Team:Bielefeld-Germany/Project/Approach
From 2010.igem.org
(→Directed mutagenesis) |
(→The Approach) |
||
| (9 intermediate revisions not shown) | |||
| Line 23: | Line 23: | ||
= The Approach = | = The Approach = | ||
| - | + | In order to create a novel biosensor we started to search for a sensor system which is able to detect certain chemicals and we looked for additional substances we could try to detect. We were looking for a system that should be taken out of a different bacteria species than ''Escherichia coli'' in order to avoid any background, when working with ''E. coli'' in the end. Therefore we checked the literature for a well reviewed sensor system, which is not part of the ''E. coli'' genome. Finally we decided to use the phenolic sensing VirA/G signaling system of ''Agrobacterium tumefaciens''. It naturally detects acetosyringone, which is a secondary metabolite of plants that affects bacteria as an attractant. After repeated research to check for any other possible substances which could be detected by the system, we got a really long list of possibilities and picked 'capsaicin', which is responsible for the spiciness of edibles. | |
| - | The biological steps for | + | The biological steps for modulating the native VirA/G sensor system, in order to detect new substances are: |
| - | # | + | # Extract the complete VirA/G signaling system out of ''A. tumefaciens'' |
| - | # | + | # Create new BioBricks out of the existing environmental parts |
| - | # | + | # Get the new BioBricks working into ''E. coli'' |
| - | # | + | # Choose a reporter gene as readout for the sensor system |
| - | + | # Modify the system for sensibility and specificity by error-prone PCR, find and select the most promising mutants (directed mutagenesis) | |
=== Preparing the system === | === Preparing the system === | ||
| - | We tried to work with the already existing | + | We tried to work with the already existing VirA receptor BioBrick from the registry (<partinfo>K238008</partinfo>). Unfortunately this VirA BioBrick did not work in our experiments. We decided to sequence this BioBrick from the registry on our own and found that the actual sequence data of this BioBrick did not fit to the published sequence. Thus, in order to create this BioBrick we had to extract the ''virA'' gene via PCR out of the TI-plasmid from ''A. tumefaciens''. |
| - | VirA | + | To construct the complete sensor system VirA is not enough, since it works only in combination with the VirG protein. The''virG'' gene was also present in the iGEM registry, but this version containes illegal restriction sites. Moreover it needs the help of the RpoA protein from ''A. tumefaciens'' to work as a transcription facor in ''E. coli''. In order to eliminate these drawbacks we changed the sequence manually and had the gene synthesized by Mr. Gene. |
| - | === Starting point for | + | === Starting point for BioBricks -> Go cloning=== |
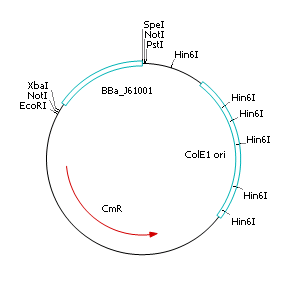
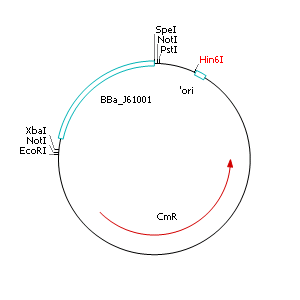
| - | + | After discussing the possibility of creating only one big plasmid for mutated ''virA'' screenings, we unified that its size would minimize transformation efficiency and would be difficult to modify via error-prone PCR. Thereby we decided to use a system with two plasmids per cell that offers certain advantages, which will be explained later. Prior to the construction of the plasmids we had to change the origin of a pSB1C3 plasmid in order to avoid incompatibility between both plasmids. Thus we cloned the R6K origin (<partinfo>J61001</partinfo>) into the <partinfo>pSB1C3</partinfo> plasmid and subsequently removed the original pMB1/ColE1 origin of replication. Therefore we digested the pSB1C3::R6K plasmid with Hin6I and seperated the resulting 6 fragments (1758 bp, 270 bp, 174 bp, 109 bp, 100 bp, 67 bp) by agarose gel electrophoresis. We extracted the largest fragment (1758 bp) and religated it. The resulting plasmid still contains 32 remaining basepairs of the pMB1 origin, but this fragment does not enable replication in pir<sup>-</sup> strains. | |
