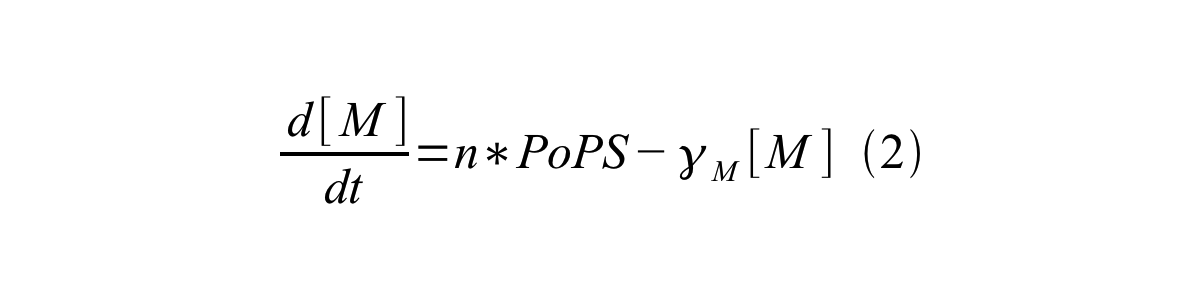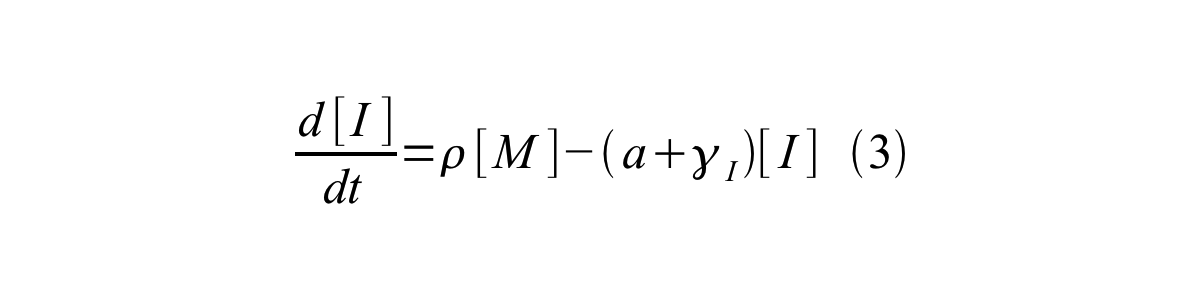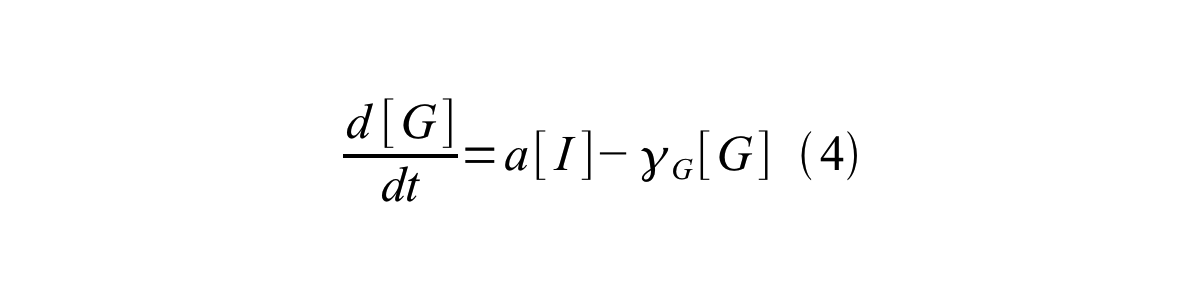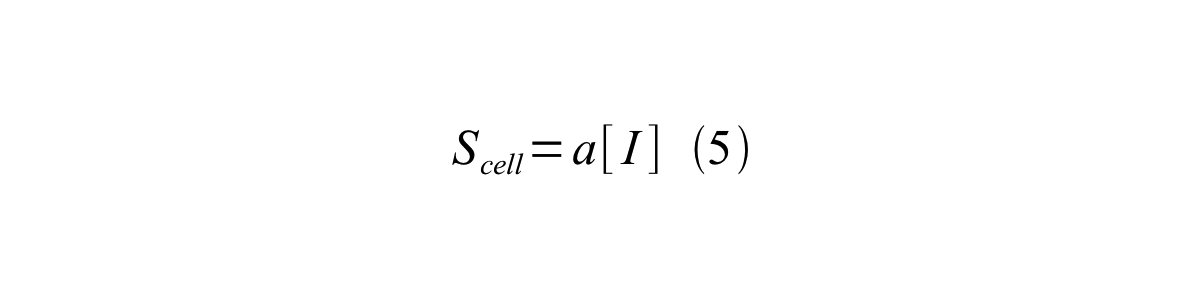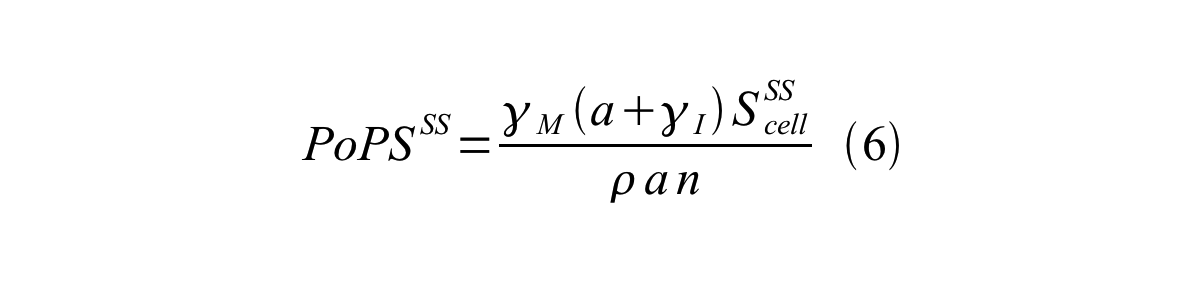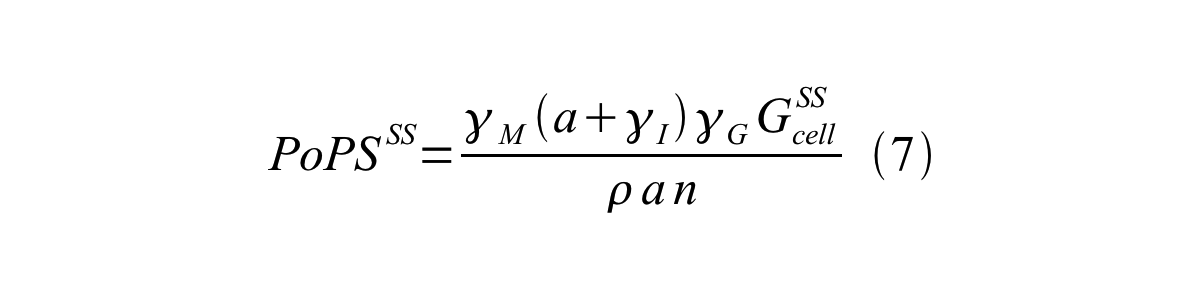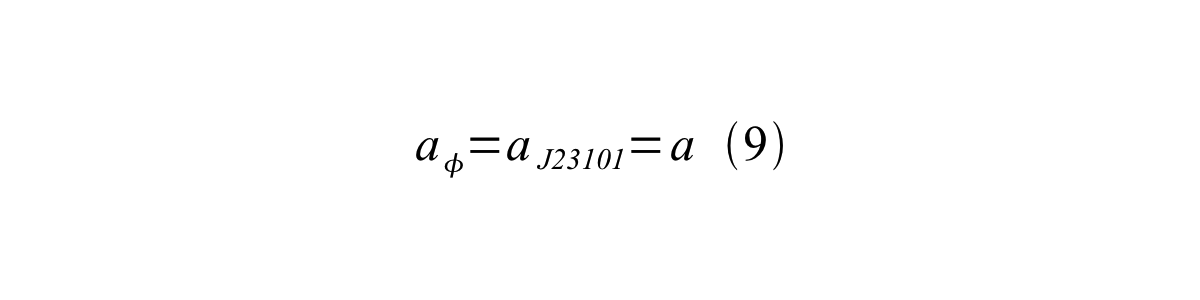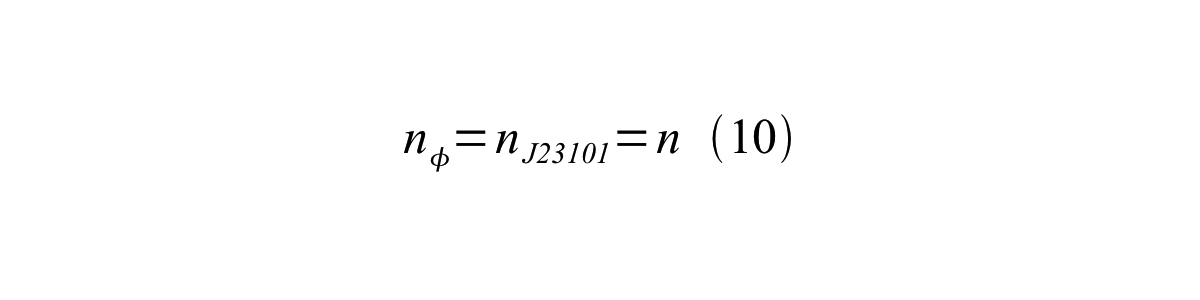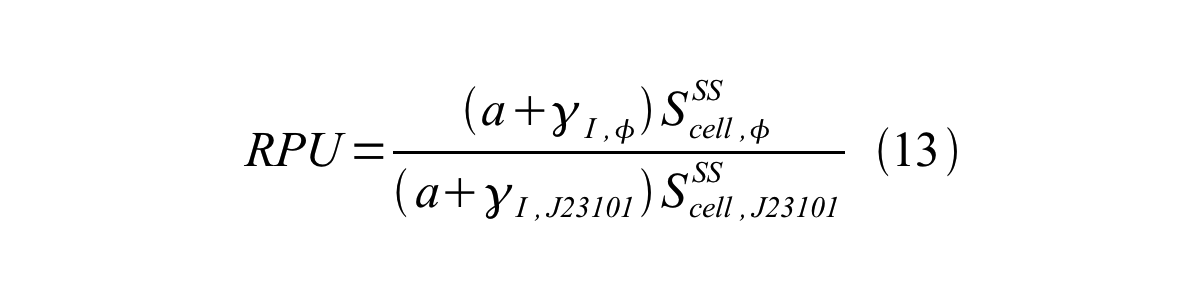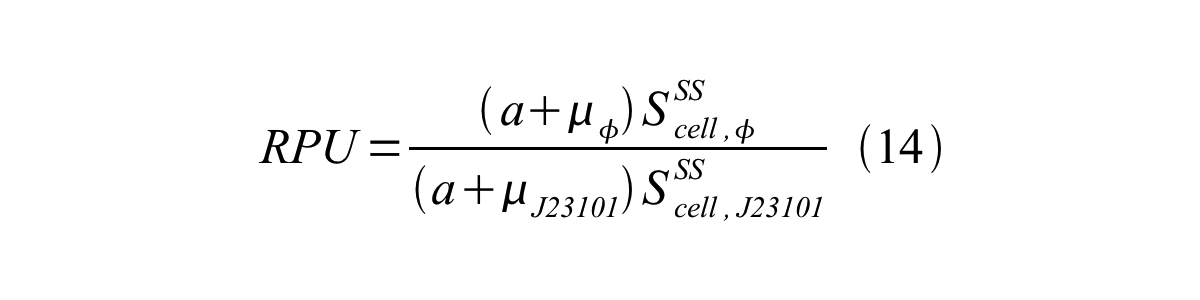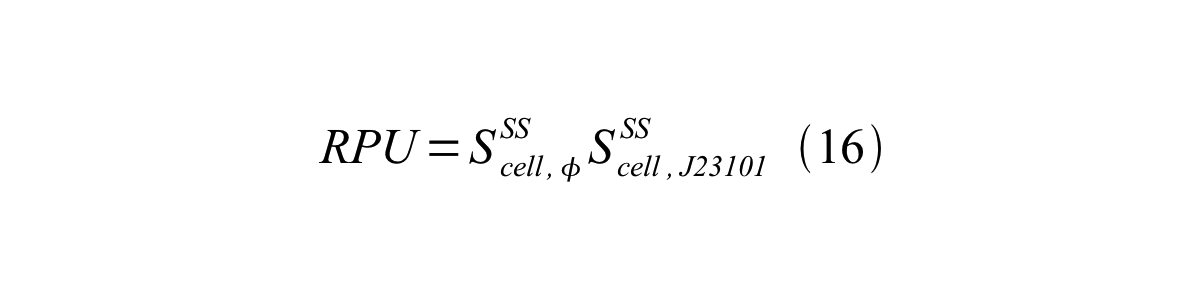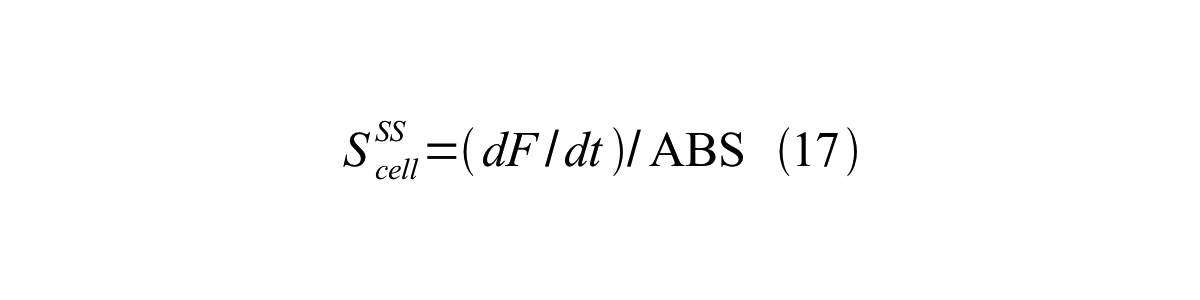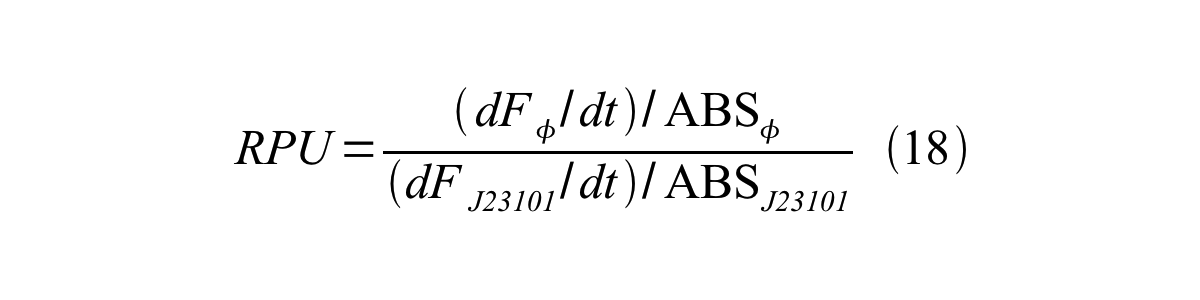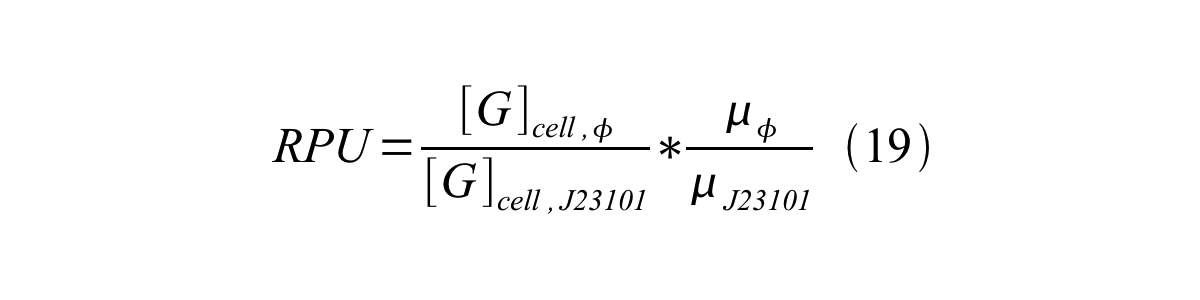Team:Kyoto/LearnMore
From 2010.igem.org
(→R0011) |
|||
| (111 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{:Team:Kyoto/Header}} | {{:Team:Kyoto/Header}} | ||
| - | == | + | ==LearnMore== |
| + | ===R0011=== | ||
| + | In our project, we utilized BioBrick part of BBa_R0011 <sup>[[#RefL001|[1]]]</sup>. This is one of the inducible promoter parts in iGEM, which is often used by a lot of iGEM teams. R0011 is composed of a unique Lac promoter (''lac''P) whose promoter activity is stronger than an ordinary ''lac''P of λ phage. First, we explain the ordinary ''lac''P and subsequently the uniqueness of R0011. | ||
| - | == | + | ====Lac promoter of λ phage==== |
| + | [[Image:KyotoFigR001.png|300px|thumb|Fig.1]] | ||
| - | + | ''lac''P of λ phage is composed of the cI binding sites, Lac operator (LacO) which is bound by Lac inhibitor (LacI) and the promoter at -10 and -35 positions upstream from the transcription start site (Fig.1). | |
| - | + | {{clear}} | |
| + | [[Image:KyotoFigR002.png|300px|thumb|Fig.2]] | ||
| + | This promoter is controlled by the interaction of LacI and lactose as inducer. ''lac''P is classified as a negatively regulated promoter. This means that ''lac''P has the strong promoter activity in the absence of LacI because RNA polymerase can approach the promoter and constitutively transcribe the downstream genes (Fig.2a). However, ''lac''P decreases the activity with increasing LacI. At this time, LacI binds to the promoter and therefore RNA polymerase is prevented from approaching the promoter. As a result, the downstream genes are hardly transcribed (Fig.2b). When lactose is introduced, it binds to LacI and prevents LacI from binding to the promoter. Then RNA polymerase can approach the promoter and transcribes the downstream genes (Fig.2c). | ||
| + | |||
| + | ====The uniqueness of R0011==== | ||
| + | [[Image:KyotoFigR003.png|300px|thumb|Fig.3]] | ||
| + | As stated above, ''lac''P of lambda phage has the cI binding sites and LacO (Figure 1). R0011 replaces cI binding sites with LacO. In other words, this part has two LacO, the original LacO and the extra LacO instead of cI binding sites (Figure 3). Nevertheless, this hybrid design has the strong promoter activity. Moreover, this part is repressed by LacI and induced by lactose as same as ''lac''P of lambda phage. | ||
| + | |||
| + | ====Reference==== | ||
| + | # <html><a name="RefL001"></a></html>Lutz R, Bujard H., [http://www.ncbi.nlm.nih.gov/pubmed/9092630 Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements.], Nucleic Acids Res. 1997 Mar 15;25(6):1203-10. | ||
| + | |||
| + | [[#top-section|^Top]] | ||
| + | |||
| + | ===Lysis Cassette=== | ||
| + | Our iGEM Kyoto team selected the λ phage’s ‘lysis cassette’, including genes S, R and so on, as killer genes (Fig.3). In this chapter, we explain this ‘lysis cassette’. | ||
| + | ====λ phage==== | ||
| + | λ phage is temperate phage that undergoes ‘lysogenic cycle’ and ‘reproductive cycle’ (Fig.1). After infection, the viral genome will integrate with host DNA and replicate along with it fairly harmlessly until host conditions deteriorate. After the deterioration, the endogenous phages (known as prophages) become active. At this point they start the reproductive cycle, resulting in lysis of the host cell. λ phage’s lysis cassette is transcribed in reproductive cycle that causes lysis of the host cell <sup>[[#RefL001|[1]]]</sup>. | ||
