Biochemical Modelling
Motivation
In order to create a flow with bacteria using their flagella movement as the motor, it is important to understand bacterial motility and the effects our genetic manipulation have on it.
Bacterial motion resembles a random walk, with periods of smooth swimming interrupted by brief tumbles that change the swimming direction. Chemotaxis is achieved by modulation of the tumbling frequency.
In our case, we have added phototaxis to E. coli MG1655 cells making them react to light as well as chemical gradients.
By creating a model describing the key features of the system, it will be possible to study the biochemical behaviour of such a system, and how different parameters will affect it, before it is implemented in a bacterial strain.
The real system
Several chemosensory systems exist in bacteria. The one most studied is the chemotaxis of E. coli. Polar clusters of membrane-spanning methyl accepting chemotaxis proteins (MCPs) are positioned in the ends of the rod shaped E. coli.
Tar (responding to aspartate), Tsr (responding to serine), Tap (responding todipetides), Trg (responding to galactose) ang Aer (responding to oxygen) are the most common of these receptors seen in E. coli strains[1]. Transduction of the signal between the MCPs and the flagella motor is managed by a particular fine-tuned signalling cascade, which is governed by several intracellular proteins.
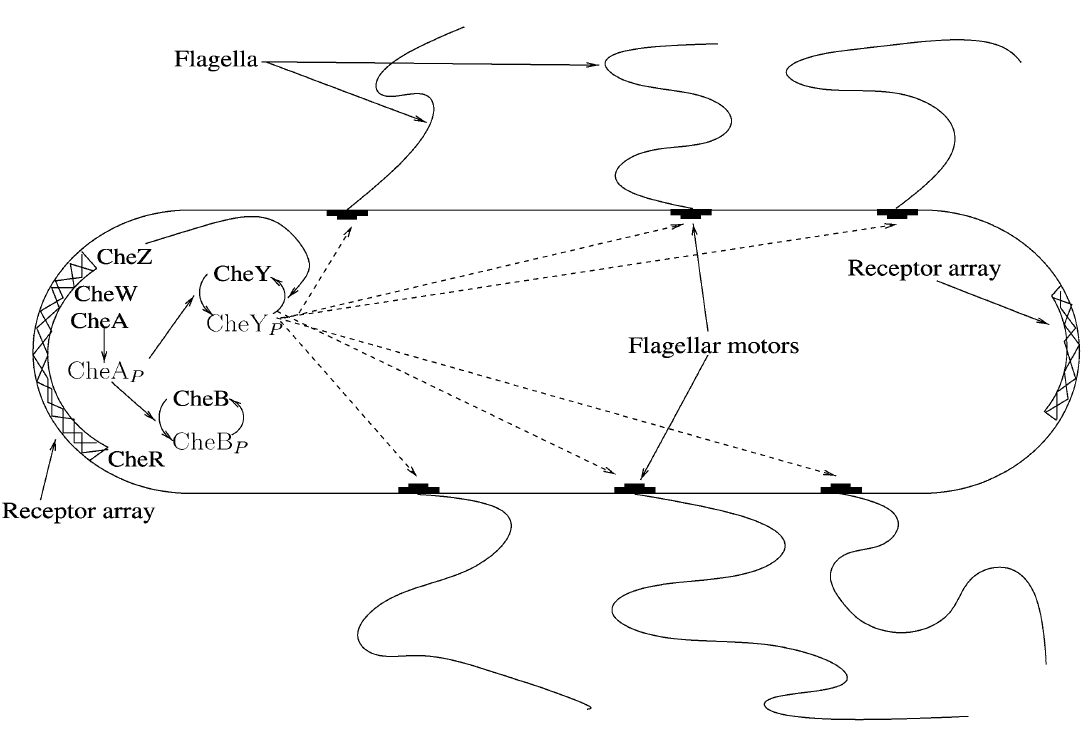
Figure 1: Schematics of the bacterial chemotaxis pathways
Chemotaxis consists of three important phases: reaction, adaptation and relaxation. In the reaction phase, one of the MCPs sense the specific chemical gradient it responds to and the membrane protein activates the CheW enzyme. The active CheW enzyme suppresses the auto-phosphorylation of the CheA enzyme and when no or very little CheA enzyme is phosphorylated, no phosphorylation of the CheY and CheB enzymes occurs. In the end, this increases run, because phosphorylated CheY binds to the flagella motors and induce tumbling. When the bacteria have reached the area with a higher concentration of the chemical in question adaptation sets in.
