Team:Cambridge/Bioluminescence/Vibrio Characterisation
From 2010.igem.org

This page describes characterisation for part BBa K325909, the lux operon from Vibrio fischeri.
Description

This page described the lux operon from Vibrio fischeri. To relieve LuxR control we placed Lux C, D, A, B, E under the pBad promoter.
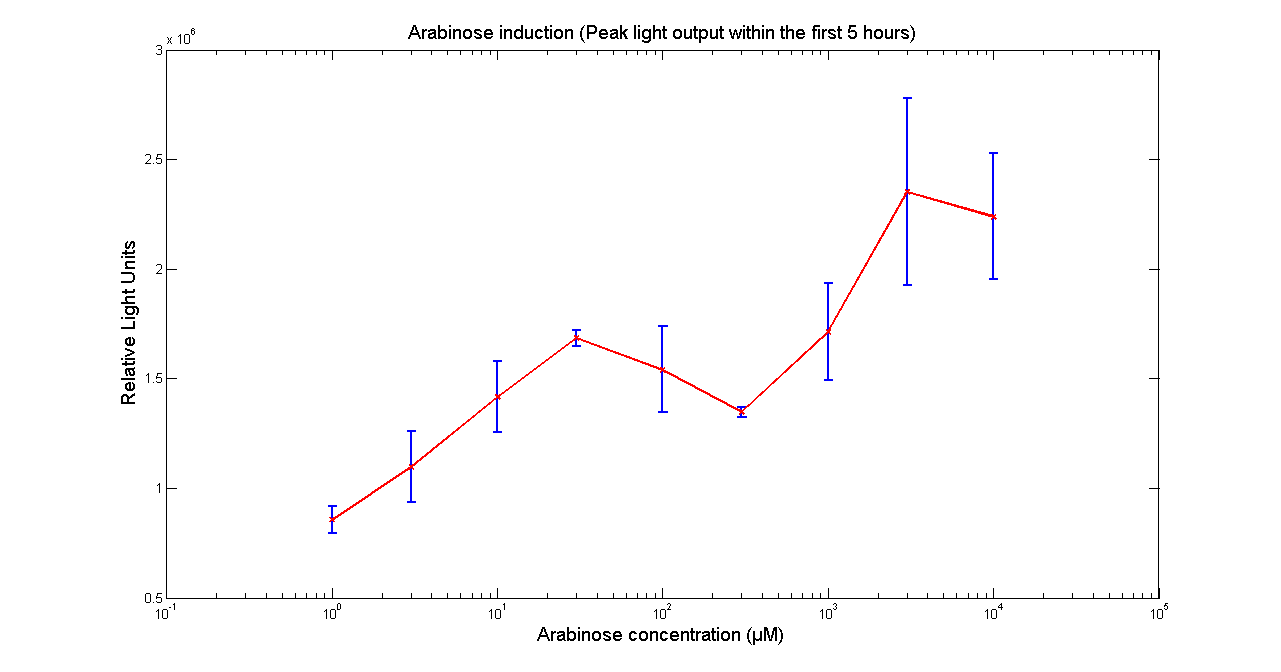
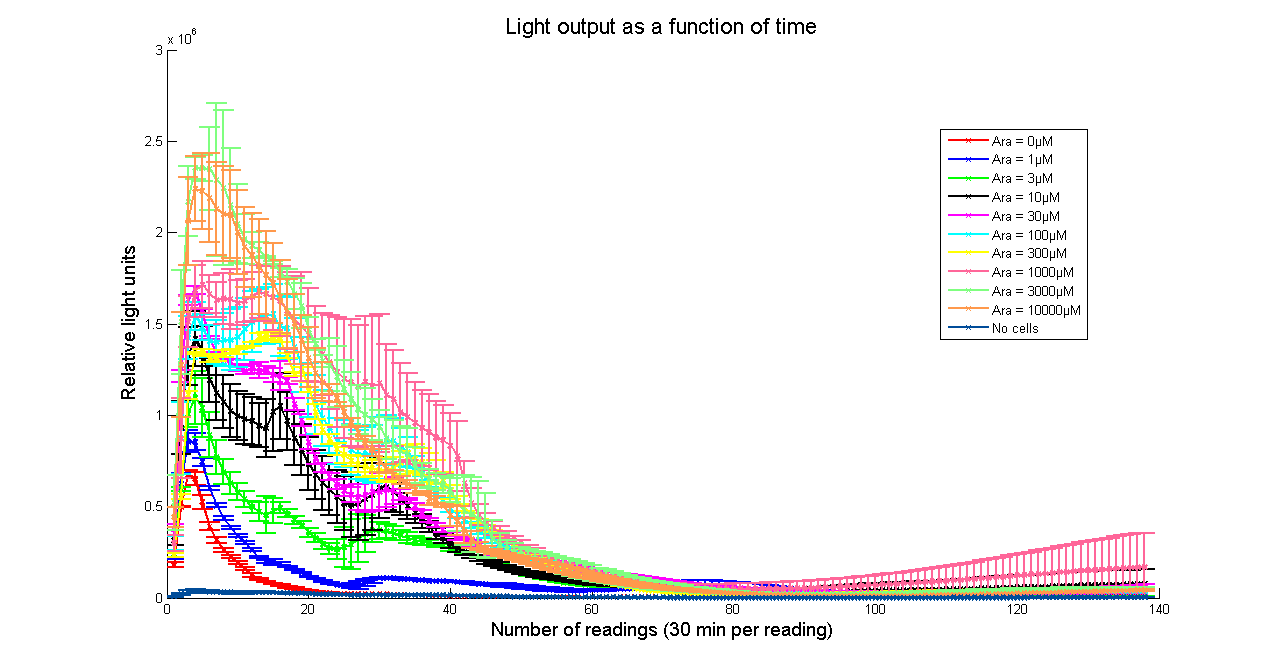
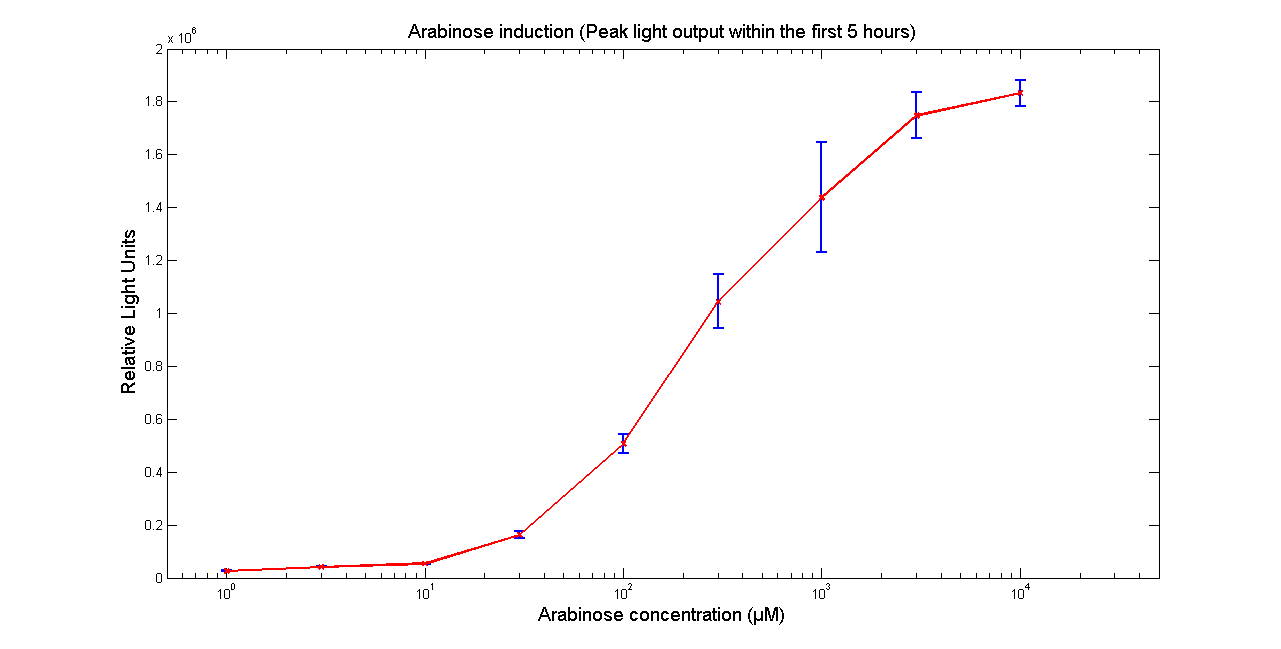
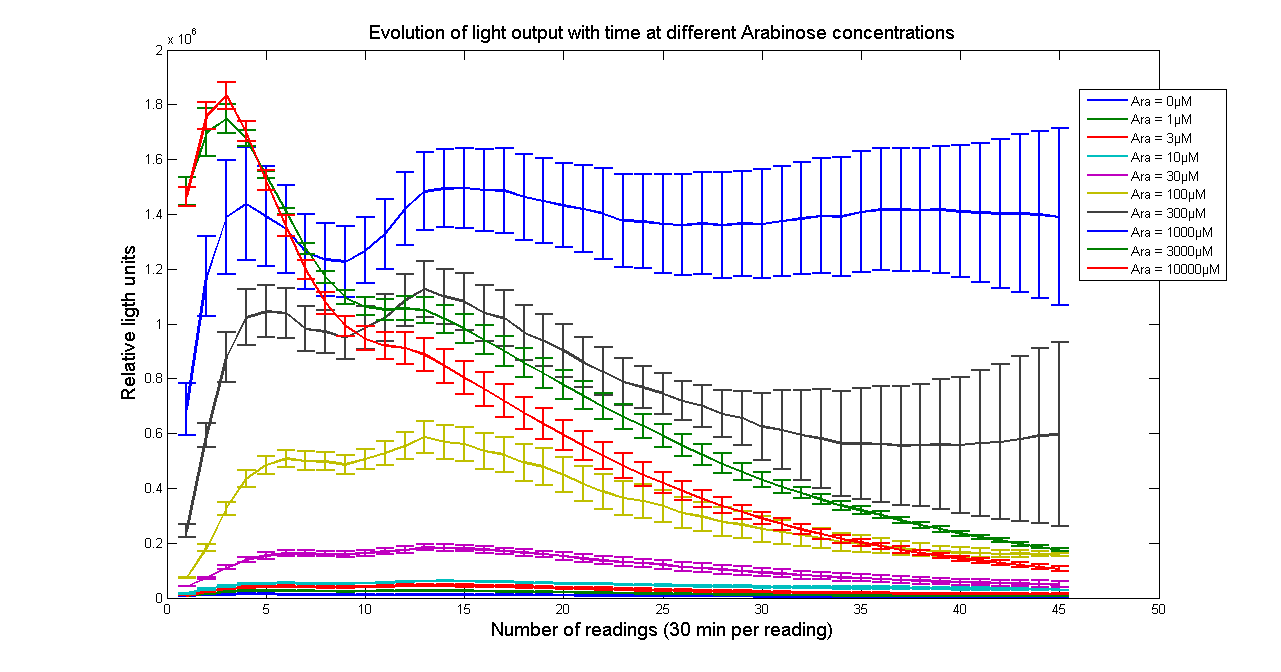
Arabinose to light
This page describes the relationship between Arabinose concentration in the medium with light output. We used a