| + | |||
| + | <div class="thumbright">[[Image:Λphage.png|thumb|none|440px||Fig. 1: λphage life cycle.<br />λ phage life cycle: λ phage binds to the target E. coli cell, the linear phage genome is injected past the cell membrane(a) and form the DNA circuit (b). At this point, if the host is in good condition, λ phage genome is inserted into host genome (c), λphage enter lysogenic cycle and repeat replication (e), while if the host is in bad condition or host condition deteriorates in lysogenic cycle, λ phage enters reproductive cycle (d), (f). In this cycle, important proteins such as structure proteins and lysis proteins are expressed and λ phages are made (g). Finally, λ phage go out of host by lysis (h). | ||
| + | ]]</div> | ||
| + | |||
| + | ====Lysis system==== | ||
| + | S and R genes, which the lysis cassette includes, perform the important roles of the lysis system. S gene encodes ‘’holin’’ S105 which is a small (Mr 5 8,500) inner membrane (IM) protein and N-out, C-in with three transmembrane domains (TMDs)<sup>[[#RefL002|[2]]] [[#RefL003|[3]]]</sup>(Fig.4a) and R, or endolysin, gene encodes a soluble transglycosylase which degrades ‘peptidoglycan’ <sup>[[#RefL004|[4]]]</sup>.Holin’s function is not understood completely, but assumed as the system below (Fig.2). After ‘’holin’’ S105 is synthesized, holin is instantly inserted into IM. While a few holins are inserted into IM, each holins remains monomers (Fig.2a), and if enough holin is synthesized and inserted into IM, then holin monomers bind each other and form holin rafts, which is stable cluster (Fig.2b). Subsequently, if more holins are inserted, and a holin raft reaches a critical size, then a hole is made within the raft (Fig.2c). As a result, this hole enables the endolysin into the periplasm, and quickly results in cell lysis <sup>[[#RefL009|[9]]]</sup>. | ||
| + | |||
| + | <div class="thumbright"> | ||
| + | [[Image:Hole-formation.png|thumb|none|440px|Fig. 2. Model for hole formation and lysis inhibition by S107.<br /><br />Model for hole formation and lysis inhibition by S107. Pink rectangle is S105, green one is S107. The upper process is a model for hole formation with only holin, the lower process is one when not only holin but also S107 is inserted into IM. (a) Holins are monomers while a few holins are inserted. (b) Holins form rafts if more holins are inserted. (c) Holins form holes if the raft reaches a critical size. If S107 is inserted into IM, it is clear that raft and hole formation is delayed due to S107 antiholin function compared with the formation by only holin (b), (c). However, once a hole is formed, S107 function as holin, S107 also forms holes with holin (d).]]</div> | ||
| + | |||
| + | ====S107 function as antiholin==== | ||
| + | S gene encodes not only S105 but also ‘’antiholin’’ S107 <sup>[[#RefL012|[12]]]</sup> which forms inactive heterodimers with the holin in the inner membrane <sup>[[#RefL007|[7]]][[#RefL008|[8]]]</sup>. | ||
| + | The expression of these two proteins is regulated by the dual-start motif. Because S gene has two start codons, S mRNA produces S105 and S107 which are different in the Met-Lys N-terminal extension of S107 <sup>[[#RefL011|[11]]]</sup> (Fig.3). This dual-start motif is regulated by two stem-loop RNA secondary structures <sup>[[#RefL010|[10]]][[#RefL013|[13]]]</sup>. | ||
