Team:Freiburg Bioware/NoteBook/Labjournal/September2
From 2010.igem.org
(→Mini-Prep and test digestion of ...) |
(→Cloning of CMV promotor into pSB1C3_VP123_inscap) |
||
| Line 831: | Line 831: | ||
<br /> | <br /> | ||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
| - | |||
===<p style="font-size:17px; background-color:#00dd77;">127. labday 22.09.2010</p>=== | ===<p style="font-size:17px; background-color:#00dd77;">127. labday 22.09.2010</p>=== | ||
====<p style="font-size:15px; background-color:#66bbff;"><b>PCR of P40</b></p>==== | ====<p style="font-size:15px; background-color:#66bbff;"><b>PCR of P40</b></p>==== | ||
Revision as of 10:39, 22 September 2010

September
124. labday 19.09.2010
Continuation with Colony-PCR of pSB1C3_CD and ViralBrick motif Z34C
Investigator: Bea
Comment: Since the colony PCR of the last try did not work exactly the way we expected it, another colony PCR approach with new primers was conducted.
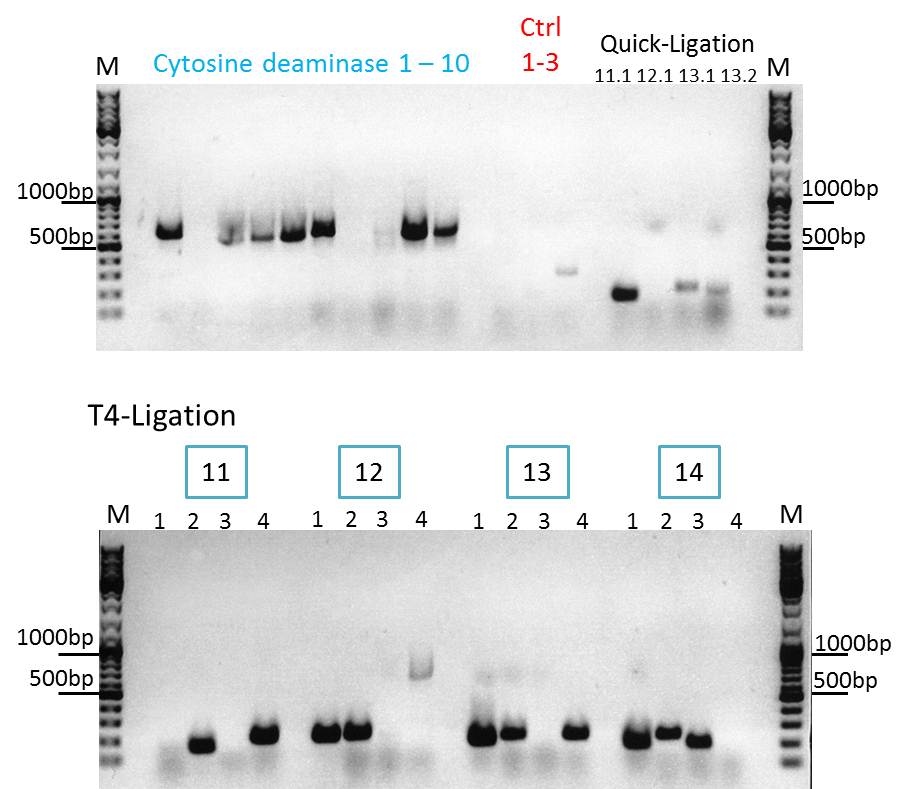
Loading plan:
upper lanes
M CD1 CD2 CD3 CD4 CD5 CD6 CD7 CD8 CD9 CD10 +ctrl(CFP) +ctrl(BAP) -ctrl(pAAV_MCS) 11Q1 12Q1 13Q1 13Q2 M
lower lanes
M 11T41 11T42 11T43 11T44 12T41 12T42 12T43 12T44 13T41 13T42 13T43 13T44 14T41 14T42 14T43 14T44 M
Expected sizes of the PCR products are:
- Cytosine deaminase (CD) = 1300bp
- Z34C loops insertion motif = 220bp
- Positive control 1 = CFP = 750bp
- Positive control 2 = BAP_587 = 170bp
Results:
The Colony PCR of the Cytosine Deaminase (CD) did not work out. The detectable bands at around 700 bp correspond to CFP. Therefore, BioBrick production of the CD must be repeated. The following three bands represent the controls. Three different controls were used. The first corresponds to pSB1C3_CFP, the second to BAP_587 and third control is the negative control. Unfortunately now bands can be detected in the postive controls. A faint band can be seen in the negative control which is the only lane in which no band should be detectable.
