Team:Heidelberg/Modeling
From 2010.igem.org
(→shRNA binding site features) |
(→shRNA binding site features) |
||
| Line 33: | Line 33: | ||
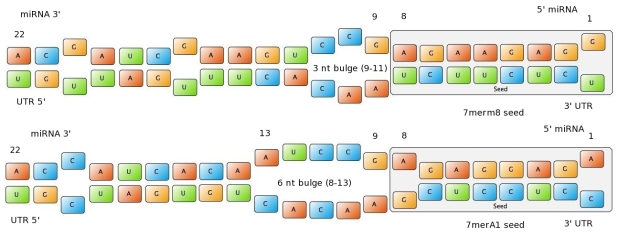
Figure 1: Interactions between two miRNAs and their binding sites. Examples to show different types of seeds. | Figure 1: Interactions between two miRNAs and their binding sites. Examples to show different types of seeds. | ||
| + | |||
| + | - 6mer (abundance 21.5%): only the nucleotides 2-7 of the miRNA match with the mRNA. | ||
| + | |||
| + | - 7merA1 (abundance 15.1%): the nucleotides 2-7 match with the mRNA, and there is an adenine in position 1. | ||
| + | |||
| + | - 7merm8 (abundance 25%): the nucleotides 2-8 match with the mRNA. | ||
| + | |||
| + | - 8mer (abundance 19.8%): the nucleotides 2-8 match with the mRNA and there is an adenine in position 1. | ||
| + | |||
| + | The percentages of abundance are calculated among conserved mammalian sites for a highly conserved miRNA (Friedman et al. 2008) | ||
=Tissue specific miRNAs= | =Tissue specific miRNAs= | ||
Revision as of 23:17, 26 October 2010

Being able to tune the expression levels of a gene in mammalian cells is a great concept but it becomes an extremely powerful tool if we can predefine the construct that can bring this about. This was the inspiration behind creating a tool that can provide us a scheme that, if followed, can make it possible to control the expression of a gene in specific cell types. miBEAT (full form) is a compilation of multiple individual models and scripts. It is shRNA binding site featuresAs the title of our project states, “DNA is not enough”. There are several upper-level regulation systems in superior organisms. Our main idea was using miRNA to tune down the expression of genes, having tissue-specific, exactly tuned gene therapy as objective. miRNA are non-coding regulatory RNAs functioning as post-transcriptional gene silencers. After they are processed, they are usually 22 nucleotides long and they usually bind to the 3’UTR region of the mRNA (although they can also bind to the ORF or to the 5'UTR), forcing the mRNA into degradation or just repressing translation. In vegetal organisms, miRNA usually bind to the mRNA with extensive complementarity. In animals, interactions are more inexact, creating a lot of uncertainty in the in silico prediction of targets. The seed of the miRNA is usually defined as the region centered in the nucleotides 2-7 in the 5’ end of the miRNA, and it usually requires extensive pairing. (For the sake of simplicity, we extended slightly the term seed to include the nucleotides 1-8.) Outside the seed, the existence of supplemental pairing (at least 3 contiguous nucleotides and centered in nucleotides 13-16 of the miRNA) stabilizes the bound complex and increases the efficacy of the binding site. Binding sites with a high local AU density around the binding site have proven to be more effective (possibly because of the destabilization of the mRNA secondary structure around the site). The position of the binding site not in the middle of the 3’UTR but either at the end or 15 nt after the stop codon increases the efficacy of repression. IS THE FOLLOWING REALLY NECESSARY? I DON'T THINK I CAN MANAGE TO EVER FIT THE TOPIC HERE... About the mechanistics of the repression, it has been shown that the repressive effect is much higher when the binding site for the miRNA is in the first 15 nt of the 3'UTR. This would match the hypothesis.... BLA BLA BLA Targetscan Scores Jan ???? Figure 1: Interactions between two miRNAs and their binding sites. Examples to show different types of seeds. - 6mer (abundance 21.5%): only the nucleotides 2-7 of the miRNA match with the mRNA. - 7merA1 (abundance 15.1%): the nucleotides 2-7 match with the mRNA, and there is an adenine in position 1. - 7merm8 (abundance 25%): the nucleotides 2-8 match with the mRNA. - 8mer (abundance 19.8%): the nucleotides 2-8 match with the mRNA and there is an adenine in position 1. The percentages of abundance are calculated among conserved mammalian sites for a highly conserved miRNA (Friedman et al. 2008) Tissue specific miRNAsA useful supplement to achieving tuned gene expression in cells is the ability to specifically target tissues where this should be carried out. Tissue targeting has, for quite some time, been an important field of research that has drawn much attention and is central to gene therapy. miBEAT[link] tool not only allows generation of binding sites that regulate the level of expression of a desired gene, but also employs strategies that help target the right tissue and exclude expression in others. This functionality is based on the principle of using tissue specific miRNA binding sites which can be introduced in the miTuner[link] construct easily. A smart way to specifically target tissues is to exploit the presence and absence of tissue specific endogenous miRNA in the target to specifically express or exclude expression of the gene of interest in the target. We make use of two of strategies based on this principle, namely, on-targeting and off-targeting. The off-targeting concept has been applied previously [http://2010.igem.org/Team:Heidelberg/Modeling#References (Wenfang Shi et.al., 2008)] wherein an endogenous miRNA is selected such that it is not present in the target tissue (therefore the gene is expressed) and is present in all the off target tissues (knockdown of the transcripts). Thus the gene is specifically expressed in the target. In addition to the off-targeting strategy, we designed a new strategy, the on-targeting. In this case, the miRNA is present in the target tissue and excluded from the off-targets. The binding site for this miRNA is present within the 3'UTR of a repressor gene (in our case TET/O2) construct. The operator for the repressor in turn precedes the gene of interest (miGene) in the miTuner or miMeasure constructs. Therefore in the presence of miRNA in the cell, repressor is degraded and miGene is expressed while in off-targets repressor is translated and represses the expression of miGene. References
|
 "
"