| + | S107 also has three TMDs and is expected to become the similar structure to S105, but only two TMDs are inserted into IM, and TMD1 of S107 is localized in the cytoplasm, blocked from insertion into IM. This is because TMD1 of S107 has the positively charged residue, Lys2 and positively charged outside surface of IM repels Lys2 <sup>[[#RefL015|[15]]]</sup>(Fig.4a). Because inserting TMD1 domain of S gene into the IM is essential for holins to form a hole, S107 prevents holins from creating a hole by forming a heterodimer with holin. | ||
| + | Thus S107 functions as antiholin and in addition, changes lysis timing <sup>[[#RefL008|[8]]] [[#RefL009|[9]]] [[#RefL010|[10]]]</sup>. | ||
| + | |||
| + | [[Image:SRRz-gene.2.png|thumb|left|440px|Fig. 3. λgene S and S mRNA’s structure. <br /><br />λ gene S and S mRNA’s structure. The lysis cassette, including genes S, R, Rz, and Rz1, is shown, along with its promoter PR’, and Q, encoding the late gene activator. The 5’ end of holin gene S has two start codons, Met1, the start for S107, and Met3, the start for S105, and two RNA structures that regulate initiations at these codons. ]] | ||
| + | [[Image:S strucure TMD1.png|thumb|440px|Fig. 4. Model for the membrane topology of S105, S107, and SΔTMD1, (a) Before hole formation events occur, (b) after they do. Over membrane is periplasm, under membrane is cytoplasm. SΔTMD1 inhibits S105 all the time, while S107 inhibits S105 only before hole formation, and forms a hole with holin after that.]] | ||
| + | |||
| + | ====SΔTMD1 is a better antiholin than S107==== | ||
| + | S107 functions as antiholin, but we cannot be satisfied with the strength of function of S107 as antiholin. | ||
| + | When an S105-mediated hole formation event does occur in a cell, the resultant collapse of the membrane potential allows insertion of TMD1 of S107 into the membrane (Fig.4b), instantly tripling the amount of active holin by making the previously inactive pool of S105-S107 complexes functional <sup>[[#RefL014|[14]]]</sup>(Fig.2). | ||
| + | That is, S107 function as holin after many S105 form holes. | ||
| + | That is why we cannot be satisfied with the strength of function of S107 as antiholin. | ||
| + | |||
| + | Then, we use SΔTMD1, which is an allele with TMD1deleted and work as antiholin(Fig.4a and 5). Unlike S107, SΔTMD1 does not function as holin because of the deletion of essential TMD1 for hole formation(Fig.4b). | ||
| + | In addition, S gene products form homodimers or heterodimers with each other by binding at TMD2. | ||
| + | So SΔTMD1 can form heterodimers with S105 and prevent holins to form holes <sup>[[#RefL011|[11]]]</sup>. S107 is temporal antiholin but SΔTMD1 is dominant-negative holin. | ||
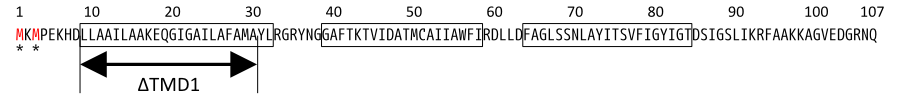
| + | [[Image:S amino sequence.png|center|900px|thumb|Fig.5. Primary structure of S proteins, missense changes relevant to the text are shown. Starts for S107 and S105 are indicated by asterisks. The three TMDs are boxed, and the extent of the TMD1 deletion is indicated.]] | ||
| + | |||
| + | ====Reference==== | ||
| + | # <html><a name="RefL001"></a></html>[http://www.microbiologyprocedure.com/ website Microbiologyprocedure] | ||
| + | # <html><a name="RefL002"></a></html>Altman, E., R. Altman, J. Garrett, R. Grimaila, and R. Young. 1983. S gene product: identification and membrane localization of a lysis control protein.J. Bacteriol. 155:1130–1137. | ||
| + | # <html><a name="RefL003"></a></html>Young, R. 2002. Bacteriophage holins: deadly diversity. J. Mol. Microbiol. Biotechnol. 4:21–36. | ||
| + | # <html><a name="RefL004"></a></html>Bienkowska-Szewczyk, K., B. Lipinska, and A. Taylor. 1981. The R gene product of bacteriophage lambda is the murein transglycosylase. Mol. Gen. Genet. 184:111–114. | ||
| + | # <html><a name="RefL005"></a></html>Young, R. 1992. Bacteriophage lysis: mechanism and regulation. Microbiol. Rev. 56:430–481. | ||