The following bands after the three controls correspond to the approaches of the Z34C Viralbrick production. Some bands in the lower and the upper lanes show positive results. The corresponding inoculated overnight culture were centrifuged and prepared in order to perform a Mini-Prep tomorrow.
Comment: hmm, seems that the CD-assambly didn't work at all.. Should i try it again or are we gonna skip it for good due to other priorities?(kira)
No, we cannot skip it. Can we change something else?? Any ideas what we can alter in the protocol? Because problem is that we are still waiting for the tk... so we should aswell focus on the other prodrug activating enzmye!! (Bea)
Testtransduction of the final modified capsid coding constructs
Investigator: Adrian
Aim of the experiment:
During the project 22 silent nucleotide exchange mutations were introduced into the capsid coding construct to make it compatible with the RFC standards and to have single cutting restriction enzymes flanking the 453 and the 587 loop sequence. Two point mutations had to be dismissed, because either a first test transduction showed that the construct was not working anymore or the insertion of the synthesized gene posed serious problems, because the restriction enzyme did not work.
Finally, we have a construct that is shown to produce infectious particles comparable to the current AAV systems and carries 20 point mutations. Now we can announce that the Adeno-associated Virus is compatible to the RFC standard and the idea to replace the loop sequences via ViralBricks works!
Impression from the moment:
Miniprep of serveral constructs
Investigator: Stefan
Glycerol stocks were prepared:
- B430 = pCerulean_ZEGFR:1907_Longlinker_VP2/3
- B431 = pCerulean_CFP_Middlelinker_VP2/3_insCap
- B432 = pCerulean_ZEGFR:1907_Shortlinker_VP2/3_insCap
Mini-Prep was performed according to the standard protocol
- P527 = pCerulean_ZEGFR:1907_Longlinker_VP2/3 c = 230,61 ng/µl
- P528 = pCerulean_CFP_Middlelinker_VP2/3_insCap c = 290,81 ng/µl
- P529 = pCerulean_ZEGFR:1907_Shortlinker_VP2/3_insCap c = 269,39 ng/µl
Several other constructs were preped, but need to be confirmed before being added to plasmid/glycerol stock.
125. labday 20.09.2010
Picking the clones and preparing the over-night culture
Investigator: Kira
VP2/3_Gsat-linker as well as VP2/3_cap_Gsat-linker plates contain colonies, thus 3 clones from each plate were inoculated into 5 ml DYT+ 5ul Chloramphenicol and incubated @ 37 C
Minipreps and Sequencing of ViralBrick clones
Investigator: Achim
The following clones were prepped:
- Quickligation:
- 11.1, c = 85,41 ng/µl
- 13.1, c = 96,93 ng/µl
- T4-Ligation:
- 11.4, c = 76,42 ng/µl
- 12.1, c = 80,36 ng/µl
- 12.4, c = 85,43 ng/µl
- 13.1, c = 90,06 ng/µl
- 13.2, c = 80,87 ng/µl
- 13.4, c = 76,62 ng/µl
- 14.1, c = 76,50 ng/µl
- 14.2, c = 98,52 ng/µl
Clones 11.4, 12.1, 13.2, 14.2 (T4) were sent for sequencing.
