Team:Heidelberg/Project/miMeasure
From 2010.igem.org
(→Flow cytometry) |
|||
| Line 1: | Line 1: | ||
| - | {{:Team:Heidelberg/ | + | {{:Team:Heidelberg/Double}} |
| - | {{:Team:Heidelberg/ | + | {{:Team:Heidelberg/Pagetop|pro_miMeasure}} |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
=miMeasure= | =miMeasure= | ||
==Abstract== | ==Abstract== | ||
| - | With the rising importance of small RNA molecules in gene therapy the identification and characterization of miRNAs and their binding sites become crucial for innovative applications. In order to exploit the miRNA ability to target and regulate specific genes, we constructed a measurement standard (see sidebar) not only to characterize existing miRNAs but also to validate potential synthetic miRNAs for a new therapeutic approach. The synthetic miRNAs we created | + | With the rising importance of small RNA molecules in gene therapy the identification and characterization of miRNAs and their binding sites become crucial for innovative applications. In order to exploit the miRNA ability to target and regulate specific genes, we constructed a measurement standard (see sidebar) not only to characterize existing miRNAs but also to validate potential synthetic miRNAs for a new therapeutic approach. The synthetic miRNAs we created are inert for endogenous targets and thus applicable for gene regulation without any direct side effects. This opens new possibilities for precise expression tuning, especially in quantitative means. |
| - | Our [https://2010.igem.org/Team:Heidelberg/Parts#final_constructs miMeasure] | + | Our [https://2010.igem.org/Team:Heidelberg/Parts#final_constructs miMeasure] construct plasmid normalizes knockdown of the green fluorescent protein (EGFP) to the blue fluorescent protein (EBFP2). This allows for an accurate study of binding site properties, since both fluorescent proteins are combined in the same construct and driven by the same [http://partsregistry.org/Part:BBa_K337017 bidirectional promoter]. Another advantage is, that any desired binding site can be cloned easily into the miMeasure plasmid with the [http://dspace.mit.edu/handle/1721.1/45139 BB-2 standard]. As the binding site is inserted downstream of EGFP, a regulation of EGFP expression is to be expected. |
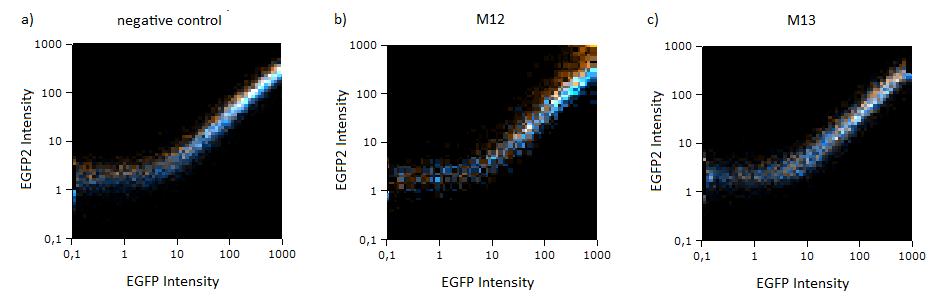
| - | The percentage of knockdown of each modified binding site can be | + | The percentage of knockdown of each modified binding site can be calculated by the ratio of EGFP to EBFP2. This ratio is derived from a linear regression curve. Therefore the knock-down efficiency can be determined by various basic methods e.g. plate reading, flow cytometry or [https://2010.igem.org/Team:Heidelberg/Notebook/Material_Methods#Microscopy microscopy]. |
==Introduction== | ==Introduction== | ||
| - | + | Micro RNAs regulate mainly the translation of their target genes by preferably interacting with regions in the 3’ untranslated region (UTR) of their target mRNA. Base-pairing with the miRNA binding site (BS) causes formation of diverse miRNA-mRNA duplexes {{HDref|reviewed by Fabian et al., 2010}}. The BS consists of a seven nucleotide long seed region that is perfectly matched to the miRNA, and surrounding regions that matched partially. The seed region is defined as being the minimal required base-pairing that can regulate the mRNA. Apart from the seed region, binding can be unspecific, creating mismatches and bulges. The position and properties of the bulges seem to play a central role in miRNA binding and therefore knockdown efficiency {{HDref|reviewed in Bartel et al., 2009}}. Since we were going to use synthetic miRNA BS in our genetherapeutic approach, we had to find a way to study their effects in a standardized manner that would be comparable and reproducible. | |
