFCC66;height:100px;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Diffusion Coefficient of Proteins</b> | ||
| + | </td> | ||
| + | <td style="background-color:#e7e7e7;height:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;">We have found two references which quote very similar values for very different media.<br />For protein in agarose gel: D<sub>average</sub> = 1.07x10<sup>-10</sup>m<sup>2</sup>/s - for a protein in agarose gel for pH=5.6 <a href="http://www.sciencedirect.com/science?_ob=MImg&_imagekey=B6V5N-4B3MXDC-2-K&_cdi=5791&_user=217827&_pii=S1369703X03002377&_origin=search&_coverDate=07%2F01%2F2004&_sk=999809998&view=c&wchp=dGLzVtb-zSkzS&md5=c17d0e7320f03931006f9b1a10a438b9&ie=/sdarticle.pdf">[2]</a> | ||
| + | <br /> | ||
| + | In the final model the following was used: | ||
| + | For protein in water: D=10<sup>-10</sup>m<sup>2</sup>/s <a href="http://www.life.illinois.edu/crofts/bioph354/diffusion1.html">[3]</a> | ||
| + | </td> | ||
| + | </tr> | ||
| + | |||
| + | <tr> | ||
| + | <td style="background-color:#FFCC66;height:100px;text-align:center; font-family: helvetica, arial, sans-serif;color:#555555;"><b>Control Volume</b> | ||
| + | </td> | ||
| + | <td style="background-color:#e7e7e7;height:100px;text-align:center;font-family: helvetica, arial, sans-serif;color:#555555;">The control volume seems to be the weakest point of this model. We have tried to rationalise it as much as we could. However, errors seem to be unavoidable. It is important to realise that the Control Volume needs to be adjusted if a bacterial concentration different than 5x10<sup>8</sup>CFU/ml is used. | ||
| + | </td> | ||
| + | </tr> | ||
| + | |||
| + | |||
| + | </table> | ||
| + | </html> | ||
| + | |||
| + | <html> | ||
| + | <h2>References</h2> | ||
| + | <ol> | ||
| + | <li>Kobayashi, G. et al (2000) Accumulation of an artificial cell wall-binding lipase by Bacillus subtilis wprA and/or sigD mutants. FEMS Microbiology Letters. [Online] 188(2000), 165-169. Available from: http://onlinelibrary.wiley.com/doi/10.1111/j.1574-6968.2000.tb09188.x/pdf [Accessed 27th August 2010]</li> | ||
| + | <li>Gutenwik, J., Nilsson, B. & Axelsson, A. (2003) Determination of protein diffusion coefficients in agarose gel with a diffusion cell. Biochemical Engineering Journal. [Online] 19(2004), 1-7. Available from: http://www.sciencedirect.com/science?_ob=MImg&_imagekey=B6V5N-4B3MXDC-2-K&_cdi=5791&_user=217827&_pii=S1369703X03002377&_origin=search&_coverDate=07%2F01%2F2004&_sk=999809998&view=c&wchp=dGLzVtb-zSkzS&md5=c17d0e7320f03931006f9b1a10a438b9&ie=/sdarticle.pdf [Accessed August 20th 2010]</li> | ||
| + | <li>Crofts, A. (1996) Biophysics 345. [Online] Available from: http://www.life.illinois.edu/crofts/bioph354/diffusion1.html [Accessed 1st September 2010]</li> | ||
| + | </ol> | ||
| + | |||
| + | |||
| + | </div> | ||
| + | |||
| + | |||
| + | <div class="accordionButton">Results & Conclusions</div> | ||
| + | <div class="accordionContent"> | ||
| + | </div> | ||
| + | |||
| + | </body> | ||
| + | <br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/><br/> | ||
</html> | </html> | ||
|} | |} | ||
Revision as of 10:33, 13 October 2010
| Output Amplification Model | ||||||||||||||||||||||||||
|
Details
The aim of this model is to determine the concentration of Schistosoma elastase or TEV protease that should be added to the bacteria to trigger the response. It is also attempted to determine how long it will take for the protease or elastase to cleave enough peptides.
