|
ProteInProgress: a cellular assembly line for protein manufacturing
Implementation and Results

Self-inducible promoters
Regulation of signal protein production
Experimental implementation: <partinfo>BBa_K300009</partinfo> part was assembled downstream of different constitutive promoters, thus obtaining a signal molecule generator. The choice of constitutive promoters was performed between the ones belonging to the [http://partsregistry.org/Part:BBa_J23101 Anderson’s promoters collection] ; we chose promoters according to their activities reported in the Registry of Standard Biological Parts, in order to have a thick mesh:
| Promoter | Strength (a.u.)
reported in the Registry
|
| <partinfo>BBa_J23100</partinfo> | 2547
|
| <partinfo>BBa_J23101</partinfo> | 1791
|
| <partinfo>BBa_J23105</partinfo> | 623
|
| <partinfo>BBa_J23106</partinfo> | 1185
|
| <partinfo>BBa_J23110</partinfo> | 844
|
| <partinfo>BBa_J23114</partinfo> | 256
|
| <partinfo>BBa_J23116</partinfo> | 396
|
| <partinfo>BBa_J23118</partinfo> | 1429
|
Before constructing the signal generators, <partinfo>BBa_K300009</partinfo> and <partinfo>BBa_K300010</partinfo> under the regulation of one of these constitutive promoters, we evaluated the promoter activities in Relative Promoter Units (R.P.U.) according to Data analysis for RPU evaluation, using the reporter protein RFP (Red Fluorescent Protein) in different experimental conditions (plasmids’ copy number and growth medium), many of them not yet explored and documented:
- high copy number plasmids and LB;
- high copy number plasmids and M9;
- low copy number plasmids and M9.
It was not possible to evaluate promoters activities in low copy number plasmids and LB because the RFP activity was too weak and not distinguishable from the background. RFP fluorescence and Optical Density at 600nm (O.D.600) were measured in 96-well microplates, as reported in Microplate reader experiments for constitutive promoters (R.P.U. evaluation) - Protocol #2 and data were analyzed as reported in Data Analysis RPU;
Results: results are shown here.
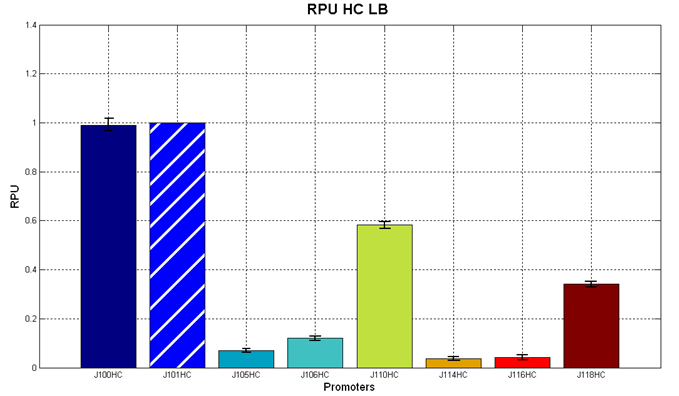 Figure 5 - R.P.U. of the studied promoters from Anderson promoters' collection, LB medium and high copy plasmid (<partinfo>BBa_J61002</partinfo>) | 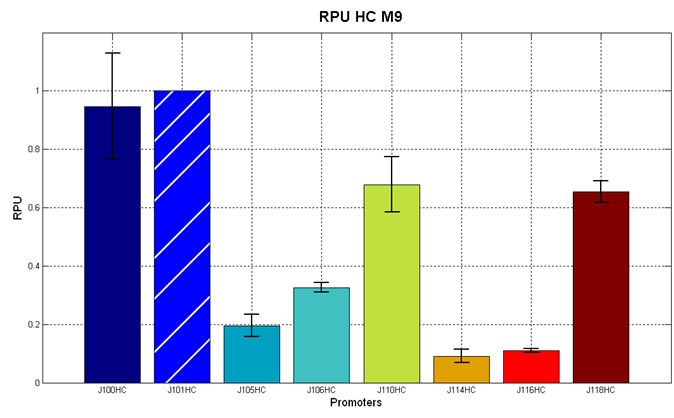 Figure 6 - R.P.U. of the studied promoters from Anderson promoters' collection, M9 medium and high copy plasmid (<partinfo>BBa_J61002</partinfo>) |
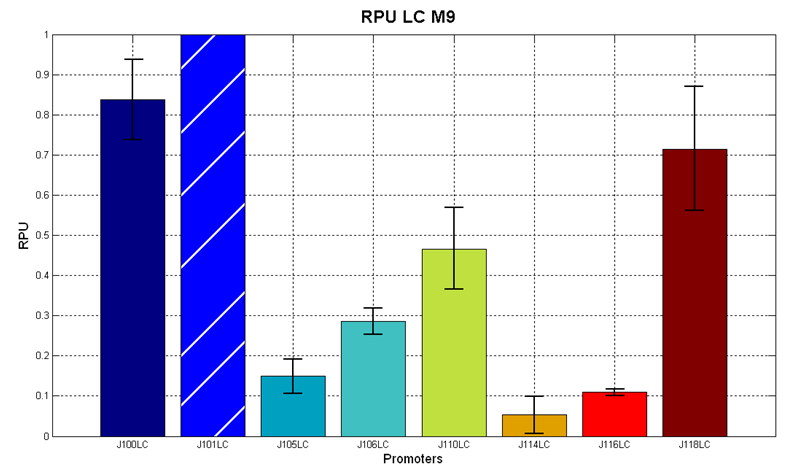 Figure 7 - R.P.U. of the studied promoters from Anderson promoters' collection, M9 medium and low copy plasmid (<partinfo>pSB4C5</partinfo>). These plasmids were constructed by assembling the EcoRI-PstI the <partinfo>BBa_J61002</partinfo>-BBa_J231xx EcoRI-PstI fragment in <partinfo>pSB4C5</partinfo>, in order to transfer the RBS-RFP-TT expression construct from <partinfo>BBa_J61002</partinfo> to <partinfo>pSB4C5</partinfo>. |
Discussion: we observed that the ranking previously documented in the Registry is not valid in all the conditions, even if a general agreement can be observed. As an example, <partinfo>BBa_J23110</partinfo> in high copy plasmid is stronger than <partinfo>BBa_J23118</partinfo>, in contrast with the ranking reported in the Registry.
After the evaluation of promoter activity, signal generators were constructed in high copy and low copy plasmids: <partinfo>BBa_K300009</partinfo> and <partinfo>BBa_K300010</partinfo> were assembled downstream of the above mentioned promoters, thus obtaining the following parts:
| BioBrick | Description
|
| <partinfo>BBa_K300030</partinfo> | 
J23118
|
| <partinfo>BBa_K300028</partinfo> | 
J23110
|
| <partinfo>BBa_K300029</partinfo> | 
J23116
|
| <partinfo>BBa_K300025</partinfo> | 
J23101
|
| <partinfo>BBa_K300026</partinfo> | 
J23105
|
| <partinfo>BBa_K300027</partinfo> | 
J23106
|
| <partinfo>BBa_K300017</partinfo> | 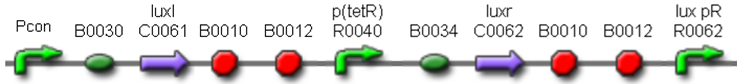
J23118
|
| <partinfo>BBa_K300014</partinfo> | 
J23110
|
| <partinfo>BBa_K300015</partinfo> | 
J23114
|
| <partinfo>BBa_K300016</partinfo> | 
J23116
|
| <partinfo>BBa_K300012</partinfo> | 
J23105
|
Some of the promoters could not be cloned upstream of these devices because they produced LuxI protein amounts that give a high metabolic burden for E. coli, so it was not possible to study all the combinations as transformans could not be obtained in some cases.
For each part, a measurement system was built, exploiting the production of the reporter gene GFP (Green Fluoresent Protein) to evaluate the "switch on" condition of every self-inducible promoter. Many different combinations were explored, in order to provide a library of promoters able to initiate transcription at the desired culture density.
Quantification of the HSL produced
Experimental implementation The new parts were, thus, characterized, measuring the HSL concentration released in the medium after a 6 hour growth of the cultures. All the details are available in this section.
<partinfo>BBa_T9002</partinfo> contained in <partinfo>pSB1A3</partinfo> in E. coli TOP10 was used as a HSL->GFP biosensor. In every experiment, a HSL-GFP calibration curve with known concentration of HSL was produced.
Results The amount of 3OC6-HSL produced after a 6 hours growth by E. coli DH5alpha bearing the parts contained in high copy plasmid <partinfo>pSB1A2</partinfo> is reported in Fig.8 and in the table:
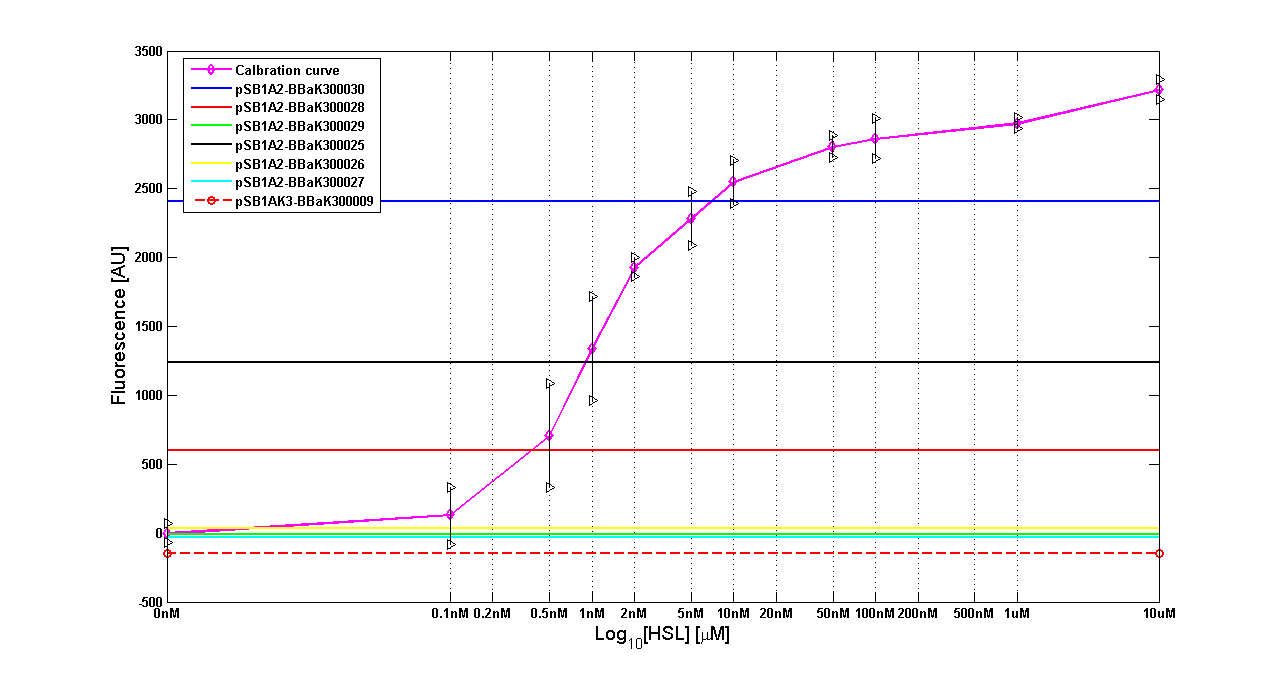 Figure 8 - <partinfo>BBa_T9002</partinfo> calibration curve for detection of [HSL] produced in high copy plasmid |
| BioBrick | Wiki name | E. coli strain | [HSL]
|
| <partinfo>BBa_K300030</partinfo> | I14 | DH5alpha | 0.7 uM
|
| <partinfo>BBa_K300028</partinfo> | I15 | DH5alpha | 0.04 uM
|
| <partinfo>BBa_K300029</partinfo> | I16 | DH5alpha | not detected
|
| <partinfo>BBa_K300025</partinfo> | I17 | DH5alpha | 0.09 uM
|
| <partinfo>BBa_K300026</partinfo> | I18 | DH5alpha | not detected
|
| <partinfo>BBa_K300027</partinfo> | I19 | DH5alpha | 0.002 uM
|
The amount of 3OC6-HSL produced after 6 hours growth by the parts contained in low copy plasmid <partinfo>pSB4C5</partinfo> is reported in Fig.9 and in the table:
 Figure 9 - <partinfo>BBa_T9002</partinfo> calibration curve for detection of [HSL] produced in low copy plasmid |
| BioBrick | Wiki name | E. coli strain | [HSL]
|
| <partinfo>BBa_K300030</partinfo> | I14 | DH5alpha | 0.005 uM
|
| <partinfo>BBa_K300028</partinfo> | I15 | DH5alpha | 0.002 uM
|
| <partinfo>BBa_K300029</partinfo> | I16 | DH5alpha | not detected
|
| <partinfo>BBa_K300025</partinfo> | I17 | DH5alpha | 0.003 uM
|
| <partinfo>BBa_K300026</partinfo> | I18 | DH5alpha | not detected
|
| <partinfo>BBa_K300027</partinfo> | I19 | DH5alpha | not detected
|
