Team:NYMU-Taipei/Experiments/Protocols
From 2010.igem.org
| Home | Project Overview | Speedy reporter | Speedy switch | Speedy protein degrader | Experiments and Parts | Applications | F.A.Q | About Us |
Contents |
Basic Cloning Protocols
The common protocols used for our cloning procedures.
Plasmid extraction
[Min 30mins]
Follow the protocol based on the kit used.
| Protocol | GeneArt | Qiagen | Viogene |
|---|---|---|---|
| harvest the plasmid: 1min @ 13200rpm | |||
| add PD1/P1/MX1 and resuspend pellet | 200uL | 200uL | 200uL |
| add PD2/P2/MX2 and wait | 200uL; 2 mins | 250uL; 2-5mins | 250uL; 2-5mins |
| add PD3/P3/MX3 and centrifuge @ 13200rpm | 300uL; 3-10mins | 350uL; 10mins | 350uL; 10mins |
| prepare the columns and the new eppendorfs and think about digestion | |||
| Transfer the supernatant into a column and centrifuge for 1min @ 13200rpm | |||
| add W1/PB/WN and centrifuge for 30sec; discard flowthrough | 400uL; 13200rpm | skipped | 500uL; 9000rpm |
| add Wash/PE/WS and centrifuge for 30sec; discard flowthrough | 600uL; 13200rpm | 700uL | 700uL; 9000rpm |
| Centrifuge for extra 3min @ 13200rpm to get rid of EtOH | |||
| Move column to a the new 1.5ml eppendorf. | |||
| Add 30uL of EB (elution buffer) in center of column and wait 5mins then centrifuge for 3min @ 13200rpm. Start thawing digestion stuff when waiting. | |||
| take the 30uL flowthrough and put it in center of column again and wait >5mins then centrifuge for 3min @ 13200rpm. Make digestion mix while waiting. | |||
Digestion
[Min 20mins]
We use Fermentas FastDigest Enzymes. Digestion Mix:
| SP | XP | XP from PCR (after purification) | |
| DNA | 10 | 16 | 26 |
| 10x FD Buffer | 2 | 2 | 2 |
| Enzyme1 | 1 | 1 | 1 |
| Enzyme2 | 1 | 1 | 1 |
| ddH20 | 6 | - | - |
| Total | 20uL | 20uL | 30uL |
Put in PCR machine incubator for anywhere between 15mins to overnight. Usually 1-3hrs (during lunch time).
Gel extraction and PCR purification
[Min 1hr; 10min]
We use Qiagen's Gel extraction and PCR purification kits.
- XP plasmid digests require gel extraction
- SP plasmid digests and XP PCR digests only need purification
- Protocol:
- Only this first step is different for the two. The other steps are the same.
- Gel extraction: Add 200/400/600uL QG depending on the size of the gel cut. Warm to 50oC; can vortex. When gel completely dissovles, then transfer to column.
- Purfication: Add 5x PB to the solution then transfer to column (using the same tip).
- Centrifuge 1min @ 13200rpm. Discard flowthrough.
- Add 700uL PE. Centrifuge for 30sec @ 13200rpm. Discard flowthrough.
- Centrifuge again for 3mins @ 13200rpm to get rid of excess EtOH.
- Put the column on a new 1.5ml eppendorf
- Add 30uL EB in center of column and wait 5mins then centrifuge for 3min @ 13200rpm.
- Take the 30uL flowthrough and put it in center of column again and wait >5mins then centrifuge for 3min @ 13200rpm.
- Measure concentration using the Nanodrop machine.
- Only this first step is different for the two. The other steps are the same.
Ligation
[Min 15min]
We use Fermentas' T4 ligase. Ligation mix:
| amount (uL) | |
|---|---|
| DNA1 | * |
| DNA2 | * |
| T4 Ligase Buffer | 1 |
| Ligase | 0.5 |
| ddH20 | * |
| Total | 10 |
DNA1+DNA2+water must not exceed 8.5uL. Total DNA concentration must be in between 25-100ng in the reaction. It is recommended that the molar weight is used to calculate the amount of DNA added.
Transformation
[Min 10min]
We use Yeastern's DH5α competent cells.
- Protocol
- aliquot 2uL of plasmid DNA or 5uL of ligation mix into a new PCR tube
- thaw 20uL of competent cell and mix it in the PCR tube.
- Put on ice for >5mins (the longer the better)
- Heatshock for 45sec @ 42oC
- Put on ice for 1min (the longer the worse)
- Put on plate.