[[Image:pSB1C3-R6K_300x300px.jpg]] [[Image:pSB1C3-R6K-ColE1del_300x300px.jpg]] | [[Image:pSB1C3-R6K_300x300px.jpg]] [[Image:pSB1C3-R6K-ColE1del_300x300px.jpg]] | ||
| - | |||
| - | After | + | === The error prone PCR and screening === |
| + | |||
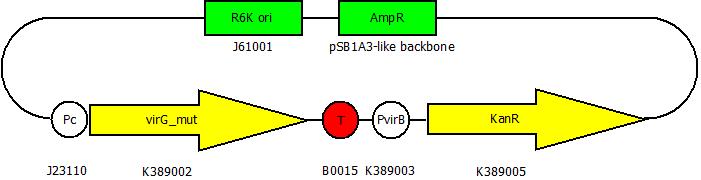
| + | After building a plasmid with R6K ori we were able to create the two constructs. The first plasmid will be found inside the competent bacteria cells and contains the ''virG'' gene under the constitutive promoter <partinfo>J23110</partinfo>, a terminator (<partinfo>B0017</partinfo>), a ''vir'' promoter and a readout or selection gene (luciferase, mRFP and kanamycin resistance, respectively): | ||
[[Image:Screening_plasmid.jpg]] | [[Image:Screening_plasmid.jpg]] | ||
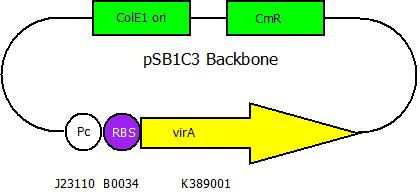
| - | The second plasmid contains | + | The second plasmid contains variants of ''virA'' ,which have been altered randomly by error-prone PCR, under the control of the constitutive promoter <partinfo>J23110</partinfo>. |
| Line 61: | Line 62: | ||
| - | The error prone PCR is a PCR under malfunction conditions | + | The error-prone PCR is a PCR using in inaccurate Taq polymerase under intesified malfunction conditions. It is possible to regulate the frequency of mutagenic events by using unbalanced concentrations of nucleotides and special salt concentrations in the PCR mixture. Using this mutagenesis we are going to create a new variants of the ''virA'' BioBrick and transform it in our cells with the read out system already present. |
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
=== Survival of the fittest === | === Survival of the fittest === | ||
| - | We | + | We used high amounts of the antibiotic kanamycin in order to select the most specific systems for the detection of the chosen substances. By varying the amount of kanamycin we will be able to carefully select some promising clones out of thousands bacteria. The mutated ''virA'' system will be induced after the error-prone PCR by a mix of possible targets for the system (like capcaicin, dopamin, homovanillic acid etc.). |
| - | === | + | === Side directed mutagenesis === |
| - | + | ||
| - | + | Alternatively to the random mutagenesis we developed two in-silico based strategies for side directed mutagenesis as described by [http://www.nature.com/nature/journal/v423/n6936/abs/nature01556.html Looger ''et al.'']. | |
| - | ''' | + | '''VirA_mut1''' |
| - | + | to develop a strategy for change in the protein sequence, we used the analysis tool Muscle, which provided an advice to change one amino acid in the receptor. This conversion of leucin 293 to tyrosine was performed using site directed mutagenesis | |
| - | + | '''VirA_mut2''' | |
| - | + | Since VirA_mut1 with its single mutation failed working in our later screening system, we decided to mutate more amino acids in the area of the predicted binding region from animal TRPV1 receptors. | |
| - | + | According to the literature the amino acid residues 288 to 293 of this TRPV1 receptor regulate phenol structural specificity. Substitutions in this region narrow the active phenols to those missing the para-acyl substituent ([http://www.springerlink.com/content/978-0-387-78884-5#section=169995&page=1 Lin ''et al.'', 2008]). Our VirA_mut2 construct contains the following substitutions: | |
| + | *A357H *R358P *R359T *L360L *D361K *Y361L | ||
| - | |||
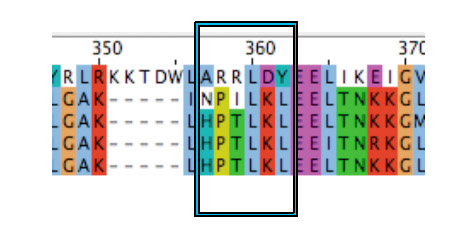
| - | + | [[Image:Bielefeld_VirA_mut2.jpg|300px|thumb|center|'''Alignment of crucial ligand binding regions from ''Agrobacterium tumefaciens'' (lane 1), chick (lane 2), ''homo sapiens'' (lane 3), ''rattus'' (lane 4) and ''mus muskulus'' (lane 5). The Box shows all six amino acids we mutagenized in our VirA_mut2 approach''']] | |