Discussion These experiments provided extremely useful informations about the capability of the signal generators to produce the 3OC6-HSL signal molecule. Data are quantitative, but incomplete because for weak promoters or medium-strength promoters contained in a low copy number plasmid the amount of 3OC6-HSL was not detectable using this system. However, this simple experiment shows that there is a strong correlation between the strength of promoter and the amount of signal molecule produced. These results confirm that the production of the autoinducer can be engineered in E. coli and different expression systems reach different amounts of 3OC6-HSL in the growth media as a function of the promoter strength. Thus, these results demonstrate that self-inducible circuits can be rationally designed from a set of well characterized standard parts.
Modulation of plasmid copy number
Signal generator and sensor device were assembled in an unique part (such as <partinfo>BBa_K300017</partinfo>, <partinfo>BBa_K300014</partinfo>, <partinfo>BBa_K300015</partinfo>, <partinfo>BBa_K300016</partinfo> and <partinfo>BBa_K300012</partinfo>) beared on high copy number plasmid <partinfo>pSB1A2</partinfo> or low copy number plasmid <partinfo>pSb4C5</partinfo>. A third alternative was the assembly of signal generator on a low copy number plasmid (<partinfo>pSB4C5</partinfo>) and the receiver device on high number plasmid (<partinfo>pSB1A2</partinfo>).
The circuits we obtained and tested are summarized here:
BioBrick
Sender | Description | Sender Vector | <partinfo>BBa_F2620</partinfo>
Receiver vector | BioBrick composite part
|
| <partinfo>BBa_K300030</partinfo>
| 
J23118
| <partinfo>pSB1A2</partinfo>
HC
| <partinfo>BBa_K300017</partinfo>
|
| <partinfo>BBa_K300028</partinfo>
| 
J23110
| <partinfo>pSB1A2</partinfo>
HC
| <partinfo>BBa_K300014</partinfo>
|
| <partinfo>BBa_K300029</partinfo>
| 
J23116
| <partinfo>pSB1A2</partinfo>
HC
| <partinfo>BBa_K300016</partinfo>
|
| <partinfo>BBa_K300026</partinfo>
| 
J23105
| <partinfo>pSB1A2</partinfo>
HC
| <partinfo>BBa_K300012</partinfo>
|
| xxx | 
J23114
| <partinfo>pSB1A2</partinfo>
HC
| <partinfo>BBa_K300015</partinfo>
|
| <partinfo>BBa_K300030</partinfo>
| 
J23118
| <partinfo>pSB4C5</partinfo>
LC
| <partinfo>BBa_K300017</partinfo>
|
| <partinfo>BBa_K300028</partinfo>
| 
J23110
| <partinfo>pSB4C5</partinfo>
LC
| <partinfo>BBa_K300014</partinfo>
|
| <partinfo>BBa_K300029</partinfo>
| 
J23116
| <partinfo>pSB4C5</partinfo>
LC
| <partinfo>BBa_K300016</partinfo>
|
| <partinfo>BBa_K300026</partinfo>
| 
J23105
| <partinfo>pSB4C5</partinfo>
LC
| <partinfo>BBa_K300012</partinfo>
|
| <partinfo>BBa_K300030</partinfo>
| 
J23118
| <partinfo>pSB4C5</partinfo>
LC
| <partinfo>pSB1A2</partinfo>
HC
| Parts are contained in two different vectors
|
| <partinfo>BBa_K300028</partinfo>
| 
J23110
| <partinfo>pSB4C5</partinfo>
LC
| <partinfo>pSB1A2</partinfo>
HC
| Parts are contained in two different vectors
|
| <partinfo>BBa_K300029</partinfo>
| 
J23116
| <partinfo>pSB4C5</partinfo>
LC
| <partinfo>pSB1A2</partinfo>
HC
| Parts are contained in two different vectors
|
| <partinfo>BBa_K300025</partinfo>
| 
J23101
| <partinfo>pSB4C5</partinfo>
LC
| <partinfo>pSB1A2</partinfo>
HC
| Parts are contained in two different vectors
|
| <partinfo>BBa_K300026</partinfo>
| 
J23105
| <partinfo>pSB4C5</partinfo>
LC
| <partinfo>pSB1A2</partinfo>
HC
| Parts are contained in two different vectors
|
| <partinfo>BBa_K300027</partinfo>
| 
J23106
| <partinfo>pSB4C5</partinfo>
LC
| <partinfo>pSB1A2</partinfo>
HC
| Parts are contained in two different vectors
|
Results
The following measurement systems were realized assembling GFP downstream of each self-inducible device. The parts characterized are reported in this table:
| Sender device
| Sensor systems with GFP
| Measurement Device
|
<partinfo>BBa_K300030</partinfo> in <partinfo>pSB1A2</partinfo>