FLUOstar OPTIMA microplate reader to quantify the light output. Protocol and plate reader settings used are given below.
Data
| Data | Notes | Date Uploaded |
|---|---|---|
| Media:BBa_K325909ArabinosetoLight.xls | Raw data from experiment | 21/10/2010 |
H-NS mutants
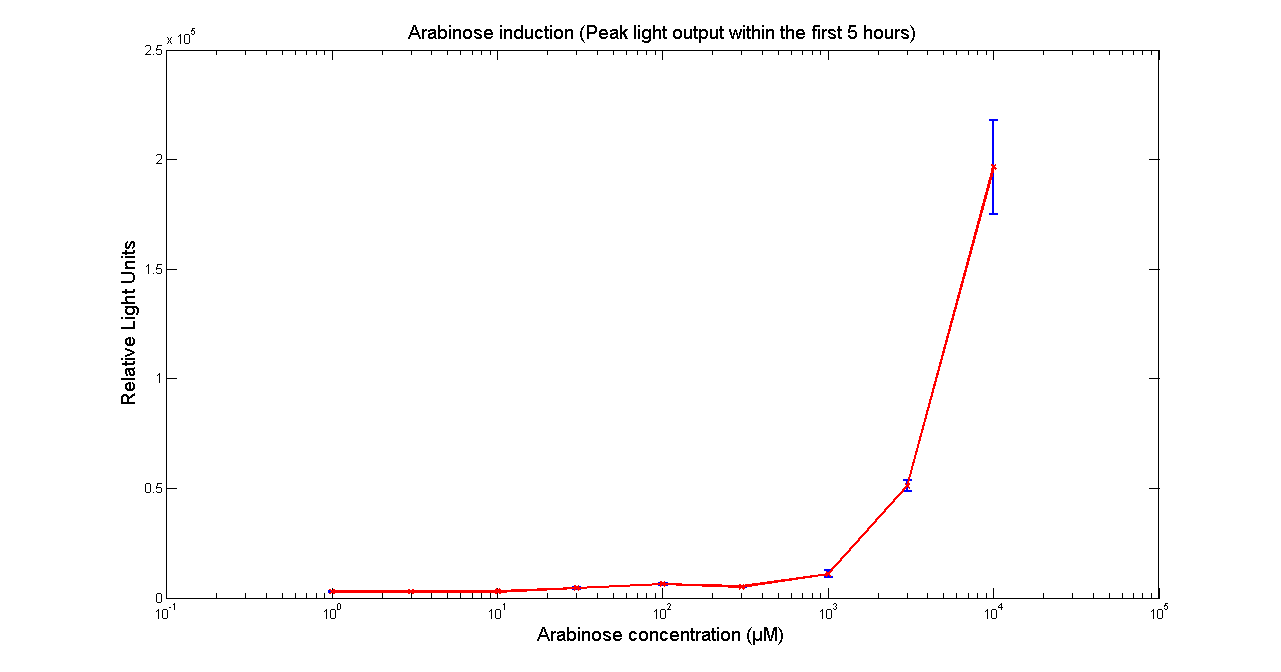
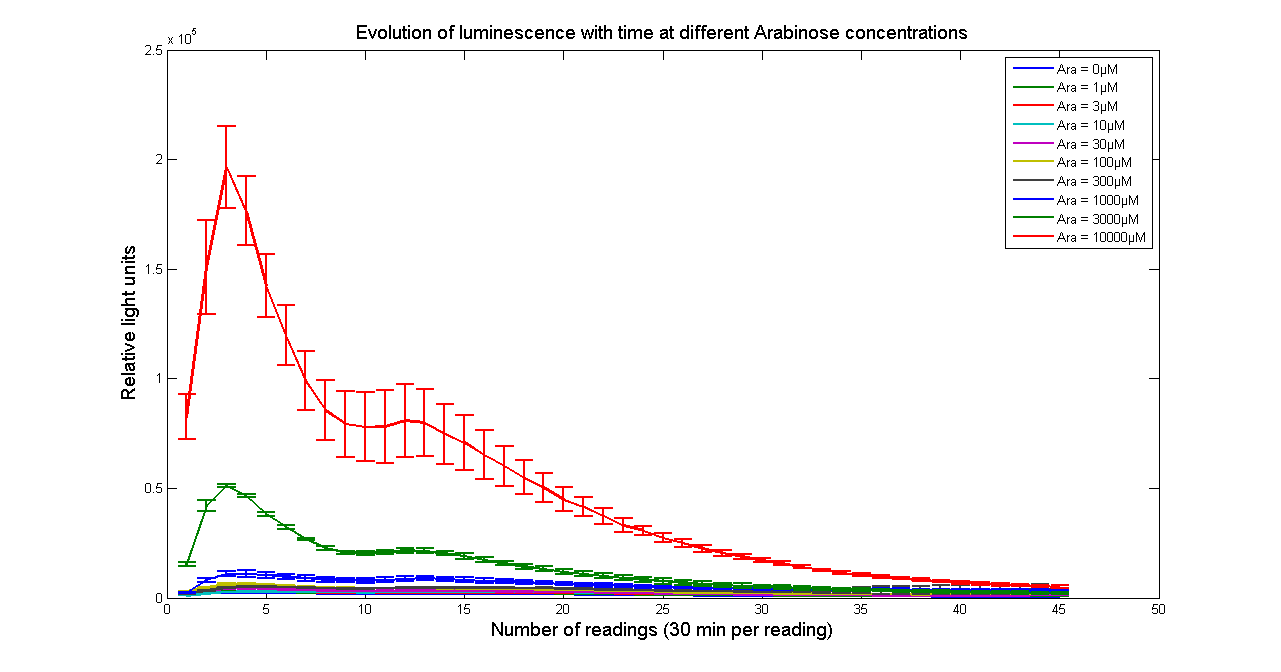
It has been shown that the expression of the Vibrio fischeri lux operon when cloned into E. coli was repressed. This repression was linked to the nucleoid protein H-NS. To investigate this effect we cloned the operon into mutant E.coli cells in which the expression of