Cleavage of protein is an enzymatic reaction, which can be written as:
This can be modelled in a very similar way to the 1-step amplification model, however, all the constants and initial concentrations will be different. [TEV](t=0) - initial concentration of TEV will be arbitrarily chosen. However, ultimately we would need to measure the concentration of elastase that schistosoma releases. Threshold concentration of peptide (20/08/2010)The optimal peptide concentration required to activate ComD is 10 ng/ml [1]. This is the threshold value for ComD activation. However, the minimum concntration of peptide to give a detectable activation is 0.5ng/ml.We want to know how long it takes until the threshold is reached:
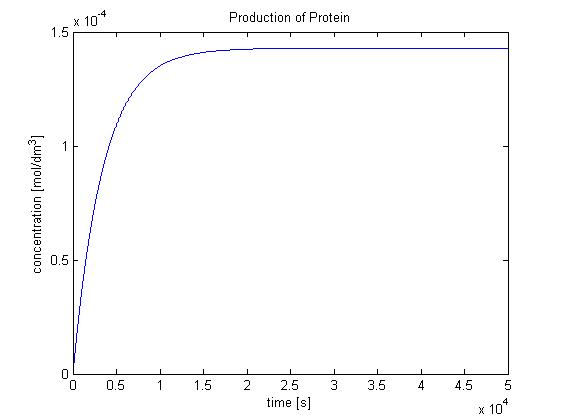
Protein production in B.sub (23/08/2010)
Hence, we can deduce that the concentration that the protein expression will tend to: c = 1.4289x10-4 mol/dm3 = cfinal. Therefore, we can model the protein production by transcription and translation and adjust the production constant so the concentration value will tend towards cfinal. Using a similar model to the simple production of Dioxygenase for the Output Amplification Model (Model preA), we obtain the following graph:
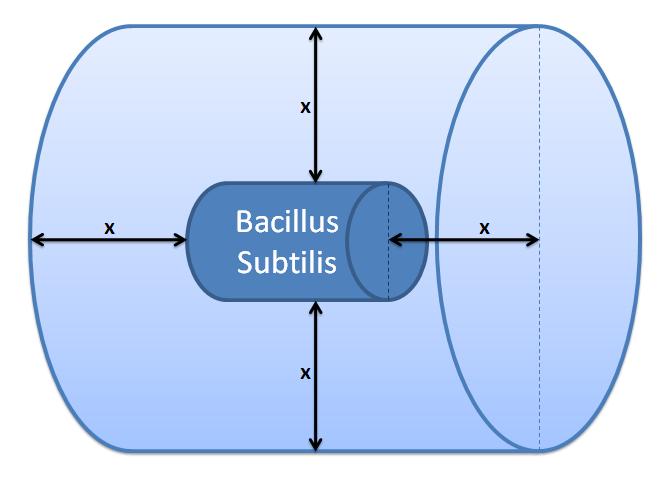
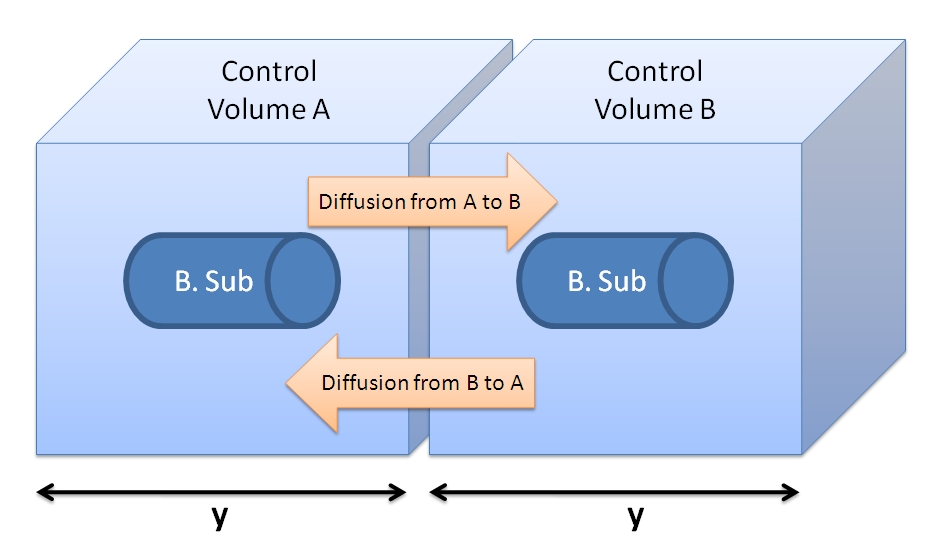
Control volume initial choice (23/08/2010)All enzymatic reactions that we have modelled so far were confined whithin the bacterial cell. However, this case is different because the molecules are not confined by the bacterial membrane and can diffuse out of the cell.The control volume: The inner boundary is determined by the bacterial cell (proteins after being displayed and cleaved cannot diffuse back into bacterium). The outer boundary is more time scale dependent. We have assumed that after mass cleavage of the display-proteins by TEV, many of these AIPs will bind to the receptors quite quickly (eg. 8 seconds). Our volume is determined by the distance that AIPs could travel outwards by diffusion within that short time. In this way, we are sure that the concentration of AIPs outside our control volume after a given time is approximately 0. This approach is not very accurate and can lead us to false negative conclusions (as in reality there will be a concentration gradient, with highest conentration on the cell wall).