Cloning of pCerulean_Affibody_VP2/3 and pSB1C3_Affibody_middlelinker_GFP_his
Investigator: Jessica
Comment:Affibody_VP2/3 will be cloned in pCerulean and Affibody_middlelinker will be cloned together with GFP_His in pSB1C3
- P407= pCerulean_CFP_middlelinker c=482,76 ng/µl
- P516= pSB1C3_ZEGFR:1907_VP2/3 clone 1 c= 191,6 ng/µl
- P290= pSB1C3_Affibody_Middlelinker clone 1 c= 227,4 ng/µl
- P518= pSB1C3_GFP_RFC25_upper band clone 1 c= 138,3 ng/µl
- P520= pSB1C3_GFP_RFC25_lower band clone 1 c= 141,6 ng/µl
Digestion:
| components | P407 | P516 | P290 | P518 | P520 |
| DNA | 3,5 | 10 | 7 | 10 | 10 |
| BSA (10x) | 2 | 2 | 2 | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 | 2 | 2 | 2 |
| EcoRI | 0,8 | 0,8 | - | - | - |
| AgeI | 0,8 | 0,8 | 0,8 | - | - |
| NgoMIV | - | - | - | 0,8 | 0,8 |
| SpeI | - | - | 0,8 | 0,8 | 0,8 |
| H2O | 10,9 | 4,4 | 7,4 | 4,4 | 4,4 |
| Total volume | 20 | 20 | 20 | 20 | 20 |
1,0 g Agarose,100 ml TAE (1%), 6 µl GELRED , at 120 Volt
P516, P518, P520 will be repeated tomorrow (no DNA after gel-ex)

Gelextraction:
The gelextraction was performed according to the standard protocol. DNA concentration of the extracts:
- P407 c= 6,5 ng/µl
- P516 c= - ng/µl
- P290 c= 7,3 ng/µl
- P518 c= - ng/µl
- P520 c= - ng/µl
stands in 4°C room for comtinuation tomorrow
Cloning of P5_TATAless downstream of pSB1C3_001_RC_insrepcap_KpnI_back and P5 upstream of pSB1C3_001_RC_insrepcap_KpnI_back
Investigator: Anissa
Digestion of the constructs:
- P320 = pSB1C3_001_RC_insrepcap_KpnI_back c=408 ng/µL
- P320 = pSB1C3_001_RC_insrepcap_KpnI_back c=408 ng/µL
- P180 = pSB1C3_pTAV2 (P5)clone 1 c=108,8 ng/µL
- P184 = pSB1C3_pAAV_RC (P5TATAless)clone 3 c=176,9 ng/µL
| Components | vP320 upstream /µL | vP180 upstream/µL | vP320 downsteam /µL | vP184 downsteam/µL |
| DNA | 2,5 | 13,8 | 2,5 | 8,5 |
| BSA (10x) | 1,5 | 2 | 1,5 | 1,5 |
| Buffer no. 4 (10x) | 1,5 | 2 | 1,5 | 1,5 |
| Enzyme 1 | XbaI 1 | SpeI 1 | SpeI 1 | XbaI 1 |
| Enzyme 2 | EcoRI 1 | EcoRI 1 | PstI 1 | PstI 1 |
| H2O | 7,5 | 0,2 | 7,5 | 1,5 |
| Total volume | 15 | 20 | 15 | 15 |
Loading plan:
M P320(upstream) P320(downstream) P180 P184
Results:
After gel extraction has been performed, the ligation was carried out.
Ligation:
- v=P320_up =7,37µL
- v=P180_up =0,63µL
- v=P320_down =7µL
- v=P184_down =1µL
The ligation mix was transformed into XL1-Blue cells and plated on agar plates containing chloramphenicol.
Next steps:
Picking clones and perform Mini-Prep.
