Team:uOttawa/Project
From 2010.igem.org
 |
||||||||
The project (mission statement)
Yeast is one of the earliest organisms used in biotechnology, even if we did not know that we were doing science at the time. Humans have been experimenting with yeast ever since our ancestors brewed their first beer. However, there is more to yeast than just making beer and bread; it also happens to be one of the simplest eukaryotic organisms. Due to its homology with mammalian organisms and its ease of genetic manipulation, yeast has become a model organism for the study of systems biology. Many key genes in mammalian organisms were first identified in yeast. For this reason, we believe it is important to develop tools that improve our ability to use yeast as a standard BioBrick chassis.
In summary we have BioBricked two yeast selection markers and have shown that they work, constructed a novel plasmid which greatly increases the efficiency of genomic integration into yeast, designed a framework based on the Gibson in vivo recombination for rapid assembly of genetic libraries, BioBricked several new yeast promoters, developed a novel strategy for dissecting promoters. Please, browse through the following sections in order to learn more about each aspect of our project. We hope that these contributions will improve BioBrick
New selection markers There are generally two types of selection markers used in yeast, auxotropic markers and selection markers that confer drug resistance. Standard yeast strains have deletions and mutations in genes in a pathway that generates an essential metabolite (such as an amino acid). These yeast strains must be grown on media supplemented with that particular metabolite. When integrating constructs into yeast, a functional copy of that gene can be brought over. Only yeast cells that contain that received that functional copy will be able to grow on media lacking that specific metabolite. From our experience we have learned that there are two major drawbacks with such selection. Firstly this type of selection is not very strong, small background colonies that do not contain the integrated construct often appear on plates. Secondly, since the selection marker shares substantial homology with the mutated / partially deleted ORF in the wild type strain, it is possible that the selection marker can undergo homologous recombination with the wild type ORF and this results in a yeast colony that can survive on the restricted media but does not contain the construct of interest. Drug selection markers are much stronger than auxotrophic markers in yeast and for this reason they have less background on plates. We BioBricked NatMX and KanMX, which are completely heterologous and do not share any homology with the yeast genome. There is less chance that they will recombinate into the incorrect locus. They also provide resistance to Nat and G418 respectively
Ade4 targeting vector
When implementing finely balanced synthetic networks in vivo it is very important to tightly control the copy number of network components{Guido, 2006, p01500}. In yeast the easiest way to control the copy number is to keep it at 1 by integrating constructs into the yeast genome. Constructs can be integrated into the yeast genome by homologous recombination, which is a natural yeast mechanism used to repair damaged DNA{Hua, 1997, p04230}. The length of the homology between the insert and the genome increases the specificity of targeting and cloning efficiency{Gray, 2001, p04021}. Although there are several yeast integrating vectors that already exist {Sikorski, 1989, p04102}, they have their own