| + | # <html><a name="RefL006"></a></html>Young, R., I. N. Wang, and W. D. Roof. 2000. Phages will out: strategies of host cell lysis. Trends Microbiol. 8:120–128. | ||
| + | # <html><a name="RefL007"></a></html>Gru¨ndling, A., D. L. Smith, U. Bla¨si, and R. Young. 2000. Dimerization between the holin and holin inhibitor of phage _. J. Bacteriol. 182:6075– 6081. | ||
| + | # <html><a name="RefL008"></a></html>Chang, C. Y., K. Nam, and R. Young. 1995. S gene expression and the timing of lysis by bacteriophage _. J. Bacteriol. 177:3283–3294. | ||
| + | # <html><a name="RefL009"></a></html>Gillian L. Ryan and Andrew D. Rutenberg.2007.Clocking Out: Modeling Phage-Induced Lysis of Escherichia coli_. J. Bacteriol. 189: 4749–4755 | ||
| + | # <html><a name="RefL010"></a></html>CHUNG-YU CHANG, KIEBANG NAM, AND RY YOUNG1. 1995.2 S Gene Expression and the Timing of Lysis by Bacteriophage λ. J. Bacteriol. 177: 3283–3294 | ||
| + | # <html><a name="RefL011"></a></html>Rebecca White, Tram Anh T. Tran, Chelsey A. Dankenbring, John Deaton, and Ry Young.2010.The N-Terminal Transmembrane Domain of _ S Is Required for Holin but Not Antiholin Function_. J. Bacteriol. 192: 725–733 | ||
| + | # <html><a name="RefL012"></a></html>Bla¨si, U., C. Y. Chang, M. T. Zagotta, K. Nam, and R. Young. 1990. The lethal lambda S gene encodes its own inhibitor. EMBO J. 9:981–989. | ||
| + | # <html><a name="RefL013"></a></html>Raab, R., G. Neal, C. Sohaskey, J. Smith, and R. Young. 1988. Dominance in lambda S mutations and evidence for translational control. J. Mol. Biol. 199:95–105. | ||
| + | # <html><a name="RefL014"></a></html>Young, R. 2002. Bacteriophage holins: deadly diversity. J. Mol. Microbiol. Biotechnol. 4:21–36. | ||
| + | # <html><a name="RefL015"></a></html>Udo Blasi and Ry Young. 1996. Two beginnings for a single purpose: the dual-start holins in the regulation of phage lysis. Mol Microbiol. 21(4), 675–682 | ||
| + | |||
| + | [[#top-section|^Top]] | ||
| + | |||
| + | ===Relative Promoter Unit (RPU)=== | ||
| + | ====Overview==== | ||
| + | Though BioBrick parts are standardized in terms of how individual parts are physically assembled into multi-component systems, many of them are not standardized in the point of characterization. | ||
| + | |||
| + | Unfortunately, it is known that the absolute activity of BioBrick promoters varies across experimental conditions and measurement instruments <sup>[[#RefP001|[1]]]</sup>. However, Kelly et al. found that this variation could be reduced by measuring the activity of promoters relative to <partinfo>J23101</partinfo> which is a promoter as an in vivo reference standard for promoter activity by ~50%. They defined a Relative Promoter Units (RPU) in order to report promoter characterization data in compatible units. In order to measure RPUs of promoters, the synthesis rate of Green Fluorescent Protein (GFP) encoded by mRNA transcribed from each promoter is observed. | ||
| + | |||
| + | RPU is defined by equation 1: | ||
| + | [[Image:KyotoPromoterActivity001.png|600px|center]] | ||
| + | |||
| + | ====Derivation==== | ||
| + | PoPS (Polymerase Per Second) is the unit of absolute “promoter activity”. It is defined as the number of RNA polymerase molecules that pass by the final base pair of the promoter and continue along DNA as an elongation complex. PoPS<sup>SS</sup> is PoPS at the steady state of following system (d[M]/dt=0, d[I]/dt=0, d[G]/dt=0: | ||
| + | [[Image:KyotoPromoterActivity002.png|600px|center]] | ||
| + | [[Image:KyotoPromoterActivity003.png|600px|center]] | ||
| + | [[Image:KyotoPromoterActivity004.png|600px|center]] | ||
| + | |||
| + | Where: | ||
| + | *[M] is the concentration of mRNA, | ||
| + | *[I] is the concentration of immature GFP, | ||
| + | *[G] is the concentration of mature GFP, | ||