the H-NS protein had been modified. We used a FLUOstar OPTIMA microplate reader to quantify the light output. Protocols and plate reader settings used are given below.
Data
| Data | Notes | Date Uploaded |
|---|---|---|
| Media:BBa_K325909Mutants.xls | Raw data from experiment | 21/10/2010 |
Compatibility
Chassis: Device has been shown to work in Top 10 (Invitrogen), BW25113 DELTA H-NS::kan and H-NS mutant JM 230 H-NS -205::tn10.
Plasmids: Device has been shown to work on <partinfo>pSB1C3</partinfo>
References
[1]: J. Slock, (1995) Transformation Experiments Using Bioluminescence Genes of Vibrio fischeri,The American Biology Teacher, 57, 225-227.
[2]: E.A. Meighen (1988) Enzymes and genes from the lux operons of bioluminescent bacteria, Annual Reviews in Microbiology 42, 151-176.
[3]: E.A. Meighen, (1994) Genetics of bacterial bioluminescence, Annual Reviews of Genetics, 28, 117-139.
[4]: S. Ulitzur, (1998) H-NS controls the transcription of three promoters of Vibrio fischeri lux cloned in Escherichia coli,Journal of Bioluminescence and Chemiluminescence, 13(4), 185-188.
[5]: R.T. Dame et al., (2006) Bacterial chromatin organization by H-NS protein unravelled using dual DNA manipulation,Nature, 444, 387-390.
 "
"