Test digestion of VP2-N-terminal fusion constructs
Investigator: Hanna
Comment: On Satureday a colony PCR was performed of:
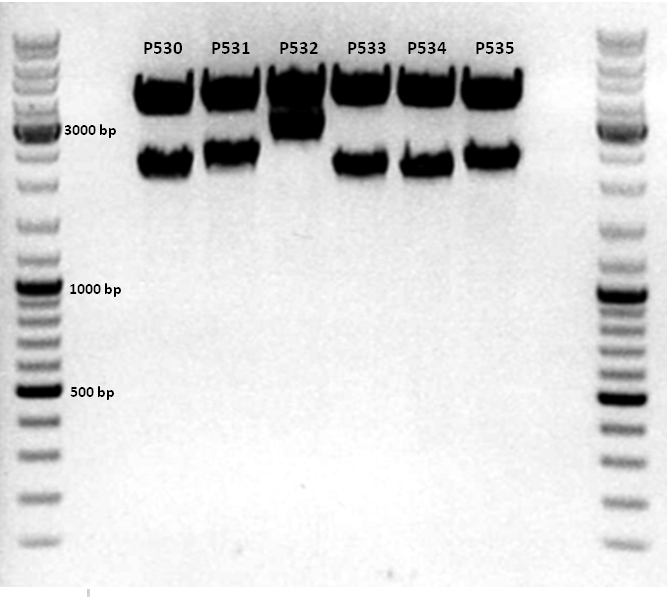
1. pCerulean_Zegfr:1907_ShortLinker_VP2/3 (P530)
2. pCerulean_Zegfr:1907_SEG_VP2/3 (P531)
3. pCerulean_CFP_MiddleLinker_VP2/3 (P532)
4. pCerulean_6xHis_MiddleLinker_VP2/3 (P533)
5. pCerulean_6xHis_MiddleLinker_VP2/3_insCap (P534)
6. pCerulean_Zegfr:1907_MiddleLinker_VP2/3_insCap (P535)
Because all constructs (4 clones of each approach), including the negative control (pAAV_RFC25), showed positive results, one test digestion of each construct was performed.
Digestion
| components | Components Master Mix /µl |
| BSA (10x) | 7 |
| Buffer 4 (10x) | 7 |
| EcoRI | 3.5 |
| SpeI | 3.5 |
| H2O | 35 |
--> 2 µL DNA + 8 µL Master Mix
- Incubation: 1.5 h
Agarose-Gel:
0.4 g Agarose, 50 mL TAE (0.8 %), 3 µL GELRED, at 100 Volt, running time: 50 minutes
| Sample | Sample/µl] | Loading dye (6x)/µl | Expected size 1 | Expected size 2 |
|---|---|---|---|---|
| P 530 | 10 µl | 2 µl | 2710 bp | 3937 bp |
| P 531 | 10 µl | 2 µl | 2710 bp | 3937 bp |
| P 532 | 10 µl | 2 µl | 2710 bp | 3937 bp |
| P 533 | 10 µl | 2 µl | 2710 bp | 3937 bp |
| P 534 | 10 µl | 2 µl | 2710 bp | 3937 bp |
| P 535 | 10 µl | 2 µl | 2710 bp | 3937 bp |
- Marker: GeneRuler ladder mix
| Marker /µL | Sample P530 /µl | Sample P531 /µl | Sample P532 /µl | Sample P533 /µl | Sample P534 /µl | Sample P535 /µl | |
|---|---|---|---|---|---|---|---|
| Lane | 4 | 12 | 12 | 12 | 12 | 12 | 12 |
Comment: Test digestion delivered positive results: Expected fragment size after VP2/3 fusion to Zegfr:1907_Linker: 2180 bp - compared to failed VP2/3 insertion: ~ 220 bp.