drawbacks. Namely, they do not comply by the BioBrick standards, they have short homologies with the yeast genome, they require special yeast deletion strains in order to function, and most do not provide colorimetric validation of successful integration. We therefore constructed a new plasmid (pADE4TA) that is able to integrate into the ADE4 locus, knocking out the ADE4 ORF with high specificity. The ADE4 locus which is flanked by ATM1 and DYN3 was chosen as the site for integration because knocking out the ADE4 gene in an ADE2 mutant strain (such as YPH500) restores the white colony phenotype by undercutting the adenine biosynthesis pathway{Ugolini, 1996, p03770}. Therefore a white colony in this background indicates successful integration at the correct locus and eliminates the need for time consuming and costly PCR validation. pADE4TA is a BioBrick backbone that conforms to Assembly standard 23 {Phillips, 2006, p04566}. The plasmid contains 200 bp homologous to the ATM1 terminator followed by the BioBrick cloning sites, followed by 200 bp homologous to the ADE4 terminator. Two version of the plasmid contain KanMX and NatMx expression cassettes (pADE4TAK and pADE4TAN respectively) between the ATM1 terminator and the BioBrick cut sites. KanMx and NatMx are completely heterologous and constructs Recombination pads included
A white colony indicates a successful recombination in the correct locus. These vectors are BioBrick compatible. Two versions of the vectors were created, one containing the expression cassette for Nat resistance, and the other containing the expression cassette for Kan resistance. Both these resistance markers completely heterologous and do not contain any homology to the yeast genome, they are less likely to integrate in a non-specific locus. They also confer resistance to drugs and allow for stronger selection over other auxotrophic markers
Proximal – Distal promoter distinction Ellis et al. developed a library of 40 variants of the inducible Gal1 promoter that contain operator sites for the binding of the TetR and the LacI repressors. These promoter libraries were generously donated to us. Eukaryotic promoters can be split into two halves, the proximal component and the distal component. The distal component is thought to be responsible for binding upstream activators. The proximal component is responsible for binding to the TATA binding protein and is directly involved in the recruitment of transcriptional machinery. We therefore hypothesized that it would be possible to shuffle around distal and proximal promoter components from various promoters. We proceeded to BioBrick the Act1 distal promoter and combine it with the proximal elements from the Ellis libraries. We were not able to finish this aspect of the project, however we have made all the BioBricks available to the registry. If this type of promoter shuffling in Eukaryotes is possible, then it can have a significant impact on eukaryotic synthetic biology. There are many interesting wild type promoters in nature that respond to a particular signal and these promoters are often used in biosensors. A problem with many of these interesting promoters is that although they respond correctly to a signal, they may not produced the desired output. For instance, a promoter that drives a GFP reporter may respond to
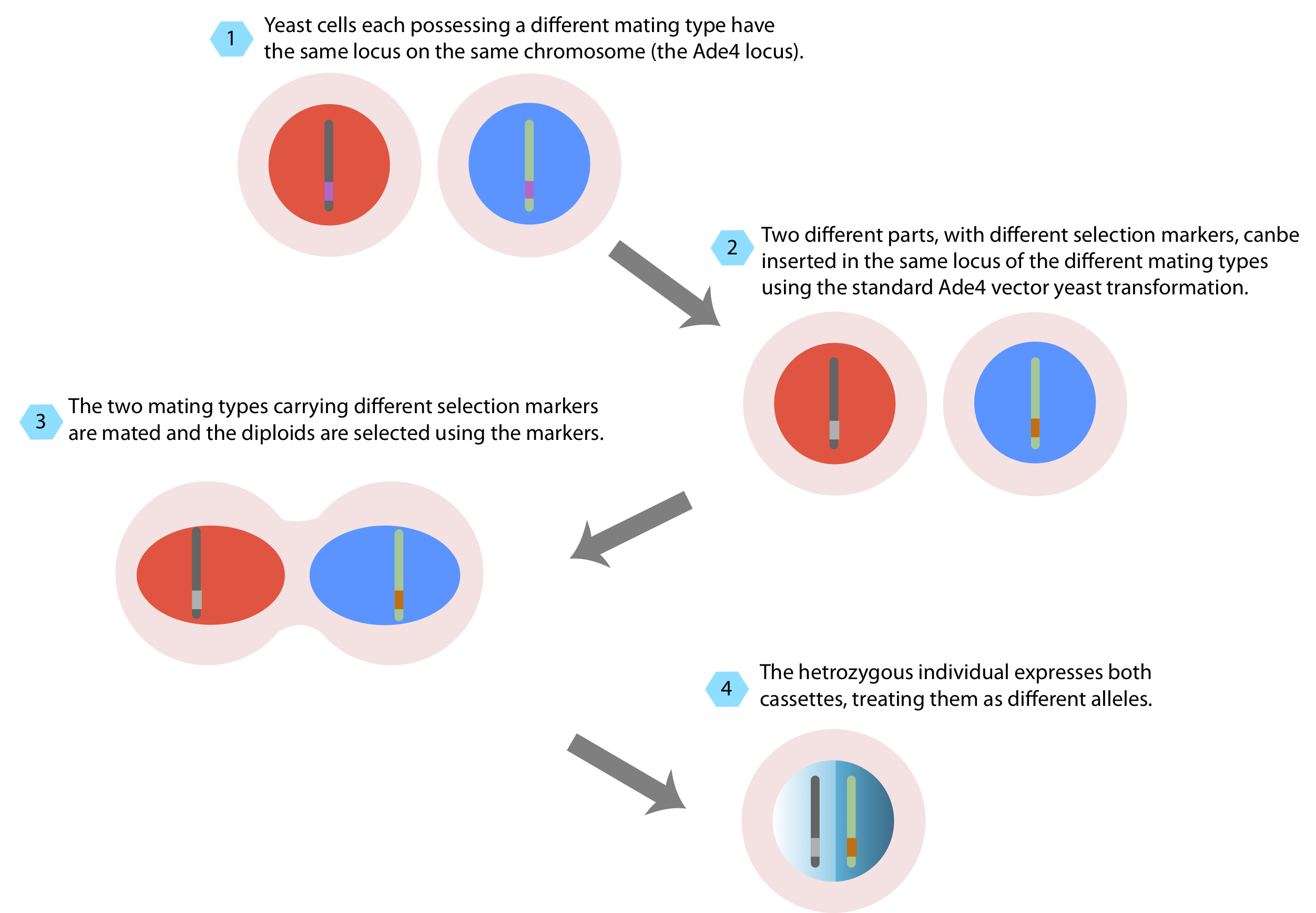
Power of yeast mating Yeast cells can live as hapoid organisms and as diploid organisms.
Promoter testing plasmids
Possible network construction with our parts (Including toggle switches)
One of the goals of the
Contents |
Overall project
Your abstract
 "
"