The 3-in-1
[Min 10min + 1min/colony]
Out 3-in-1 protocol consists of a combination of three things:
- Colony PCR
- Liquid culture
- 2nd time plate
Preparation
First, make the Colony PCR mix: (We use Fermentas' DreamTaq)
| uL | |
| Template | 1 |
| FP (VR) | 1 |
| RP (VF2) | 1 |
| dNTP (2.5mM) | 2 |
| 10x DreamTaq buffer | 5 |
| Taq Polymerase | 0.2 |
| ddH20 | 39.8 |
| Total | 50 |
However, we only use 10-15uL for each colony. And we make the mix in the Negative control if total is less than ~220uL. Also 5uL/reaction is not enough.
Second, divide up the second time plate and Add Ampicillin (or another antibiotic) into the liquid at 1uL per 1ml (so 4uL for 4ml liquid culture).
Protocol
- 3-in-1:
- Select a colony using a tip.
- Dip it in a PCR tube and swirl it around.
- Put it on a pipette and dip it in the liquid culture OR dip it in 10uL ddH20 for later use when colony is confirmed to be correct.
- Take the tip off the pipette and use it to draw on the right place on the 2nd time plate.
- Repeat.
- Put the PCR tubes in the PCR machine with this protocol:
| 94oC | 60sec | |
| 94oC | 15sec | 30 cycles |
| 55oC | 20sec | |
| 72oC | 60sec/1kb + ~0-15sec | |
| 72oC | 300sec |
Put the liquid and the 2nd time plate in a 37oC incubator, and the liquid shaking at 180-200 rpm.
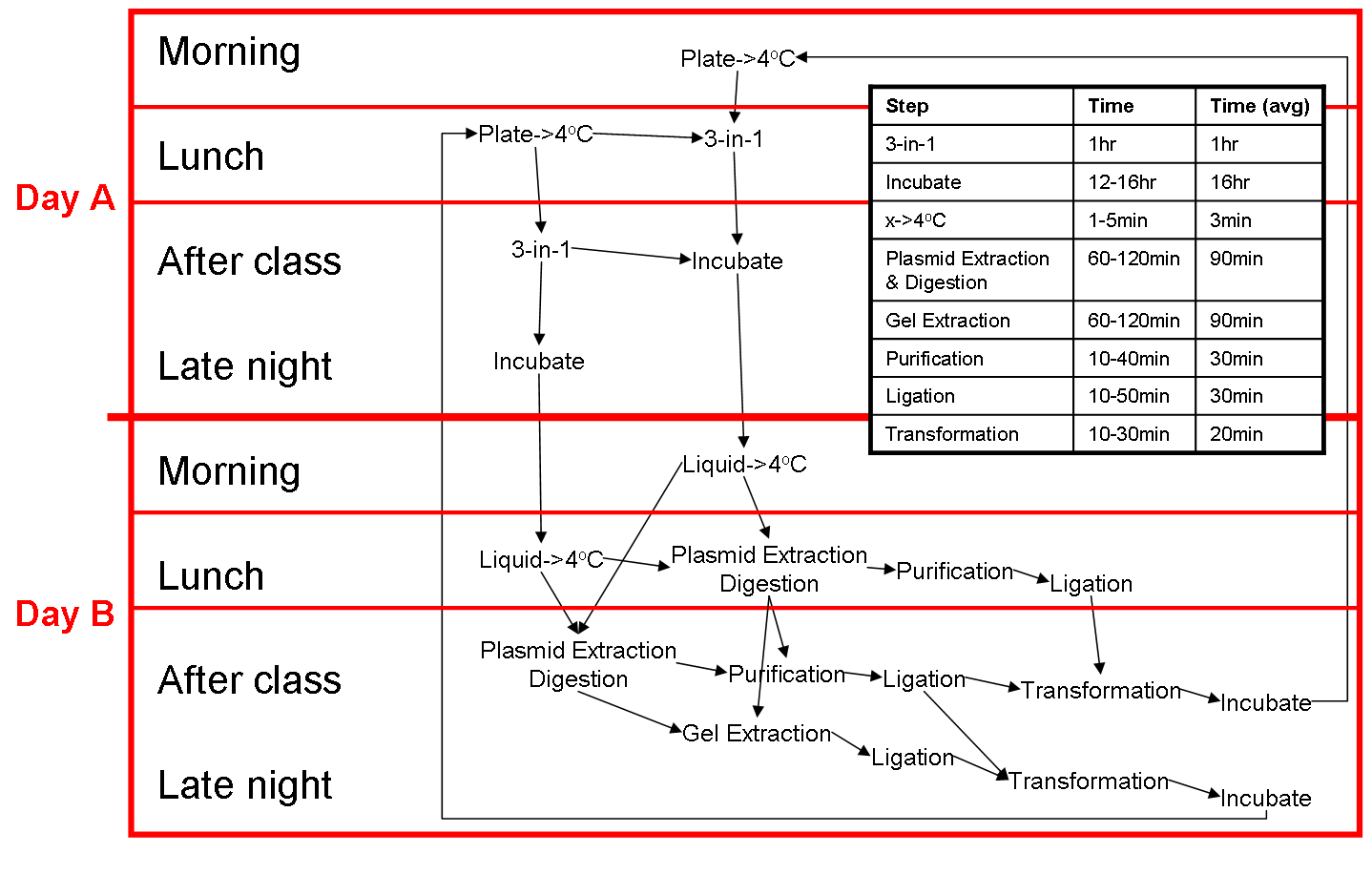
Two Day cloning cycle
Our two day cloning cycle on a typical summers day (when we're free all day) and on a typical semester's day (when our daytime is restricted). This two day protocol can be modified into a 1.5 day protocol by reducing the time of overnights. However, the concentration of the extracted plasmid will be reduced, the colonies are smaller and harder to pick.
Typical Summer's Day
Day A
- 10am (5min): Plate->4oC
- 10:05am(~1hr): 3-in-1, put liquid and plate in fridge.
- Lunch
- After lunch (~1hr): Run Gel
- 5pm: Incubate
Day B
- 10am (2-3hrs): Plasmid extraction & Digestion
- Lunch (while digesting)
- After lunch (2-3min): Gel Extraction/Purification + Ligation + Transformation
- 5pm: Incubate plate
Typical Semester's Day
Gene Reporter Assay
This section describes the gene reporter assay protocol we used. The first subsection is a generalized one, and the other subsections are specific modifications with parameters for the specified experiment.
Generalized protocol
The final raw data obtained using this method is:
- Two OD points, one at n1 hours, and one at n1+n2 hours. Where:
- n1: The amount of time between the dilution of the overnight culture and the first fluorescence measurement.
- n1+n2: The amount of time between the dilution of the overnight culture and the final fluorescence measurement.
- The continuous measurement of fluorescence with one fluorescence data point every m minutes
Protocol:
- Selected genes to be reported are incubated overnight in an LB liquid culture at 37oC and 180-200rpm. This makes sure they are fresh in the morning. Positive and negative controls are also needed.
- The OD600 of each liquid culture and all diluted to the same OD600 ["OD at 0 hours"] and incubated for n1 hours.
- Measurement of OD at n1 hours: For each used well in the 96-well plate:
- Take 200uL from the liquid (make sure you pipette this step well) and put it in a cuvette to read the OD600.
- Note down the OD600 ["OD at n1 hours"], then take the liquid in the cuvette and put it in the right place in the 96-well plate.
- Measurement of fluorescence:
- Take the 96-well plate, and directly culture it and perform reads directly in the Eliza reader for n2 hours. Measurement is done every m minutes.
- Measurement of OD at n1+n2 hours: For each used well in the 96-well plate:
- Take the liquid from the well and put it in the cuvette to measure the OD ["OD at n1+n2 hours"].
Speedy switch testing protocol
The parameters have a range because we performed the testing with a different parameters.
- Overnight liquid culture is diluted to OD600 of 0.1, theophylline is added at concentrations ranging from 0.01mM to 20mM, and the mix incubated for 2-2.5 hours.
- Continuous measurement of fluorescence with the excitation/emission wavelengths 488/511nm for 2 hours, with one fluorescence data point every 2 minutes.
Speedy protein degrader testing protocol
- Overnight LB liquid culture is diluted to OD600 of 0.1 and incubated for 1-2 hours.
- After the short incubation, the medium is changed from LB to ABT (AB medium+2.5ug/uL Thiamine) by:
- Centifuging the liquid culture in an Eppendorf at 13200rpm for 1 min (same as plasmid extraction) to obtain the pellet.
- The LB is removed; the pellet washed with ABT medium which is then removed.
- The pellet is resuspended in pre-heated 37oC ABT medium, then shaken for a minute to make sure it's mixed.
- Continuous measurement of fluorescence for 1-2 hours, with one fluorescence data point every 2 minutes. Excitation/emission wavelengths:
 "
"