J23118
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A2</partinfo>
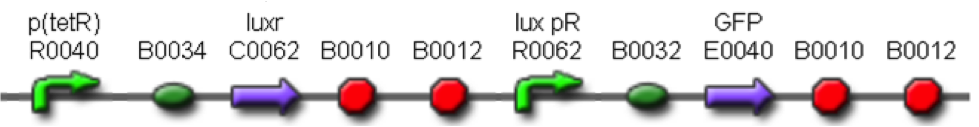
in <partinfo>pSB1A2</partinfo>
| <partinfo>BBa_K300024</partinfo>
in <partinfo>pSB1A2</partinfo>
|
<partinfo>BBa_K300028</partinfo> in <partinfo>pSB1A2</partinfo>

J23110
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A2</partinfo>

in <partinfo>pSB1A2</partinfo>
| <partinfo>BBa_K300021</partinfo>
in <partinfo>pSB1A2</partinfo>
|
<partinfo>BBa_K300029</partinfo> in <partinfo>pSB1A2</partinfo>

J23116
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A2</partinfo>

in <partinfo>pSB1A2</partinfo>
| <partinfo>BBa_K300022</partinfo>
in <partinfo>pSB1A2</partinfo>
|
<partinfo>BBa_K300026</partinfo> in <partinfo>pSB1A2</partinfo>

J23105
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A2</partinfo>

in <partinfo>pSB1A2</partinfo>
| <partinfo>BBa_K300019</partinfo>
in <partinfo>pSB1A2</partinfo>
|
xxx

J23114
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A2</partinfo>

in <partinfo>pSB1A2</partinfo>
| <partinfo>BBa_K300023</partinfo>
in <partinfo>pSB1A2</partinfo>
|
<partinfo>BBa_K300030</partinfo> in <partinfo>pSB4C5</partinfo>