| + | *γ<sub>M</sub> is the mRNA degradation rate, | ||
| + | *a is the GFP maturation rate, | ||
| + | *γ<sub>I</sub> is the degradation rate of immature GFP | ||
| + | *n is the copy number of the plasmid containing the promoter | ||
| + | *ρ is the rate of synthesis of immature GFP in absolute units of protein per second per mRNA. | ||
| + | |||
| + | By combining equation 2, 3, 4 and 5 at the steady state, equation 6 and 7 are obtained: | ||
| + | [[Image:KyotoPromoterActivity005.png|600px|center]] | ||
| + | |||
| + | Here Scell is define as the per cell mature GFP synthesis term. | ||
| + | [[Image:KyotoPromoterActivity006.png|600px|center]] | ||
| + | [[Image:KyotoPromoterActivity007.png|600px|center]] | ||
| + | |||
| + | Here G<sub>cell</sub> is define as the per cell mature GFP synthesis term. | ||
| + | |||
| + | From equation 1 and 6 RPU is described as equation8. | ||
| + | |||
| + | [[Image:KyotoPromoterActivity008.png|600px|center]] | ||
| + | |||
| + | If the test promoter φ and the reference standard promoter are measured under the same culture conditions and both promoters are carried on the same backbone plasmid, following equations are approved. | ||
| + | [[Image:KyotoPromoterActivity009.png|600px|center]] | ||
| + | [[Image:KyotoPromoterActivity010.png|600px|center]] | ||
| + | [[Image:KyotoPromoterActivity011.png|600px|center]] | ||
| + | [[Image:KyotoPromoterActivity012.png|600px|center]] | ||
| + | |||
| + | From equation 8, 9, 10, 11, and 12, equation 13 is accomplished. | ||
| + | [[Image:KyotoPromoterActivity013.png|600px|center]] | ||
| + | |||
| + | In addition, immature GFP is stable so that protein degradation is negligible compared to dilution due to cell growth (γ<sub>I</sub>, φ=μ<sub>φ</sub> and γ<sub>I,,J23101</sub>=μ<sub>J23101</sub>, where μis the cellular growth rate). Therefore, equation 13 can be modified to equation 14: | ||
| + | [[Image:KyotoPromoterActivity014.png|600px|center]] | ||
| + | |||
| + | And if the growth rates of both strains are almost same, equation 15 is approved. | ||
| + | [[Image:KyotoPromoterActivity015.png|600px|center]] | ||
| + | |||
| + | Accordingly, we can make equation 16 from equation15. | ||
| + | [[Image:KyotoPromoterActivity016.png|600px|center]] | ||
| + | |||
| + | S<sub>cell</sub><sup>SS</sup> can be described by equation17: | ||
| + | [[Image:KyotoPromoterActivity017.png|600px|center]] | ||
| + | |||
| + | Therefore, RPU is described by equation 18: | ||
| + | [[Image:KyotoPromoterActivity018.png|600px|center]] | ||
| + | |||
| + | We can also establish equation20 from equation1, 6, 9, 10, 11 and 12. | ||
| + | [[Image:KyotoPromoterActivity019.png|600px|center]] | ||
| + | |||
| + | RPU can be calculated from equation 18 or equation 19. Therefore, we have to observe GFP synthesis rate and ABS (OD600) of both strains or to measure GFP concentration per cell and growth rate of both strains to measure RPU. | ||
| + | |||
| + | ====Reference==== | ||
| + | # <html><a name="RefP001"></a></html>[http://www.ncbi.nlm.nih.gov/pubmed/19298678 PMID: 19298678] Kelly JR, Rubin AJ, Davis JH, Ajo-Franklin CM, Cumbers J, Czar MJ, de Mora K, Glieberman AL, Monie DD, Endy D., ''Measuring the activity of BioBrick promoters using an in vivo reference standard.'', J Biol Eng. 2009 Mar 20;3:4. | ||
| + | [[#top-section|^Top]] | ||
---- | ---- | ||
Latest revision as of 20:45, 27 October 2010
Contents |
LearnMore
R0011
In our project, we utilized BioBrick part of BBa_R0011 [1]. This is one of the inducible promoter parts in iGEM, which is often used by a lot of iGEM teams. R0011 is composed of a unique Lac promoter (lacP) whose promoter activity is stronger than an ordinary lacP of λ phage. First, we explain the ordinary lacP and subsequently the uniqueness of R0011.