| - | + | === The final construct=== | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | In order to get a visible light signal by the reporter gene, we integrated the quantifiable proteins luciferase and mRFP. We also experimented with certain sensitivity tuners to be able to enhance the detected signal. | |
| - | |||
| - | |||
| + | For a demonstration of this final system see the animation below: | ||
[[Image:Bielefeld_Vorgehen.gif]] | [[Image:Bielefeld_Vorgehen.gif]] | ||
Latest revision as of 03:10, 28 October 2010
Contents |
The Approach
In order to create a novel biosensor we started to search for a sensor system which is able to detect certain chemicals and we looked for additional substances we could try to detect. We were looking for a system that should be taken out of a different bacteria species than Escherichia coli in order to avoid any background, when working with E. coli in the end. Therefore we checked the literature for a well reviewed sensor system, which is not part of the E. coli genome. Finally we decided to use the phenolic sensing VirA/G signaling system of Agrobacterium tumefaciens. It naturally detects acetosyringone, which is a secondary metabolite of plants that affects bacteria as an attractant. After repeated research to check for any other possible substances which could be detected by the system, we got a really long list of possibilities and picked 'capsaicin', which is responsible for the spiciness of edibles.
The biological steps for modulating the native VirA/G sensor system, in order to detect new substances are:
- Extract the complete VirA/G signaling system out of A. tumefaciens
- Create new BioBricks out of the existing environmental parts
- Get the new BioBricks working into E. coli
- Choose a reporter gene as readout for the sensor system
- Modify the system for sensibility and specificity by error-prone PCR, find and select the most promising mutants (directed mutagenesis)
Preparing the system
We tried to work with the already existing VirA receptor BioBrick from the registry (<partinfo>K238008</partinfo>). Unfortunately this VirA BioBrick did not work in our experiments. We decided to sequence this BioBrick from the registry on our own and found that the actual sequence data of this BioBrick did not fit to the published sequence. Thus, in order to create this BioBrick we had to extract the virA gene via PCR out of the TI-plasmid from A. tumefaciens.
To construct the complete sensor system VirA is not enough, since it works only in combination with the VirG protein. ThevirG gene was also present in the iGEM registry, but this version containes illegal restriction sites. Moreover it needs the help of the RpoA protein from A. tumefaciens to work as a transcription facor in E. coli. In order to eliminate these drawbacks we changed the sequence manually and had the gene synthesized by Mr. Gene.
Starting point for BioBricks -> Go cloning
After discussing the possibility of creating only one big plasmid for mutated virA screenings, we unified that its size would minimize transformation efficiency and would be difficult to modify via error-prone PCR. Thereby we decided to use a system with two plasmids per cell that offers certain advantages, which will be explained later. Prior to the construction of the plasmids we had to change the origin of a pSB1C3 plasmid in order to avoid incompatibility between both plasmids. Thus we cloned the R6K origin (<partinfo>J61001</partinfo>) into the <partinfo>pSB1C3</partinfo> plasmid and subsequently removed the original pMB1/ColE1 origin of replication. Therefore we digested the pSB1C3::R6K plasmid with Hin6I and seperated the resulting 6 fragments (1758 bp, 270 bp, 174 bp, 109 bp, 100 bp, 67 bp) by agarose gel electrophoresis. We extracted the largest fragment (1758 bp) and religated it. The resulting plasmid still contains 32 remaining basepairs of the pMB1 origin, but this fragment does not enable replication in pir- strains.