Expected fragment size after VP2/3 fusion to CFP_MiddleLinker: 2710 bp - compared to failed VP2/3 insertion: ~ 760 bp.
pCerulean_ZEGFR:1907_Longlinker_VP2/3
pCerulean_CFP_Middlelinker_VP2/3_insCap
pCerulean_ZEGFR:1907_Shortlinker_VP2/3_insCap
pCerulean_ZEGFR:1907_Shortlinker_VP2/3
pCerulean_ZEGFR:1907_SEG_VP2/3
pCerulean_CFP_MiddleLinker_VP2/3
pCerulean_6xHis_MiddleLinker_VP2/3
pCerulean_6xHis_MiddleLinker_VP2/3_insCap
pCerulean_ZEGFR:1907_MiddleLinker_VP2/3_insCap
will be sent for sequencing to GATC.
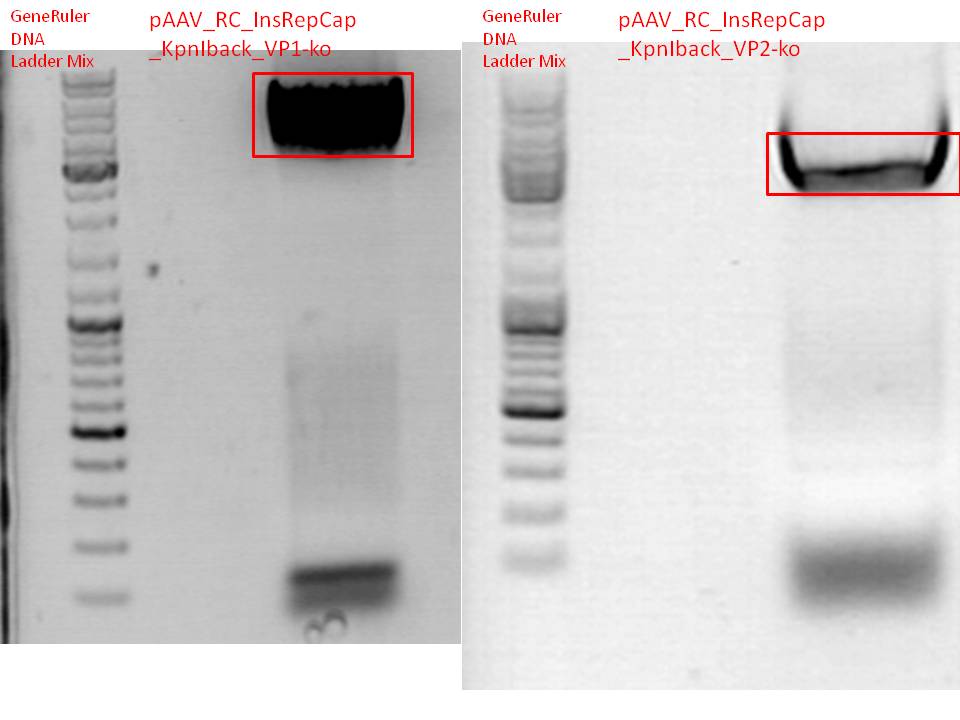
BioBrick production: PCR of pAAV_RC_InsRepCap_KpnIback_VP1-ko and VP2-ko
Investigator: Stefan
DNA samples were diluted 1:1000
- P455 = pAAV_RC_1.2 SDM SalI
- P463 = pAAV_RC_CapIns_prepSDM clone 3
PCR progam:
| Ingredients | Volume P455 / µl | Volume P463 / µl |
| 5X Phusion HF buffer | 10 | 10 |
| 10 mM dNTP mix | 1 | 1 |
| forward primer: O93 | 2,5 | 2,5 |
| reverse primer: O115 | 2,5 | 2,5 |
| DNA Template | 3 | 3 |
| DMSO | 2 | 2 |
| Phusion Polymerase | 0,5 | 0,5 |
| H2O | 28,5 | 28,5 |
| Total volume | 50 | 50 |
PCR program:
| Cycles | Temperature / °C | Time / s |
| 1 | 98 | 60 |
| 2 ( step 2-4: 8x) | 98 | 15 |
| 3 | 59 | 25 |
| 4 | 72 | 75 |
| 5 (step 5-6: 17x) | 98 | 15 |
| 6 | 72 | 85 |
| 7 | 72 | 300 |
| Hold | 4 | 4 |
Gel:
0,5 g agarose (1%), 50 ml TAE, 3µl GELRED running time: 45 minutes at 110V
Gel-Extraction:
Gel-extraction was performed according to standard protocol. The complete amount eluted was used for PCR digestion.