J23118
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB4C5</partinfo>

in <partinfo>pSB1A2</partinfo>
| <partinfo>BBa_K300024</partinfo>
in <partinfo>pSB4C5</partinfo>
|
<partinfo>BBa_K300028</partinfo> in <partinfo>pSB4C5</partinfo>

J23110
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB4C5</partinfo>

in <partinfo>pSB1A2</partinfo>
| <partinfo>BBa_K300021</partinfo>
in <partinfo>pSB4C5</partinfo>
|
<partinfo>BBa_K300029</partinfo> in <partinfo>pSB4C5</partinfo>

J23116
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB4C5</partinfo>

in <partinfo>pSB1A2</partinfo>
| <partinfo>BBa_K300022</partinfo>
in <partinfo>pSB4C5</partinfo>
|
<partinfo>BBa_K300026</partinfo> in <partinfo>pSB4C5</partinfo>

J23105
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB4C5</partinfo>

in <partinfo>pSB1A2</partinfo>
| <partinfo>BBa_K300019</partinfo>
in <partinfo>pSB4C5</partinfo>
|
<partinfo>BBa_K300030</partinfo> in <partinfo>pSB4C5</partinfo>

J23118
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo>

in <partinfo>pSB1A2</partinfo>
| Sender and Receiver are contained in two different plasmids,
cotransformed in the same cell
|
<partinfo>BBa_K300028</partinfo> in <partinfo>pSB4C5</partinfo>

J23110
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo>

in <partinfo>pSB1A2</partinfo>
| Sender and Receiver are contained in two different plasmids,
cotransformed in the same cell
|
<partinfo>BBa_K300029</partinfo> in <partinfo>pSB4C5</partinfo>

J23116
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo>

in <partinfo>pSB1A2</partinfo>
| Sender and Receiver are contained
in two different plasmids,
cotransformed in the same cell
|
<partinfo>BBa_K300026</partinfo> in <partinfo>pSB4C5</partinfo>

J23105
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo>

in <partinfo>pSB1A2</partinfo>
| Sender and Receiver are contained
in two different plasmids,
cotransformed in the same cell
|
<partinfo>BBa_K300025</partinfo> in <partinfo>pSB4C5</partinfo>

J23101
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo>

in <partinfo>pSB1A2</partinfo>
| Sender and Receiver are contained
in two different plasmids,
cotransformed in the same cell
|
<partinfo>BBa_K300027</partinfo> in <partinfo>pSB4C5</partinfo>