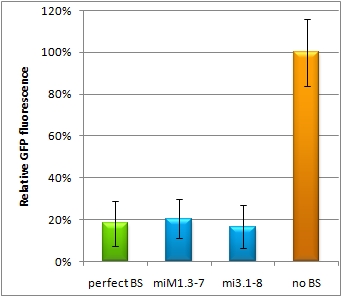
| - | Micro RNAs regulate the translation of their target genes by preferably | + | One goal of the iGEM Team Heidelberg 2010 was to test the effects of changes in BS sequences on expression control. Thereby miRNA BS should be characterized. To standardize our measurements of knockdown according to BS specificity, we had to come up with a new standard that is independent from the endogenous cell machinery. We decided to bring synthetic miRNAs into play, hence we engineered BS for them creating an artificial regulatory circuit<!--, simulating naturally occurring miRNAs and miRNA BS without having to worry about the effect of endogenous targets-->. Of course there are also differences that arise through the availability of the enzymes involved in the miRNA pathway that may differ slightly from cell to cell. Therefore, we also measured the knockdown achieved by the perfect binding site and set this as 100% knockdown efficiency. Ideally, the miRNA would be stably expressed in the cell line, but a uniform co-transfection also leads to an even distribution of synthetic shRNA-like miRNAs (shRNA miRs). Additionally, miRNA levels can be adjusted by differing transfection ratios. |
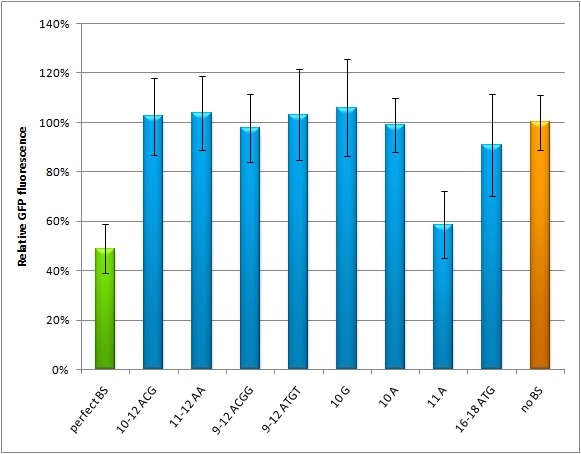
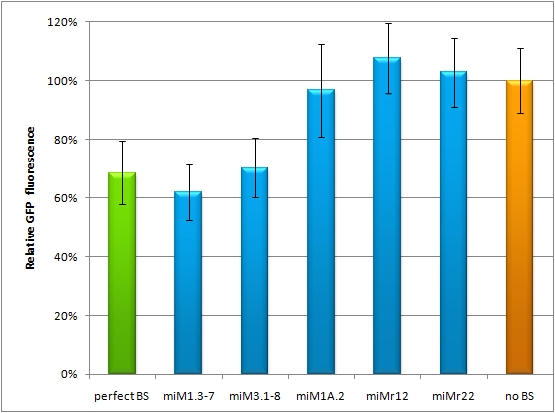
| - | + | The main purpose of our measurement standard, miMeasure, is to express two nearly identical but discernible proteins: one of them tagged with a BS, the other one unregulated (even though the possibility exists to clone in a reference binding site). The two reporters are expressed by a bidirectional CMV promoter to make sure their transcription rate is comparable. We used a destabilized version of GFP, dsEGFP (by Clontech) and a dsEBFP2 that was derived from the same sequence. Thus, we could make sure that both proteins exhibit the same synthesis and degradation properties, making them directly comparable. Hereby, we can also neglect the difference between mRNA and protein knockdown and can take the fluorescence of the marker protein as a direct, linear output of mRNA down-regulation. We included a BBb standard site into our plasmid, which allows to clone BS behind the GFP. If co-transfected with the corresponding shRNA miR, GFP will be down-regulated, while BFP expression is maintained. The ratio of GFP to BFP expression can be used to conclude the knockdown efficiency (in percent, compared to perfect binding site=100% and no binding site=0%) of the BS. Having destabilized marker proteins with a turnover time of two hours enables us not only to avoid accumulation of marker proteins, which would make the knockdown harder to observe, but also to conduct time-lapse experiments. In the future, this could be for example a way to observe dynamic activity patterns of endogenous miRNAs. | |
| - | Since we were going to use synthetic miRNA BS in our genetherapeutic approach, we had to find a way to study their effects in a standardized manner that would be comparable and reproducible. | + | |
| - | + | ||
| - | One goal of the iGEM Team Heidelberg 2010 was to test the effects of changes in BS | + | |
| - | + | ||
| - | The main | + | |
<html> | <html> | ||
<div class="backtop"> | <div class="backtop"> | ||
Revision as of 18:18, 27 October 2010

|
 "
"