Lac promoter of λ phage
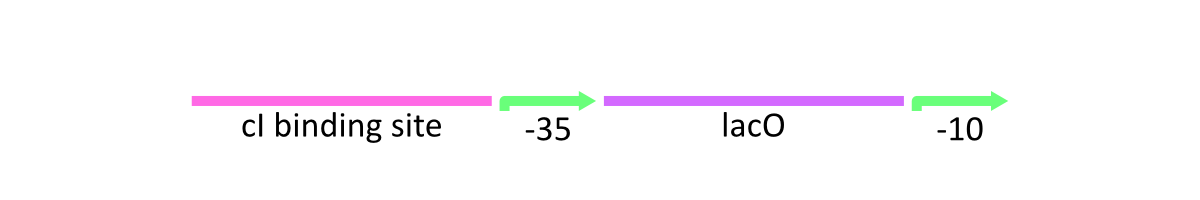
lacP of λ phage is composed of the cI binding sites, Lac operator (LacO) which is bound by Lac inhibitor (LacI) and the promoter at -10 and -35 positions upstream from the transcription start site (Fig.1).
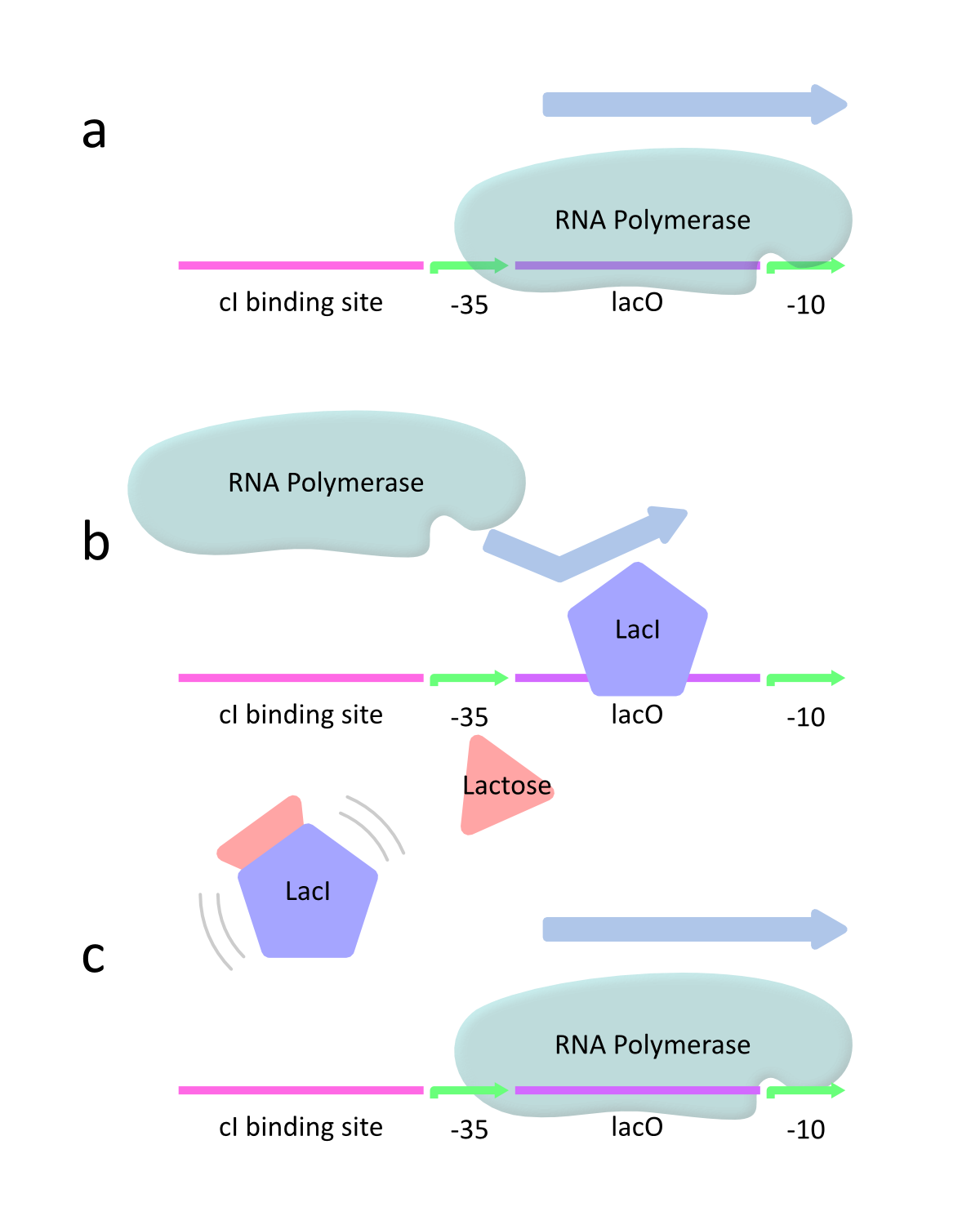
This promoter is controlled by the interaction of LacI and lactose as inducer. lacP is classified as a negatively regulated promoter. This means that lacP has the strong promoter activity in the absence of LacI because RNA polymerase can approach the promoter and constitutively transcribe the downstream genes (Fig.2a). However, lacP decreases the activity with increasing LacI. At this time, LacI binds to the promoter and therefore RNA polymerase is prevented from approaching the promoter. As a result, the downstream genes are hardly transcribed (Fig.2b). When lactose is introduced, it binds to LacI and prevents LacI from binding to the promoter. Then RNA polymerase can approach the promoter and transcribes the downstream genes (Fig.2c).
The uniqueness of R0011
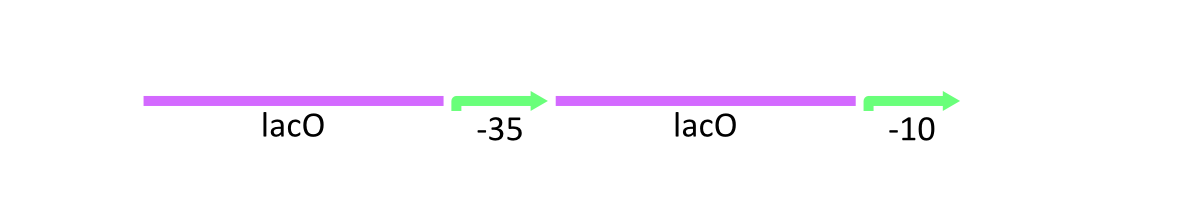
As stated above, lacP of lambda phage has the cI binding sites and LacO (Figure 1). R0011 replaces cI binding sites with LacO. In other words, this part has two LacO, the original LacO and the extra LacO instead of cI binding sites (Figure 3). Nevertheless, this hybrid design has the strong promoter activity. Moreover, this part is repressed by LacI and induced by lactose as same as lacP of lambda phage.
Reference
- Lutz R, Bujard H., [http://www.ncbi.nlm.nih.gov/pubmed/9092630 Independent and tight regulation of transcriptional units in Escherichia coli via the LacR/O, the TetR/O and AraC/I1-I2 regulatory elements.], Nucleic Acids Res. 1997 Mar 15;25(6):1203-10.