J23106
| <partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo>

in <partinfo>pSB1A2</partinfo>
| Sender and Receiver are contained
in two different plasmids,
cotransformed in the same cell
|
Cultures of E. coli TOP10 bearing the plasmids containing the self-inducible devices expressing G.F.P. were grown according to this protocol and all data collected were analyzed as explained in this section
 Growth curve of <partinfo>BBa_K300019</partinfo> (O.D.600) | 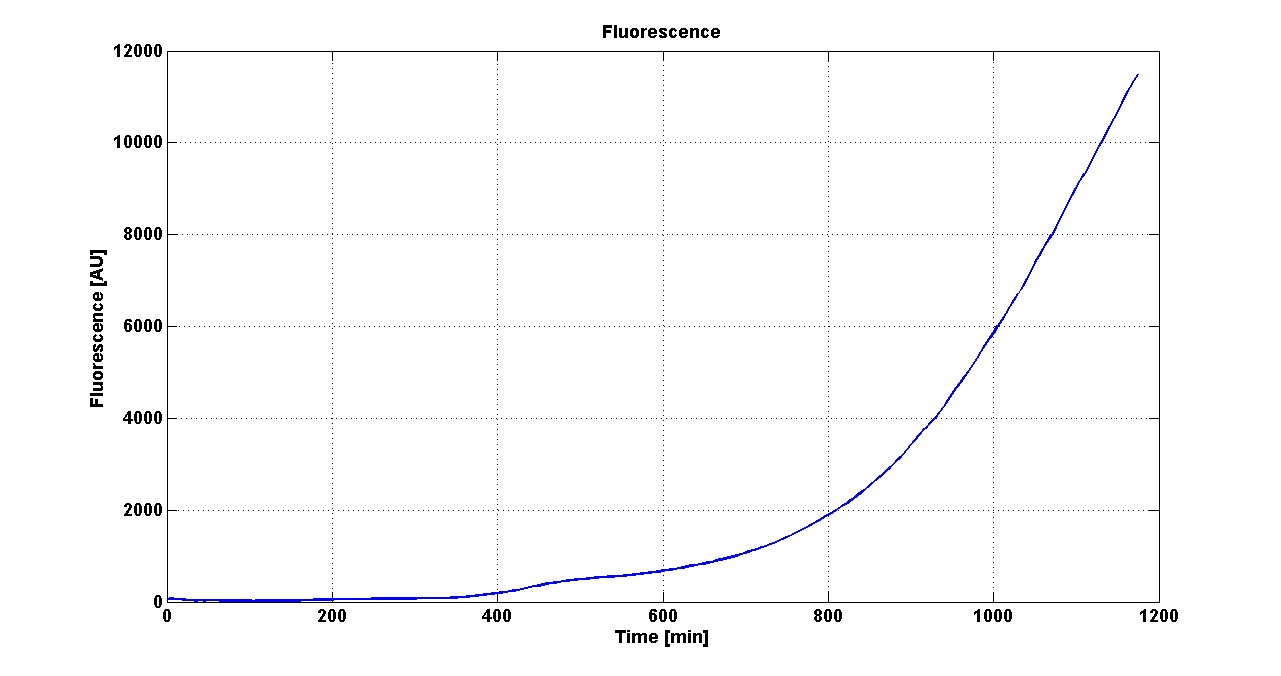 Fluorescence curve of <partinfo>BBa_K300019</partinfo> (G.F.P.) |
 Fluorescence VS Optical density curve of <partinfo>BBa_K300019</partinfo> | 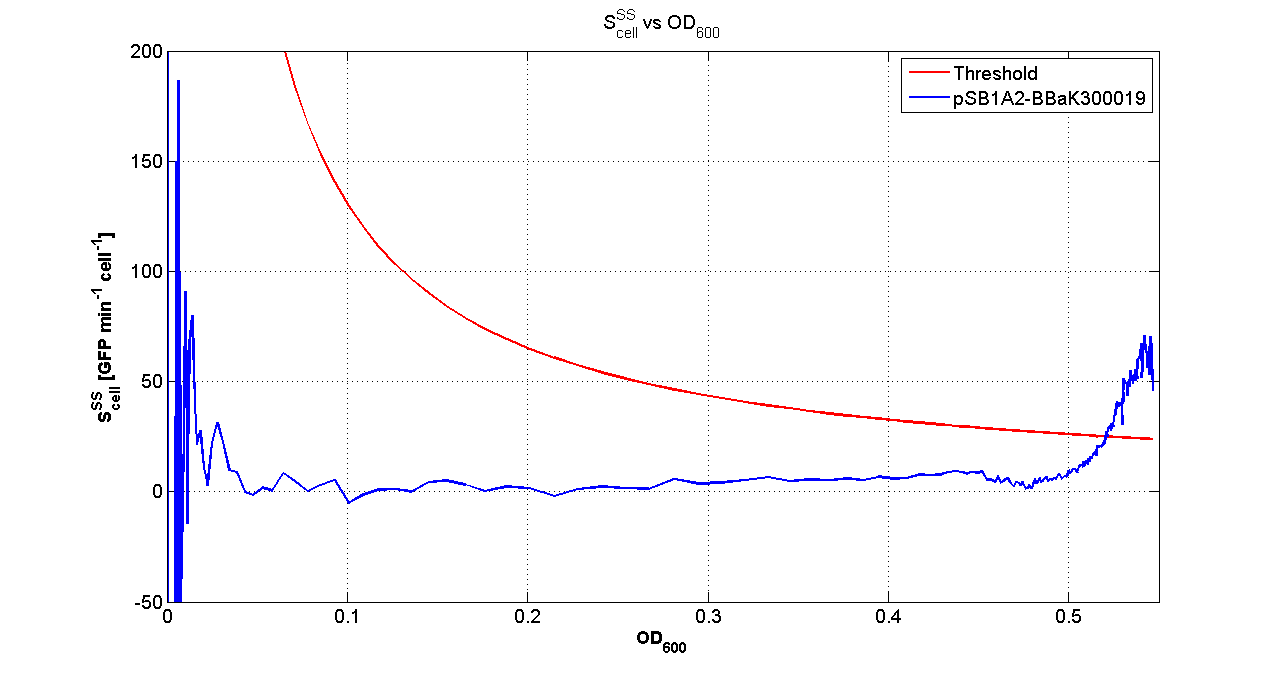 Scell=(dGFP/dt)/O.D.600 and threshold |
Doubling times were estimated as explained here
Thus, these BioBrick parts can be used to express recombinant proteins without adding an inducer to trigger the transcription of their genes; in large-scale production of such proteins this strategy could be also cost saving.
For every self-inducible device, several parameters were evaluated, as reported in this section. Results are summarized in the following tables:
Tab. 1 - Sender and Receiver on high copy plasmid <partinfo>pSB1A2</partinfo>
| Self-inducible device | Description | O.D.start LB | K_HSL LB | Doubling time LB | O.D.start M9 | K_HSL M9 | Doubling time M9
|
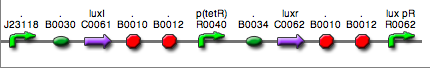
| <partinfo>BBa_K300017</partinfo> (wiki name: I7) in <partinfo>pSB1A2</partinfo> plasmid
| 
|
|
|
|
|
|
|
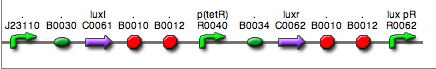
| <partinfo>BBa_K300014</partinfo> (wiki name: I8) in <partinfo>pSB1A2</partinfo> plasmid
| 
|
|
|
|
|
|
|
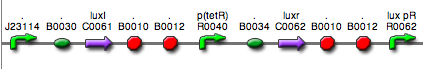
| <partinfo>BBa_K300015</partinfo> (wiki name: I9) in <partinfo>pSB1A2</partinfo> plasmid
| 
|
|
|
|
|
|
|
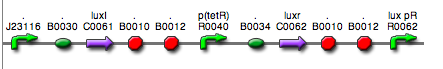
| <partinfo>BBa_K300016</partinfo> (wiki name: I10) in <partinfo>pSB1A2</partinfo> plasmid
| 
|
|
|
|
|
|
|
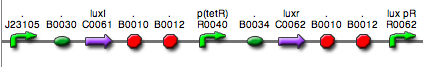
| <partinfo>BBa_K300012</partinfo> (wiki name: I12) in <partinfo>pSB1A2</partinfo> plasmid
| 
|
|
|
|
|
|
|
Tab. 2 - Sender and Receiver on low copy plasmid <partinfo>pSB4C5</partinfo>