The error prone PCR and screening
After building a plasmid with R6K ori we were able to create the two constructs. The first plasmid will be found inside the competent bacteria cells and contains the virG gene under the constitutive promoter <partinfo>J23110</partinfo>, a terminator (<partinfo>B0017</partinfo>), a vir promoter and a readout or selection gene (luciferase, mRFP and kanamycin resistance, respectively):
The second plasmid contains variants of virA ,which have been altered randomly by error-prone PCR, under the control of the constitutive promoter <partinfo>J23110</partinfo>.
The error-prone PCR is a PCR using in inaccurate Taq polymerase under intesified malfunction conditions. It is possible to regulate the frequency of mutagenic events by using unbalanced concentrations of nucleotides and special salt concentrations in the PCR mixture. Using this mutagenesis we are going to create a new variants of the virA BioBrick and transform it in our cells with the read out system already present.
Survival of the fittest
We used high amounts of the antibiotic kanamycin in order to select the most specific systems for the detection of the chosen substances. By varying the amount of kanamycin we will be able to carefully select some promising clones out of thousands bacteria. The mutated virA system will be induced after the error-prone PCR by a mix of possible targets for the system (like capcaicin, dopamin, homovanillic acid etc.).
Side directed mutagenesis
Alternatively to the random mutagenesis we developed two in-silico based strategies for side directed mutagenesis as described by [http://www.nature.com/nature/journal/v423/n6936/abs/nature01556.html Looger et al.].
VirA_mut1
to develop a strategy for change in the protein sequence, we used the analysis tool Muscle, which provided an advice to change one amino acid in the receptor. This conversion of leucin 293 to tyrosine was performed using site directed mutagenesis
VirA_mut2
Since VirA_mut1 with its single mutation failed working in our later screening system, we decided to mutate more amino acids in the area of the predicted binding region from animal TRPV1 receptors.
According to the literature the amino acid residues 288 to 293 of this TRPV1 receptor regulate phenol structural specificity. Substitutions in this region narrow the active phenols to those missing the para-acyl substituent ([http://www.springerlink.com/content/978-0-387-78884-5#section=169995&page=1 Lin et al., 2008]). Our VirA_mut2 construct contains the following substitutions:
- A357H *R358P *R359T *L360L *D361K *Y361L
The final construct
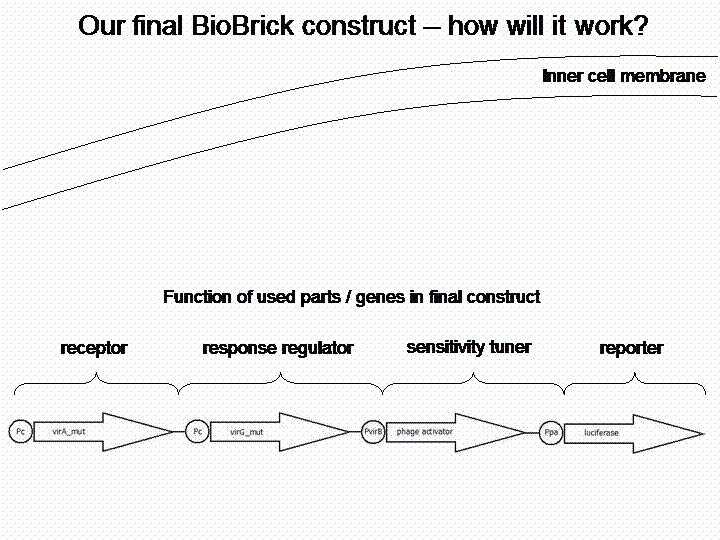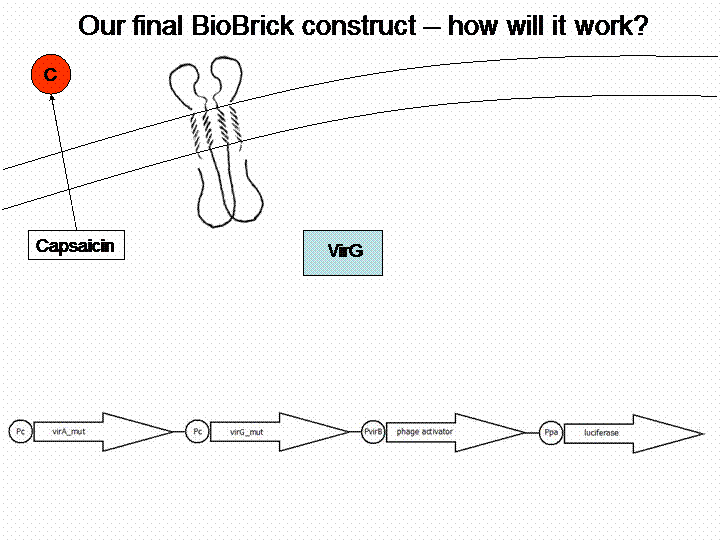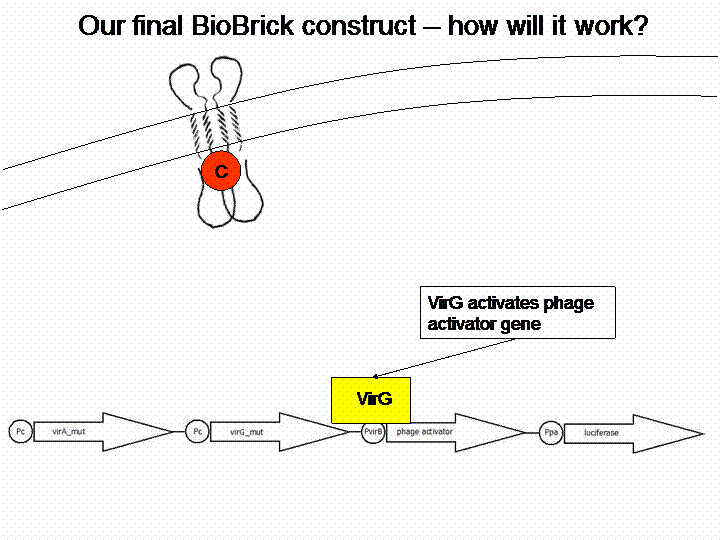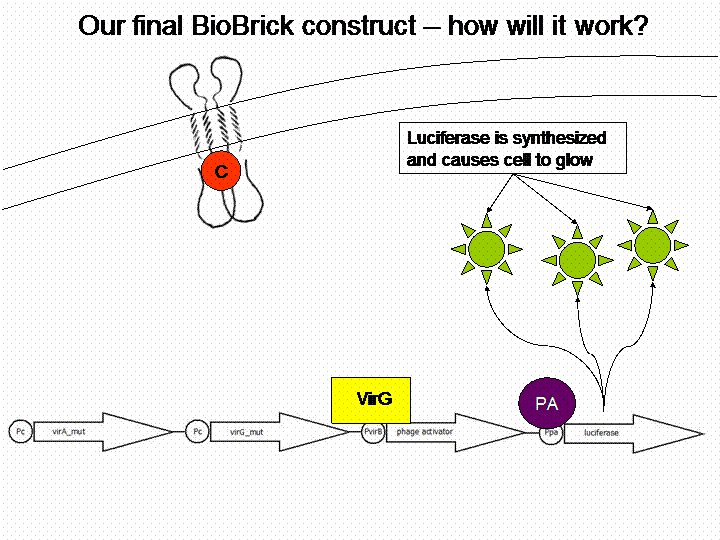
In order to get a visible light signal by the reporter gene, we integrated the quantifiable proteins luciferase and mRFP. We also experimented with certain sensitivity tuners to be able to enhance the detected signal.
For a demonstration of this final system see the animation below:
 "
"