Lysis Cassette
Our iGEM Kyoto team selected the λ phage’s ‘lysis cassette’, including genes S, R and so on, as killer genes (Fig.3). In this chapter, we explain this ‘lysis cassette’.
λ phage
λ phage is temperate phage that undergoes ‘lysogenic cycle’ and ‘reproductive cycle’ (Fig.1). After infection, the viral genome will integrate with host DNA and replicate along with it fairly harmlessly until host conditions deteriorate. After the deterioration, the endogenous phages (known as prophages) become active. At this point they start the reproductive cycle, resulting in lysis of the host cell. λ phage’s lysis cassette is transcribed in reproductive cycle that causes lysis of the host cell [1].

λ phage life cycle: λ phage binds to the target E. coli cell, the linear phage genome is injected past the cell membrane(a) and form the DNA circuit (b). At this point, if the host is in good condition, λ phage genome is inserted into host genome (c), λphage enter lysogenic cycle and repeat replication (e), while if the host is in bad condition or host condition deteriorates in lysogenic cycle, λ phage enters reproductive cycle (d), (f). In this cycle, important proteins such as structure proteins and lysis proteins are expressed and λ phages are made (g). Finally, λ phage go out of host by lysis (h).
Lysis system
S and R genes, which the lysis cassette includes, perform the important roles of the lysis system. S gene encodes ‘’holin’’ S105 which is a small (Mr 5 8,500) inner membrane (IM) protein and N-out, C-in with three transmembrane domains (TMDs)[2] [3](Fig.4a) and R, or endolysin, gene encodes a soluble transglycosylase which degrades ‘peptidoglycan’ [4].Holin’s function is not understood completely, but assumed as the system below (Fig.2). After ‘’holin’’ S105 is synthesized, holin is instantly inserted into IM. While a few holins are inserted into IM, each holins remains monomers (Fig.2a), and if enough holin is synthesized and inserted into IM, then holin monomers bind each other and form holin rafts, which is stable cluster (Fig.2b). Subsequently, if more holins are inserted, and a holin raft reaches a critical size, then a hole is made within the raft (Fig.2c). As a result, this hole enables the endolysin into the periplasm, and quickly results in cell lysis [9].

Model for hole formation and lysis inhibition by S107. Pink rectangle is S105, green one is S107. The upper process is a model for hole formation with only holin, the lower process is one when not only holin but also S107 is inserted into IM. (a) Holins are monomers while a few holins are inserted. (b) Holins form rafts if more holins are inserted. (c) Holins form holes if the raft reaches a critical size. If S107 is inserted into IM, it is clear that raft and hole formation is delayed due to S107 antiholin function compared with the formation by only holin (b), (c). However, once a hole is formed, S107 function as holin, S107 also forms holes with holin (d).
S107 function as antiholin
S gene encodes not only S105 but also ‘’antiholin’’ S107 [12] which forms inactive heterodimers with the holin in the inner membrane [7][8]. The expression of these two proteins is regulated by the dual-start motif. Because S gene has two start codons, S mRNA produces S105 and S107 which are different in the Met-Lys N-terminal extension of S107 [11] (Fig.3). This dual-start motif is regulated by two stem-loop RNA secondary structures [10][13]. S107 also has three TMDs and is expected to become the similar structure to S105, but only two TMDs are inserted into IM, and TMD1 of S107 is localized in the cytoplasm, blocked from insertion into IM. This is because TMD1 of S107 has the positively charged residue, Lys2 and positively charged outside surface of IM repels Lys2 [15](Fig.4a). Because inserting TMD1 domain of S gene into the IM is essential for holins to form a hole, S107 prevents holins from creating a hole by forming a heterodimer with holin. Thus S107 functions as antiholin and in addition, changes lysis timing [8] [9] [10].

λ gene S and S mRNA’s structure. The lysis cassette, including genes S, R, Rz, and Rz1, is shown, along with its promoter PR’, and Q, encoding the late gene activator. The 5’ end of holin gene S has two start codons, Met1, the start for S107, and Met3, the start for S105, and two RNA structures that regulate initiations at these codons.
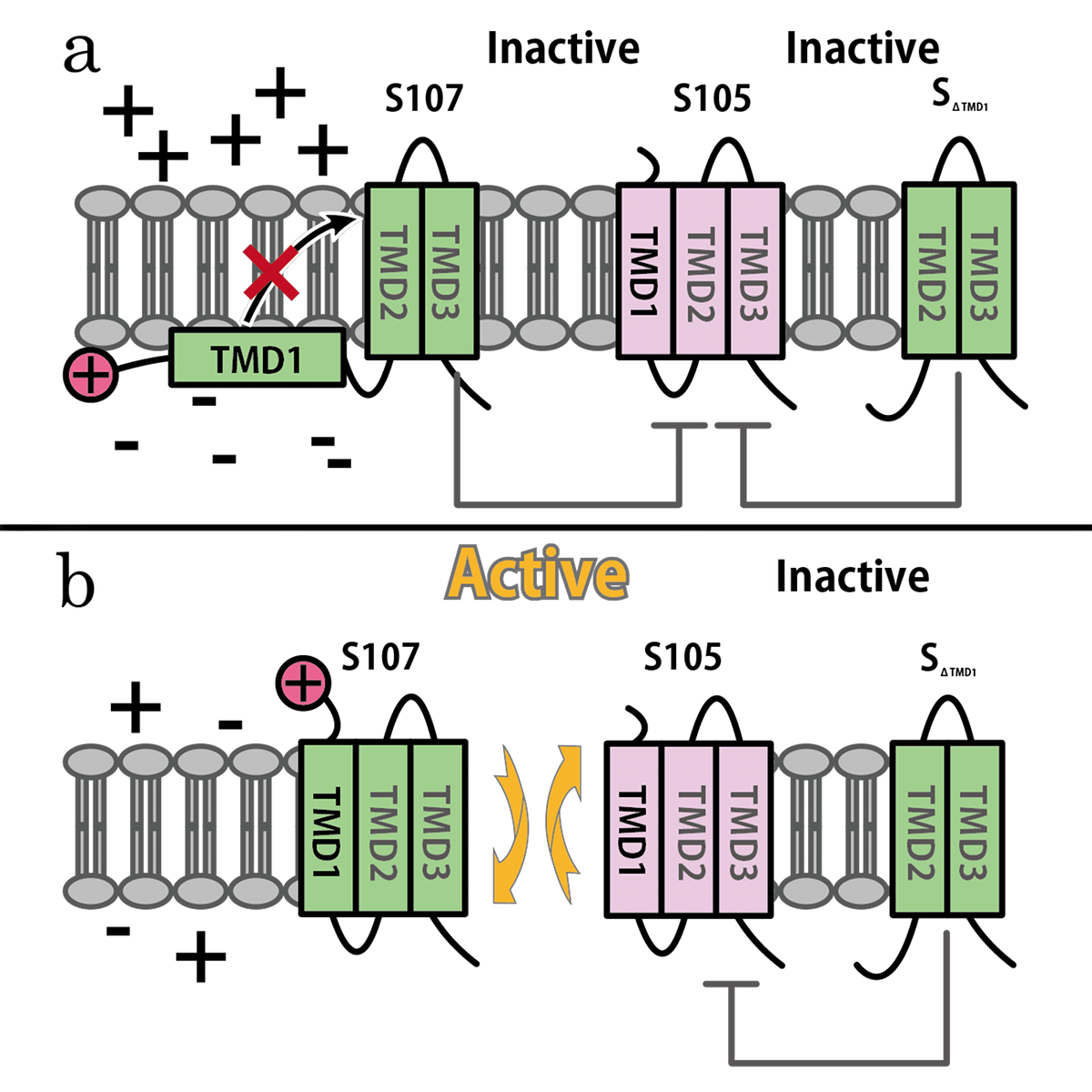
SΔTMD1 is a better antiholin than S107
S107 functions as antiholin, but we cannot be satisfied with the strength of function of S107 as antiholin. When an S105-mediated hole formation event does occur in a cell, the resultant collapse of the membrane potential allows insertion of TMD1 of S107 into the membrane (Fig.4b), instantly tripling the amount of active holin by making the previously inactive pool of S105-S107 complexes functional [14](Fig.2). That is, S107 function as holin after many S105 form holes. That is why we cannot be satisfied with the strength of function of S107 as antiholin.