Because no CheB have been phosphorylated during the reaction period CheR is “allowed” to methylate the MCPs, which increases the auto-phosphorylation of the CheA enzyme. This increase in auto-phosphorylation of CheA results in phosphorylation of CheY that leads to tumbling of the bacteria. In the last phase, relaxation, CheB has again been phosphorylated by CheA and is now able to demethylate the MCPs, returning the bacteria to its normal run/tumble frequency with a 0,1 second tumbling period every 1 second.
One other thing to consider in the model is the distance from the CheY phosphorylation site to the flagella motors that are randomly located on the entire cell body. Several modelling methods have been proposed and nearly all of them include these considerations:
"Attractant is assumed to be in excess and binding is assumed to be rapid". One of the most common receptor types to be modelled is the Tar receptor. This is not unexpected given that a large degree of the experimental literature has focused on the response of E. coli to aspartate which is detected by this receptor. Tar is also one of the most abundant receptors within the cytoplasmic membrane.
In models which have focused on the adaptation and/or the phosphorylation cascade, receptors are commonly modelled as complexes consisting of MCP, CheW and CheA. This is justified by the tight association of CheW and CheA to the MCP receptors.
Descriptions of the motor bias
Given the limited detail available on the interactions of both CheY-p and CheY with the cytoplasmic FliM end of flagella motors and that this interaction may be quite complex, it has been a common, simplifying assumption of a number of authors that the fraction of time a motor spends spinning counter-clockwise can be expressed in terms of CheY-p by a Hill function. Such reactions are based on experimental observations.
Description of the model
The model we will be using to predict biochemical behaviour of the system is a simplification of the real system.
The simplified model does not consider the normal metabolism in the cell. It only includes the most important enzymes in the normal E. coli chemotaxis pathway and the proteins responsible for coupling the phototaxis to the chemotaxis.
