Team:Heidelberg/Modeling
From 2010.igem.org
(Difference between revisions)
(→miRockdown on miBEAT) |
(→miRockdown on miBEAT) |
||
| Line 54: | Line 54: | ||
===miRockdown on miBEAT=== | ===miRockdown on miBEAT=== | ||
Right from the beginning of our modeling project, we knew we would have to integrate our trained models into an online GUI. We realized it in the most user friendly way we could think of: The user only needs to input the desired knockdown percentage (kd%) and choose an sh/miRNA sequence, to get a binding site that satisfies the users needs. | Right from the beginning of our modeling project, we knew we would have to integrate our trained models into an online GUI. We realized it in the most user friendly way we could think of: The user only needs to input the desired knockdown percentage (kd%) and choose an sh/miRNA sequence, to get a binding site that satisfies the users needs. | ||
| + | <br> | ||
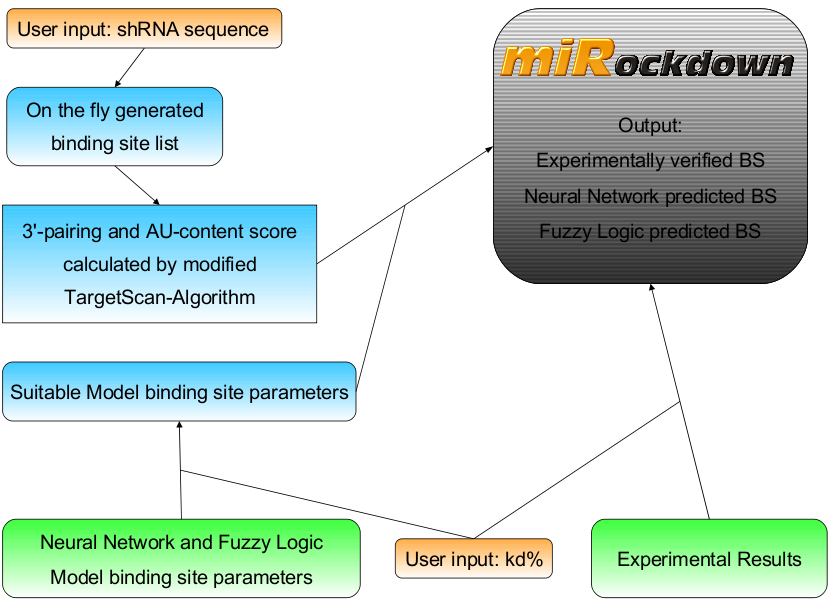
[[Image:MiRockdownOverview.png|630px]] | [[Image:MiRockdownOverview.png|630px]] | ||
| - | The results of both of our models and the experimentally verified binding sites are integrated in [miRockdown] (see Figure: miRockdown) on the [miBEAT] GUI. For every binding site request of a user there are the results of the three different concepts displayed. Thus the users can always choose which of the three differently generated binding to use. The binding site with the most similar experimentally observed knockdown percentage is given out, together with its properties and oligos ready to clone into the [miTuner]-construct. | + | <br> |
| - | The binding sites generated from the model results come into play, when the user wants to use his or her own sh/miRNA, or when the experimentally verified binding sites have a knockdown, that is not sufficiently similar to the desired knockdown. | + | The results of both of our models and the experimentally verified binding sites are integrated in [miRockdown] (see Figure: miRockdown) on the [miBEAT] GUI. For every binding site request of a user there are the results of the three different concepts displayed. Thus the users can always choose which of the three differently generated binding to use. The binding site with the most similar experimentally observed knockdown percentage is given out, together with its properties and oligos ready to clone into the [miTuner]-construct.<br> |
| - | A script integrated into miRockdown will correlate the desired kd% with a database file for every model. The content of the database files consists of a set of binding site parameters objects spanning the complete range of the model input binding site parameters. Additionally the database files contain the models kd% result calculated for the whole set of objects. | + | The binding sites generated from the model results come into play, when the user wants to use his or her own sh/miRNA, or when the experimentally verified binding sites have a knockdown, that is not sufficiently similar to the desired knockdown.<br> |
| - | With the user-chosen sh/miRNA sequence as input a binding site generator script is invoked, which varies the seed-type, 3'-pairing, AU-content and bulge-size of on the fly generated binding sites. The 3'-pairing and the AU-content score of the generated BS are characterized by a modified version of the targetscan_50_context_scores – Algorithm [Rodriguez et al.]. The input and output functions were adapted to the mode of operation of miRockdown, thus no files have to be generated while running miRockdown. | + | A script integrated into miRockdown will correlate the desired kd% with a database file for every model. The content of the database files consists of a set of binding site parameters objects spanning the complete range of the model input binding site parameters. Additionally the database files contain the models kd% result calculated for the whole set of objects.<br> |
| + | With the user-chosen sh/miRNA sequence as input a binding site generator script is invoked, which varies the seed-type, 3'-pairing, AU-content and bulge-size of on the fly generated binding sites. The 3'-pairing and the AU-content score of the generated BS are characterized by a modified version of the targetscan_50_context_scores – Algorithm [Rodriguez et al.]. The input and output functions were adapted to the mode of operation of miRockdown, thus no files have to be generated while running miRockdown.<br> | ||
Now, that the generated binding sites are completely characterized, they can be compared with the parameters of the suitable model BS. The generated BS that fits the parameters of the suitable model BS best is selected as the output BS of miRockdown. | Now, that the generated binding sites are completely characterized, they can be compared with the parameters of the suitable model BS. The generated BS that fits the parameters of the suitable model BS best is selected as the output BS of miRockdown. | ||
Revision as of 00:16, 26 October 2010

 "
"