| Self-inducible device | Description | O.D.start LB | K_HSL LB | Doubling time LB | O.D.start M9 | K_HSL M9 | Doubling time M9
|
| <partinfo>BBa_K300017</partinfo> (wiki name: I7) in <partinfo>pSB1A2</partinfo> plasmid
| 
| X
| X
| X
|
|
|
|
<partinfo>BBa_K300014</partinfo> (wiki name: I8) in <partinfo>pSB1A2</partinfo> plasmid
| 
| X
| X
| X
| X
| X
| X
|
<partinfo>BBa_K300015</partinfo> (wiki name: I9) in <partinfo>pSB1A2</partinfo> plasmid
| 
| X
| X
| X
| X
| X
| X
|
<partinfo>BBa_K300016</partinfo> (wiki name: I10) in <partinfo>pSB1A2</partinfo> plasmid
| 
| X
| X
| X
| X
| X
| X
|
<partinfo>BBa_K300012</partinfo> (wiki name: I12) in <partinfo>pSB1A2</partinfo> plasmid
| 
| X
| X
| X
| X
| X
| X
|
Tab. 3 - Sender on low copy plasmid <partinfo>pSB4C5</partinfo> and Receiver on high copy plasmid <partinfo>pSB1A3</partinfo>
| Self-inducible device
| Sender Description
| Receiver Description
| O.D.start LB
| K_HSL LB
| Doubling time LB
| O.D.start M9
| K_HSL M9
| Doubling time M9
|
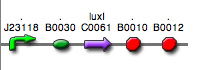
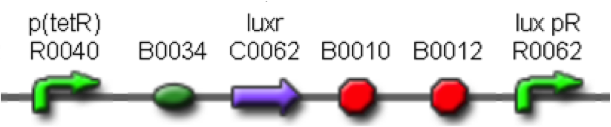
<partinfo>BBa_K300030</partinfo>
(wiki name: I14)
in <partinfo>pSB4C5</partinfo> plasmid
| 
LC
| 
HC
|
|
|
|
|
|
|
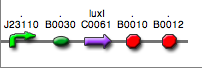
<partinfo>BBa_K300028</partinfo>
(wiki name: I15)
in <partinfo>pSB4C5</partinfo> plasmid
| 
LC
| 
HC
|
|
|
|
|
|
|
<partinfo>BBa_K300029</partinfo>
(wiki name: I16)
in <partinfo>pSB4C5</partinfo> plasmid
| 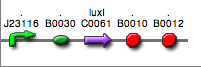
LC
| 
HC
|
|
|
|
|
|
|
<partinfo>BBa_K300025</partinfo>
(wiki name: I17)
in <partinfo>pSB4C5</partinfo> plasmid
| 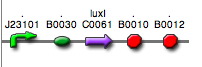
LC
| 
HC
|
|
|
|
|
|
|
<partinfo>BBa_K300026</partinfo>
(wiki name: I18)
in <partinfo>pSB4C5</partinfo> plasmid
| 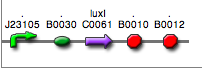
LC
| 
HC
|
|
|
|
|
|
|
<partinfo>BBa_K300027</partinfo>
(wiki name: I19)
in <partinfo>pSB4C5</partinfo> plasmid
| 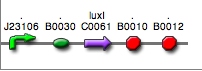
LC
| 
HC
|
|
|
|
|
|
|
 Average growth curve with ODstart evaluated by Threshold algorithm in LB |  Average growth curve with ODstart evaluated by Threshold algorithm in M9 |