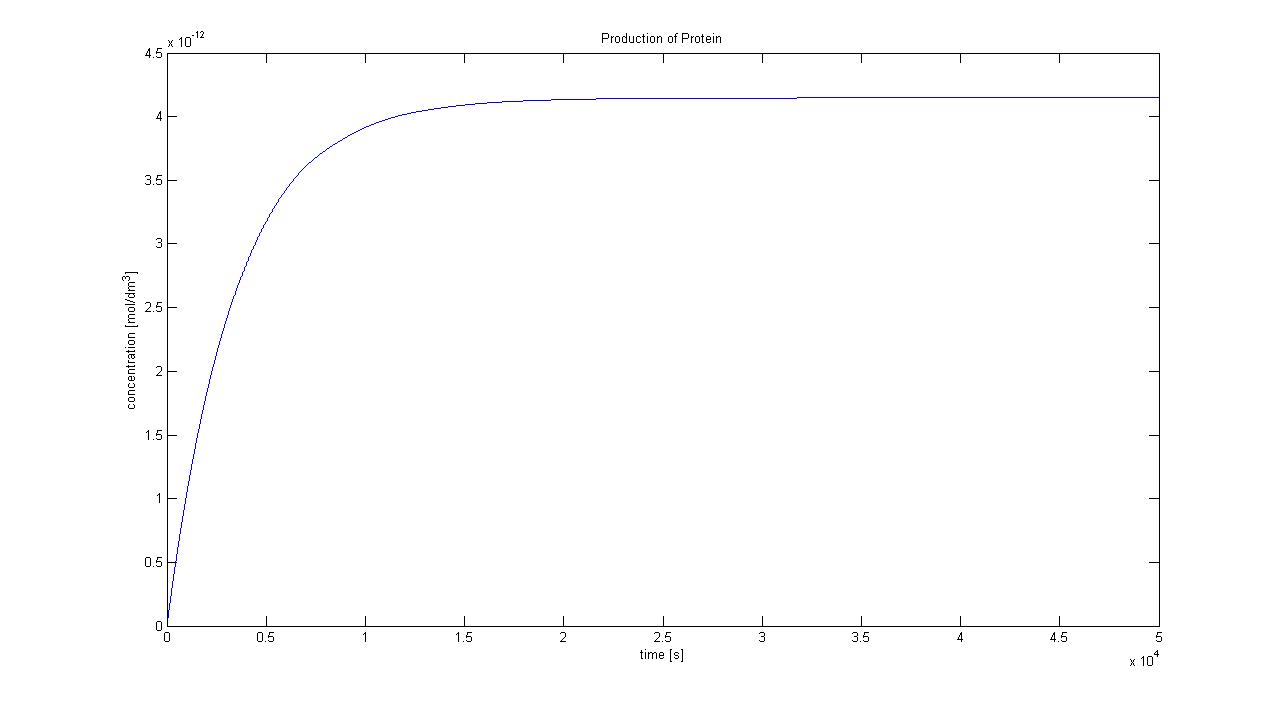
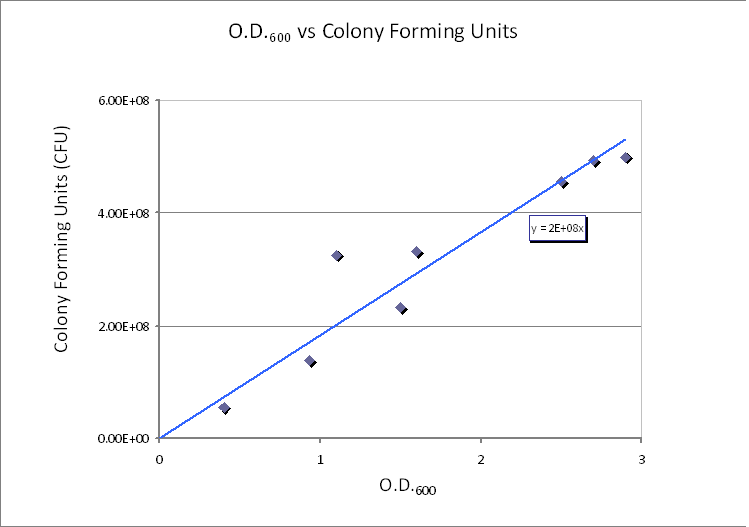
Protein production in Control Volume (23/08/2010)The previously determined constants of protein production in B.sub to obtain the concentration of proteins are not valid in the Control Volume. It has to be adjusted (multiplied) by the following factor:factor=Vbacillus/VCV = 5.7974x10-6 (for the particular numbers presented above) Control volume final choice (23/08/2010)We realized that our initial choice of control volume was not accurate because this assumption was treating bacteria as the medium. However, in reality bacteria live in colonies very close to each other. They are much closer to each other than the diffusion distance (1.9596x10−5m) derived above even if placed in water solution.Using CFU to estimate the spacing between cells (24/08/2010)CFU stands for Colony-forming unit. It is a measure of bacterial numbers. For liquids, CFU is measured per ml. Since we already have data of CFU/ml from the Imperial iGEM 2008, this is an easy way to estimate the number of cells in a given volume using a spectrometer at 600nm wavelength. The graph below is taken from the Imperial iGEM 2008 Wiki page [4].
Choice of Control Volume allows simplifications (24/08/2010)
Matlab Simulation (24/08/2010)Here is the Matlab code for the Matlab simulation
Sensitivity of our model (24/08/2010)
Whether the threshold concentration of AIP is reached is highly dependent on the initial concentration of TEV. The smallest initial concentration of TEV, [TEV>]