Digestion of PCR product:
| Components | PCR products Volume/µL |
| DNA | 30 |
| BSA (10x) | 4 |
| Buffer no. 4 (10x) | 4 |
| EcoRI | 1 |
| SpeI | 1 |
| H2O | - |
| Total volume | 40 |
Digestion was performed at 37 °C for 2 hours.
PCR-Purification:
Gel-extraction was performed according to standard protocol.
- c (P455) = 13,5 ng/µl
- c (P463) = ng/µl
Digestion of plasmid backbone:
Plasmid used: pSB1C3_VCK_Bla (P320)
c (pSB1C3_VCK_Bla) = 408,0 ng/ µl
| Components | vector Volume/µL |
| DNA | 3 |
| BSA (10x) | 2 |
| Buffer no. 4 (10x) | 2 |
| EcoRI | 1 |
| SpeI | 1 |
| H2O | 11 |
| Total volume | 20 |
Digestion was performed at 37 °C for 2 hours.
Gel:
0,5 g agarose (1%), 50 ml TAE, 3µl GELRED running time: 50 minutes at 120V
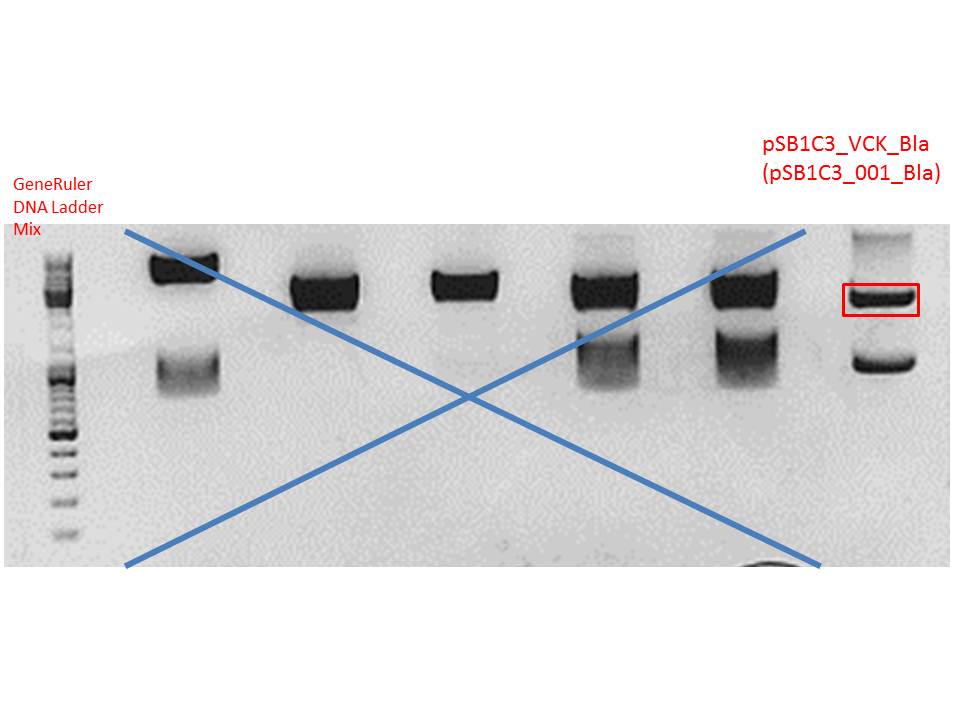
Gel-Extraction:
Gel-extraction was performed according to standard protocol.
c(P320) = 0,8 ng/µl
Ligation:
- 1 µl T4 DNA ligase
- 1 µl 10x Buffer
- 8 µl DNA mix (vector + insert)
Amounts of DNA used:
- c(P320) = 7,73 µl
- c(P455) = 0,27 µl
- c(P320) = 7,19 µl
- c(P463) = 0,81 µl
Incubation time: 45 minutes at room temperature.