Then, we use SΔTMD1, which is an allele with TMD1deleted and work as antiholin(Fig.4a and 5). Unlike S107, SΔTMD1 does not function as holin because of the deletion of essential TMD1 for hole formation(Fig.4b). In addition, S gene products form homodimers or heterodimers with each other by binding at TMD2. So SΔTMD1 can form heterodimers with S105 and prevent holins to form holes [11]. S107 is temporal antiholin but SΔTMD1 is dominant-negative holin.
Reference
- [http://www.microbiologyprocedure.com/ website Microbiologyprocedure]
- Altman, E., R. Altman, J. Garrett, R. Grimaila, and R. Young. 1983. S gene product: identification and membrane localization of a lysis control protein.J. Bacteriol. 155:1130–1137.
- Young, R. 2002. Bacteriophage holins: deadly diversity. J. Mol. Microbiol. Biotechnol. 4:21–36.
- Bienkowska-Szewczyk, K., B. Lipinska, and A. Taylor. 1981. The R gene product of bacteriophage lambda is the murein transglycosylase. Mol. Gen. Genet. 184:111–114.
- Young, R. 1992. Bacteriophage lysis: mechanism and regulation. Microbiol. Rev. 56:430–481.
- Young, R., I. N. Wang, and W. D. Roof. 2000. Phages will out: strategies of host cell lysis. Trends Microbiol. 8:120–128.
- Gru¨ndling, A., D. L. Smith, U. Bla¨si, and R. Young. 2000. Dimerization between the holin and holin inhibitor of phage _. J. Bacteriol. 182:6075– 6081.
- Chang, C. Y., K. Nam, and R. Young. 1995. S gene expression and the timing of lysis by bacteriophage _. J. Bacteriol. 177:3283–3294.
- Gillian L. Ryan and Andrew D. Rutenberg.2007.Clocking Out: Modeling Phage-Induced Lysis of Escherichia coli_. J. Bacteriol. 189: 4749–4755
- CHUNG-YU CHANG, KIEBANG NAM, AND RY YOUNG1. 1995.2 S Gene Expression and the Timing of Lysis by Bacteriophage λ. J. Bacteriol. 177: 3283–3294
- Rebecca White, Tram Anh T. Tran, Chelsey A. Dankenbring, John Deaton, and Ry Young.2010.The N-Terminal Transmembrane Domain of _ S Is Required for Holin but Not Antiholin Function_. J. Bacteriol. 192: 725–733
- Bla¨si, U., C. Y. Chang, M. T. Zagotta, K. Nam, and R. Young. 1990. The lethal lambda S gene encodes its own inhibitor. EMBO J. 9:981–989.
- Raab, R., G. Neal, C. Sohaskey, J. Smith, and R. Young. 1988. Dominance in lambda S mutations and evidence for translational control. J. Mol. Biol. 199:95–105.
- Young, R. 2002. Bacteriophage holins: deadly diversity. J. Mol. Microbiol. Biotechnol. 4:21–36.
- Udo Blasi and Ry Young. 1996. Two beginnings for a single purpose: the dual-start holins in the regulation of phage lysis. Mol Microbiol. 21(4), 675–682
Relative Promoter Unit (RPU)
Overview
Though BioBrick parts are standardized in terms of how individual parts are physically assembled into multi-component systems, many of them are not standardized in the point of characterization.
Unfortunately, it is known that the absolute activity of BioBrick promoters varies across experimental conditions and measurement instruments [1]. However, Kelly et al. found that this variation could be reduced by measuring the activity of promoters relative to <partinfo>J23101</partinfo> which is a promoter as an in vivo reference standard for promoter activity by ~50%. They defined a Relative Promoter Units (RPU) in order to report promoter characterization data in compatible units. In order to measure RPUs of promoters, the synthesis rate of Green Fluorescent Protein (GFP) encoded by mRNA transcribed from each promoter is observed.
RPU is defined by equation 1:
Derivation
PoPS (Polymerase Per Second) is the unit of absolute “promoter activity”. It is defined as the number of RNA polymerase molecules that pass by the final base pair of the promoter and continue along DNA as an elongation complex. PoPSSS is PoPS at the steady state of following system (d[M]/dt=0, d[I]/dt=0, d[G]/dt=0:
Where:
- [M] is the concentration of mRNA,
- [I] is the concentration of immature GFP,
- [G] is the concentration of mature GFP,
- γM is the mRNA degradation rate,
- a is the GFP maturation rate,
- γI is the degradation rate of immature GFP
- n is the copy number of the plasmid containing the promoter
- ρ is the rate of synthesis of immature GFP in absolute units of protein per second per mRNA.
By combining equation 2, 3, 4 and 5 at the steady state, equation 6 and 7 are obtained:
Here Scell is define as the per cell mature GFP synthesis term.
Here Gcell is define as the per cell mature GFP synthesis term.
From equation 1 and 6 RPU is described as equation8.
If the test promoter φ and the reference standard promoter are measured under the same culture conditions and both promoters are carried on the same backbone plasmid, following equations are approved.
From equation 8, 9, 10, 11, and 12, equation 13 is accomplished.
In addition, immature GFP is stable so that protein degradation is negligible compared to dilution due to cell growth (γI, φ=μφ and γI,,J23101=μJ23101, where μis the cellular growth rate). Therefore, equation 13 can be modified to equation 14:
And if the growth rates of both strains are almost same, equation 15 is approved.
Accordingly, we can make equation 16 from equation15.
ScellSS can be described by equation17:
Therefore, RPU is described by equation 18:
We can also establish equation20 from equation1, 6, 9, 10, 11 and 12.
RPU can be calculated from equation 18 or equation 19. Therefore, we have to observe GFP synthesis rate and ABS (OD600) of both strains or to measure GFP concentration per cell and growth rate of both strains to measure RPU.
Reference
- [http://www.ncbi.nlm.nih.gov/pubmed/19298678 PMID: 19298678] Kelly JR, Rubin AJ, Davis JH, Ajo-Franklin CM, Cumbers J, Czar MJ, de Mora K, Glieberman AL, Monie DD, Endy D., Measuring the activity of BioBrick promoters using an in vivo reference standard., J Biol Eng. 2009 Mar 20;3:4.
 "
"