One order of magnitude change in the production rate results in at least 50s delay of the AIP concentration reaching the threshold concentration. Changing the production rate influences the time duration of the AIP concentration above the threshold level. The higher it is, the shorter the receptor will be activated (at extreme values, AIP concentration does not reach the threshold). However, the production rate has not much influence on how fast the threshold will be reached. Our model is extremely sensitive to this factor. One order of magnitude change in CV results in several orders of magnitude change in AIP concentration. Hence, special care should be taken in determination of this value. If the model is to be compared with the experimental results, the CFU/ml has to be the same as the one used in the model. Otherwise, the CV has to be readjusted. Risk of False positives (31/08/2010)It was pointed out that we should assess the risk of false positive activation of the receptor. We are particularly concerned about the display protein not binding to the cell wall, but instead diffusing into the extra-cellular environment. In order to be able to assess the risk of false positives, we need to do further research into the affinity of AIP with attached linker and transmembrane proteins for the receptor as compared to the affinity of the AIP itself for the receptor.This paper [5] might have some information on affinity comparison. We need to know how proteins are being transported from intracellular to transmembrane space. Understanding this concept could give us an idea of what could go wrong. References
Parameters & Constants
Constants for the Protein Display Model
References
Results & Conclusions
|
 "
"




![Graphs showing the simulation using [TEV]<sub>0</sub> = 4x10<sup>-4</sup> mol/dm<sup>3</sup>. The graph on the right hand-side below shows that the AIP threshold (red line) is reached after 22 s.](http://www.openwetware.org/images/7/77/Protein_display.png)
