Team:Stockholm/30 August 2010
From 2010.igem.org
Contents |
Andreas
Cloning of SOD into pMA.His
Transformation results
From 28 28/8 Good colony yield. Four colonies (SH1-SH4) picked for colony PCR.
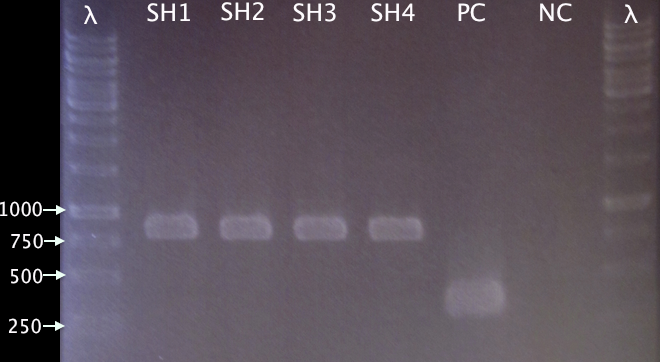
Colony PCR
- SH1-SH4: pMA.SOD⋅His
- PC: Positive control; pMA.His
- NC: Negative control; blank
Procedures according to standard colony PCR protocol. Elongation time 1:00.
Gel verification
1 % agarose, 100 V
Expected bands:
- pMA.SOD⋅His: 831 bp
- pMA.His: 348 bp
Results
Well corresponding bands indicating successful insertion of SOD into the vector.
ON cultures
SH1 and SH2 selected for plasmid prep and sequencing. Set 5 ml LB + 100 Amp ON cultures. 37 °C, 225 rpm.
N-CPP extraction
Gel extraction
From 28/8 samples Purification using the E.Z.N.A. Gel Extraction kit. Elution in 30 μl dH2O; double elution.
| DNA concentrations | ||
|---|---|---|
| Sample | Conc. [ng/μl] | A260/A280 |
| Tra10 | 13.56 | 1.86 |
| TAT | 1.736 | 1.20 |
| LMWP † | 2.523 | 2.69 |
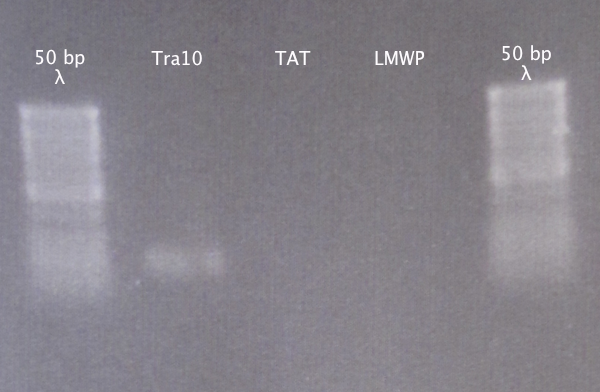
Gel verification
Ran a gel to verify that the presence and size of our extracted DNA fragments.
1 % agarose, 100 V
Results
Weak band for Tra10, no bands for TAT and LMWP. Proceeded to cloning anyway.
Cloning of N-CPPs into pSB1C3
A last-minute decision was made to also make a bulk cloning of all three N-CPPs by digesting directly from the N-CPP cluster vector.
Digestion
[N-CPP plasmid]=672 ng/μl (28/8)
| [ng/μl] | Tra10 | TAT | LMWP | N-CPP |
|---|---|---|---|---|
| 10X FD buffer | 3 | 3 | 3 | 3 |
| dH2O | 3 | 3 | 8 | 23 |
| FD XbaI | 0.5 | 0.5 | 0.5 | 0.5 |
| FD AgeI | 0.5 | 0.5 | 0.5 | 0.5 |
| DNA | 23 | 23 | 18 | 3 |
| 30 | 30 | 30 | 30 |
Incubation: 37 °C, 30 min
Inactivation: 80 °C, 10 min
Dephosphorylation
Treated the N-CPP sample with FastAP alkaline phosphatase to prevent multiple insertions (3, 5, etc...) into target vector.
- 3 μl FastAP
- Incubation: 37 °C, 10 min
- Inactivation at 75 °C, 5 min
Ligation
[Dig. pSB1X3 X+A EXTR]=13.72 ng/μl (digested and extracted vector from 9/8) [Dig. N-CPP X+A]=60 ng/μl
| [ng/μl] | N-CPP | Tra10 | TAT | LMWP |
|---|---|---|---|---|
| Vector DNA | 3 | 3 | 3 | 3 |
| Insert DNA | 9 | 12 | 12 | 12 |
| 5X Rapid Lig. buf. | 4 | 4 | 4 | 4 |
| dH2O | 3 | 0 | 0 | 0 |
| T4 DNA Ligase | 1 | 1 | 1 | 1 |
| 20 | 20 | 20 | 20 |
Transformation
Standard transformation protocol.
- 2 μl ligation mix
- Cm 25
Mimmi
MITF-M
Colony PCR
- Made by Andreas --> ~280bp bands, empty vector? should not be possible...
- Try again, more colonies
| Mix | (µl) | X8 | Primers | Conditions | ||||
|---|---|---|---|---|---|---|---|---|
| sH2O | 22.5 | 180 | pSB1_VF2 | time | °C | |||
| F primer | 1 | 8 | pSB1_VR | 2m | 94 | |||
| R primer | 1 | 8 | 30s | 94 | ) | |||
| DNA | 0.5 | 8x0.5 | 30s | 50 | > 30 cycles | |||
| tot | 25µl | 2m40s | 72 | ) | ||||
| 10m | 72 | |||||||
| oo | 10 | |||||||
Gel
| well | sample |
|---|---|
| 1 | ladder |
| 2 | pSB1C3.MITF-M 1 |
| 3 | pSB1C3.MITF-M 2 |
| 4 | pSB1C3.MITF-M 3 |
| 5 | pSB1C3.MITF-M 4 |
| 6 | pSB1C3.MITF-M 5 |
| 7 | pSB1C3.MITF-M 6 |
| 8 | pSB1C3.MITF-M 7 |
| 9 | blank |
- No product, trying another 7 colonies over night...
 |

|
 |

|
 |

|

|

|
 "
"