HKU-Hong Kong/13 July 2010
From 2010.igem.org
(Difference between revisions)
| Line 1: | Line 1: | ||
| - | {{:Team:HKU-Hong_Kong/ | + | {{:Team:HKU-Hong_Kong/header1}} |
=13 July 2010:Con't BBa_k112808 isolation= | =13 July 2010:Con't BBa_k112808 isolation= | ||
Different combinations of enzymes were used for the isolation. | Different combinations of enzymes were used for the isolation. | ||
Latest revision as of 18:20, 27 October 2010

| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
| |||||||||||||||||||||||||||||||||||||||||||||||||||
13 July 2010:Con't BBa_k112808 isolation
Different combinations of enzymes were used for the isolation.
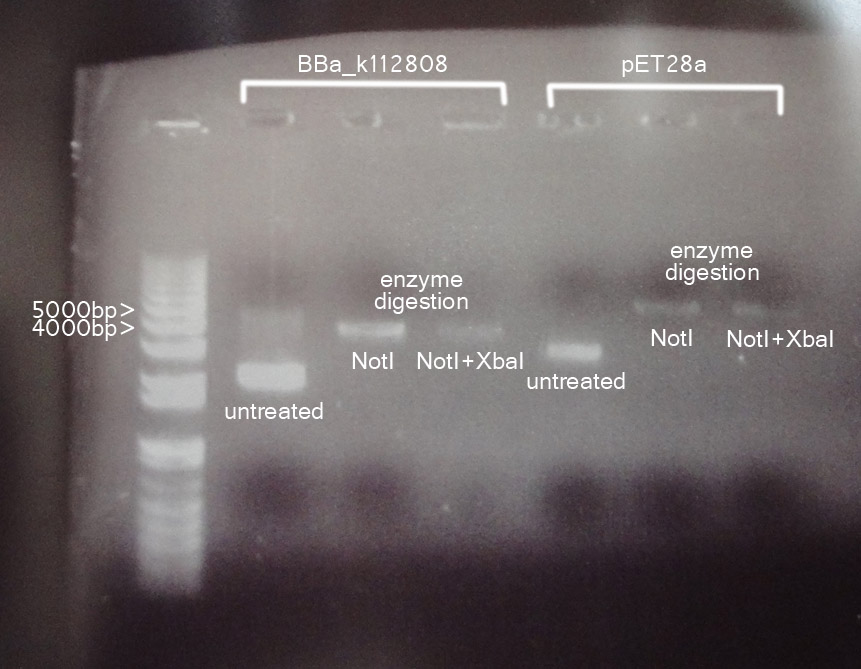
Result
Controls were run in parallel to the plasmid after various treatments. An upward shift indicated the presence of a functional cutting site to turn the faster-moving circular plasmid to slower-moving linear DNA fragments. The results of single enzyme digestion were the same for double enzyme digestion, suggesting that one of the site might be absent.
 "
"