Below, you will se a schematic representation of the system our biochemical modelling was performed on. The crossed-over enzymes and proteins have not been modelled as a differential equation but considered as a constant in the system.
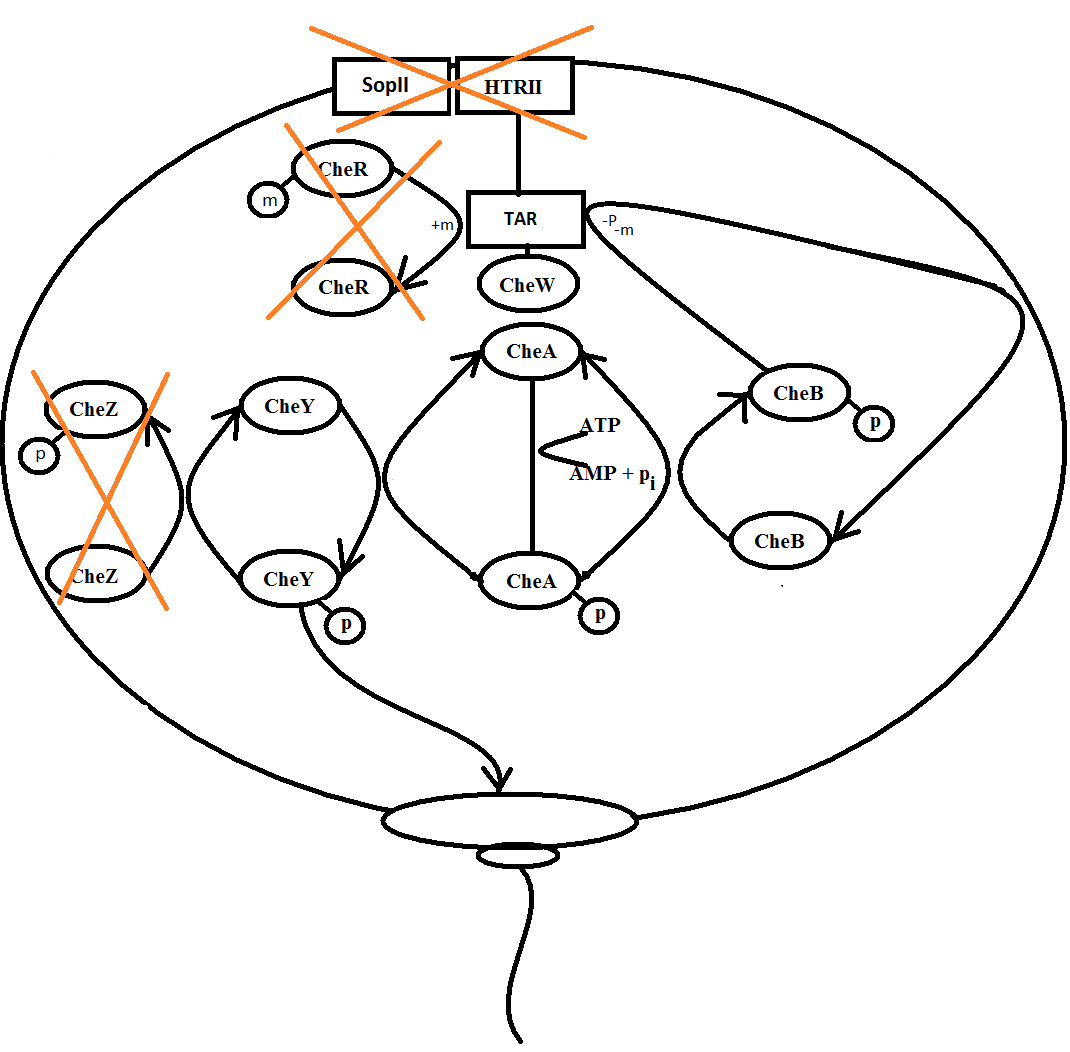
Figure 2: Schematics of the system we want make. The enzymes that are crossed-over are considered constant in our system
The important enzymes in the system are mathematically represented by a differential equation under the assumption of steady state in the system and modelled in a demo version of the program [http://www.berkeleymadonna.com/ Berkeley Madonna].
When light hits the photo receptor SRII, the signal is transduced through the HtrII and Tsr and on to the enzyme CheW. CheW is then activated because it is a straight-forward signal transduction. The proteins SRII and HtrII are not considered in the model and the equation for the activation of CheW is:
When the CheW enzyme is activated, it induces auto-phosphorylation of CheA. The phosphorylated CheA can then phosphorylate the enzymes CheY and CheB. The equation for CheA-p concentration is given by:
Phosphorylated CheY is produced by phosphorylation of CheA and is removed by CheZ which, in this model, is defined by a constant. This can be described mathematically by this equation:
Phosphorylated CheB is dephosphorylated when methylated Tsr is demethylated, this can be described mathematically by this equation:
Methylated Tsr is constantly produced by the CheR enzyme which, in this model, is also considered a constant. The demethylation is performed by CheB-p. The concentration of methylated Tsr can be described by this equation:
In Berkeley Madonna it is possible to define different parameters as sliders which enable the user to change parameters continually while the model is running. The sliders used in the model are:
Model results
References
[1] M.J. Tindall, S.L. Porter, P.K. Maini, G. Gaglia, J.P. Armitage [http://www.springerlink.com/content/f602r72767124602/ Overview of Mathematical Approaches Used to Model Bacterial Chemotaxis I: The Single Cell] Bulletin of Mathematical Biology (2008) 70: 1525–1569
[2] Nikita Vladimirov, Linda Løvdok, Dirk Lebiedz, Victor Sourjik [http://www.ploscompbiol.org/article/info%3Adoi%2F10.1371%2Fjournal.pcbi.1000242 Dependence of Bacterial Chemotaxis on Gradient Shape and Adaptation Rate] Computational Biology (2008) 4(12): e1000242
 "
"