Transformation:
Transformation was performed according to standard protocol using XL1b cells and Chloramphenicol as antibiotic.
2x repetition of CD biobrick
Investigator: Kira
| Ingredients | CD sample |
| 5X Phusion HF buffer | 10 µl |
| 10 mM dNTP mix | 1µl |
| forward primer: O158 | 2,5µl |
| reverse primer: O159 | 2,5 µl |
| DNA Template (1:100 dil) | 0,5 µl |
| DMSO | 0 µl |
| Phusion Polymerase | 0,5 µl |
| H2O | 33µl |
| Total volume | 50 µl |
PCR program:
| Cycles | Temperature | Time |
| 98°C | 1 | |
| 10x | 98°C | 15" |
| 58°C | 25" | |
| 72°C | 40" | |
| 17x | 98°C | 15" |
| 65°C | 25" | |
| 72°C | 40" | |
| 1x | 72°C | 5' |
| Hold 4°C |
Digestion of plasmid backbone:
c (pSB1C3) = 151, 1 ng/ µl
| Components | vector Volume/µL |
| DNA 1 µg | 6,0 µl |
| BSA (100x) | 0,2 µl |
| Buffer no. 4 (10x) | 2,0 µl |
| Enzyme 1 XbaI | 0,5 µl |
| Enzyme 2 AgeI HF | 0,5 µl |
| H2O | 10,8 µl |
| Total volume | 20 |
incubation @ 37 C for approx. 6 h
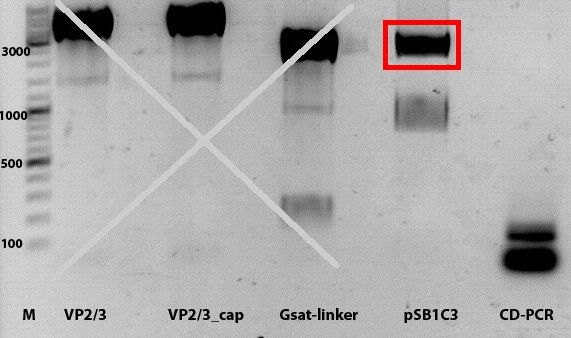
1% agarose gel
According to the gel results, PCR was repeated twice.. But both tries revealed the same unexpected results. Insert from the last cloning procedure was found and used for the ligation and further transformation.
Ligation
T4 ligase was used
1 ul T4 Buffer
1 ul T4 Ligase
8 ul (0,9 ul vector+ 7,1 ul insert) DNA-mix
incubation @ RT for 45 min
Transformation was performed according to the standard protocol and the cells were spread on the agar plate containing Chloramphenicol.
126. labday 21.09.2010
Trafo evalutation of CD biobrick plate
Investigator: Kira
the agar plate was checked at approx. 4 pm and there were no colonies visible. Taking in account that the plate was incubated from 10.30 pm till 8 am @ 37C due to the electricity failure, the plate was put back @ 37C at 4.30 pm
Comment: the plate was picked this morning by a lab member and put in the coldroom
Continuation of cloning of pCerulean_Affibody_VP2/3 and pSB1C3_Affibody_middlelinker_GFP_his
Investigator: Jessica
Comment:Affibody_VP2/3 will be cloned in pCerulean and Affibody_middlelinker will be cloned together with GFP_His in pSB1C3
- P407= pCerulean_CFP_middlelinker c=482,76 ng/µl
- P516= pSB1C3_ZEGFR:1907_VP2/3 clone 1 c= 191,6 ng/µl
- P290= pSB1C3_Affibody_Middlelinker clone 1 c= 227,4 ng/µl
- P518= pSB1C3_GFP_RFC25_upper band clone 1 c= 138,3 ng/µl
- P520= pSB1C3_GFP_RFC25_lower band clone 1 c= 141,6 ng/µl
Digestion:
| components | P516 | P518 | P520 |
| DNA | 10 | 10 | 10 |
| BSA (10x) | 2 | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 | 2 |
| EcoRI | 0,8 | - | - |
| AgeI | 0,8 | - | - |
| MscI | 0,8 | - | - |
| NgoMIV | - | 0,8 | 0,8 |
| SpeI | - | 0,8 | 0,8 |
| H2O | 3,6 | 4,4 | 4,4 |
| Total volume | 20 | 20 | 20 |
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 120 Volt
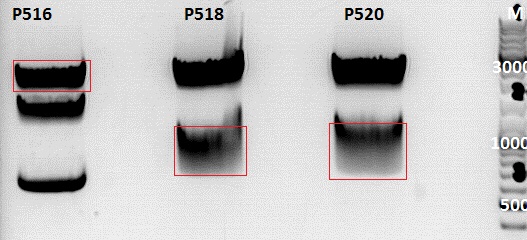
Gelextraction:
The gelextraction was performed according to the standard protocol. DNA concentration of the extracts:
- P516= c= ng/µl
- P518= c= ng/µl
- P520= c= ng/µl
T4 Ligation:
The Ligation was performed as following:
- Vector Volume: µl
- Insert Volume: µl
- 1µl T4-Ligase buffer (10x)
- 8µl (Vector + Insert) mix
- 1µl T4-Ligase
Incubating for 40 minutes.
Transformation:
Trafo was performed according to the standard protocol (XL1blue). The cells were plated on a agar plate with Kana/Cm
Sequenzing result of pSB1C3_EGFP_His upper and (!) lower band
Investigator: Jessica
1. pSB1C3_EGFP_His suffix:
2. pSB1C3_EGFP_His prefix:
results look in both plasmids and are used for cloning of pSB1C3_Affibody_middlelinker_GFP_His
Midi-Prep of pHelper & pSB1C3_leftITR_CMV_beta-globin_mVenus_hGH_rightITR
Investigators: Chris W.
Midi-Preps of pHelper=P540 and pSB1C3_leftITR_CMV_beta-globin_mVenus_hGH_rightITR=P541
The Midi-Preps were performed according to the standard protocol yielding the following concentrations:
| plasmid-no. | P540 | P541 |
| concentration (ng/µl) | 480,45 | 1264,20 |
New aliquots (60µl) of XL1blue can be used ! (Jessica)
127. labday 22.09.2010
PCR of P40
Investigator: Jessica
DNA sample was diluted 1:1000
PCR progam:
| Ingredients | Volume / µl |
| 5X Phusion HF buffer | 10 |
| 10 mM dNTP mix | 1 |
| forward primer: O152 | 2,5 |
| reverse primer: O153 | 2,5 |
| DNA Template | 2,5 |
| DMSO | - |
| Phusion Polymerase | 0,5 |
| H2O | 31 |
| Total volume | 50 |
PCR program:
| Cycles | Temperature / °C | Time / s |
| 98 | 60 | |
| 8x | 98 | 15 |
| 61 | 25 | |
| 72 | 8 | |
| 17x | 98 | 15 |
| 69 | 25 | |
| 72 | 8 | |
| 1x | 72 | 300 |
| Hold | 4 |
Digestion of plasmid backbone:
Plasmid used: pSB1C3_CFP (P51.2)
c (pSB1C3_CFP) = 151, 1 ng/ µl
| Components | vector Volume/µL |
| DNA | 10 |
| BSA (10x) | 2 |
| Buffer no. 4 (10x) | 2 |
| XbaI | 1 |
| PstI | 1 |
| H2O | 4 |
| Total volume | 20 |
incubation @ 37 C for approx. 2 h
Comment:
Mini-Prep and test digestion of pSB1C3_001_RC_InsRepCap_KpnIback_VP1-ko
Investigator: Stefan
 "
"