Integrative standard vector for E. coli
Materials and Methods
Plasmids and strains: the <partinfo>BBa_J72008</partinfo> helper plasmid was kindly given by Prof. JC Anderson (UC Berkeley). MC1061 (<partinfo>BBa_K300078</partinfo>) and MG1655 (<partinfo>BBa_V1000</partinfo>) E. coli strains and the pCP20 helper plasmid were purchased from the Coli Genetic Stock Center (Yale University). DH5alpha (<partinfo>BBa_V1001</partinfo>) strain was purchased from Invitrogen.
Verification primers: all the oligonucleotides were purchased from Primm (San Raffaele Biomedical Science Park, Milan, Italy). The P1 (<partinfo>BBa_K300975</partinfo>) and P4 (<partinfo>BBa_K300978</partinfo>) primers had already been used in [Anderson JC et al., 2010]. The P2 (<partinfo>BBa_K300976</partinfo>) and P3 (<partinfo>BBa_K300977</partinfo>) primers have been newly designed using ApE and Amplify 3X. P2 and P3 have been designed also considering the previously used verification primers P2 and P3 in the pG80ko integrative plasmid, described in [DeLoache W, 2009].
The relative position of the P1, P2, P3 and P4 primers is shown in Fig.1:
 Figure 1: Relative position of the verification primers. a) no integrants; b) single integrant and c) integrant with multiple tandem copies. P1/P2 and P3/P4 pairs give an amplicon when at least one copy of the vector is integrated in the Phi80 locus. P2/P3 pair show an amplicon only when multiple tandem copies occur. |
Competent cells preparation: all the E. coli strains were made competent following a slightly modified version of the protocol described in [Sambrook J et al., 1989]. Briefly, cells were grown to and OD600 of ~0.4-0.6, harvested (4000 rpm, 10 min, 4°C) and the supernatant discarded. Cells were resuspended in (30 ml for each 50 ml of initial culture) pre-chilled Mg-Ca buffer (80 mM MgCl2, 20 mM CaCl2), centrifuged as before and the supernatant discarded. Cells were resuspended in (2 ml for each 50 ml of initial culture) pre-chilled Ca buffer (100 mM CaCl2, 15% glycerol), aliquoted in 0.5 ml tubes and freezed immediately at -80°C. Test the transformation efficiency as:
efficiency [CFU/ug of DNA]= # CFU * 1000 ng of DNA / amount of transformed DNA [ng]
Pir strains validation: in order to test the capability of BW25141 (pir+) and BW23474 (pir-116) to propagate this conditional replication origin, they were made competent, as well as three control strains: MC1061, MG1655 and DH5alpha. Then, a vial of 100 ul of competent cells was transformed with 2-4 ng of:
- no DNA (negative control);
- a pSB*** series vector (positive control);
- self-ligated <partinfo>BBa_K300008</partinfo> (a R6K plasmid with Cm resistance).
Self-ligated <partinfo>BBa_K300008</partinfo> was prepared by digesting <partinfo>pSB1A2</partinfo>-<partinfo>BBa_K300008</partinfo> (yielded by BioBrick Standard Assembly) with XbaI-SpeI. The insert was isolated and purified from a 1% agarose gel. Then, it was self-ligated to generate a Cm-resistant R6K plasmid. The colonies were counted in each plate and the transformation efficiency was estimated as described before.
The Chloramphenicol concentration in plates was 34 ug/ml for the high copy plasmids, 12.5 ug/ml for the medium/low copy plasmids and 12.5 for the three control strains transformed with the R6K plasmid.
BBa_K300000 construction: This section describes how this vector backbone was assembled using BioBrick parts.
- First step - <partinfo>BBa_K300982</partinfo> intermediate part construction:
- <partinfo>BBa_K300983</partinfo>, provided by Mr Gene DNA synthesis service (www.mrgene.com), was excised from its original shipping vector (pMK-RQ) through digestion with AvrII restriction enzyme (Roche). It was then isolated through a 1% agarose gel electrophoresis and gel-extracted (Macherey-Nagel NucleoSpin Extract II).
- <partinfo>BBa_K300008</partinfo> was assembled via BioBrick Standard Assembly from available parts and was excised from its vector (<partinfo>pSB1A2</partinfo>) through digestion with XbaI-SpeI (Roche). It was then isolated using the same procedure described above and it was dephosphorylated by using Antarctic Phosphatase (NEB).
- Digested <partinfo>BBa_K300983</partinfo> and <partinfo>BBa_K300008</partinfo>, all having compatible sticky ends, were ligated with T4 Ligase (Roche) and transformed into competent BW23474 E. coli strain (<partinfo>BBa_K300985</partinfo>) in order to allow the ligated plasmid propagation at high copy number. This strain was necessary because the replication origin of the resulting plasmid was the conditional R6K origin (<partinfo>BBa_J61001</partinfo>).
- Positive transformants, grown on Chloramphenicol (34 ug/ml) plates, were identified by restriction mapping with EcoRI-HindIII (Roche). The yielded plasmid was <partinfo>BBa_K300982</partinfo>.
- Second step - <partinfo>BBa_K300000</partinfo> final vector construction:
- <partinfo>BBa_K300982</partinfo> DNA was miniprepped, digested with EcoRI-PstI (Roche), run on agarose gel and gel-extracted as described above, in order to cut out and eliminate the RBS (<partinfo>BBa_B0033</partinfo>) between EcoRI and PstI.
- <partinfo>BBa_I52002</partinfo> DNA was excised from <partinfo>pSB4C5</partinfo> by EcoRI-PstI (Roche) digestion and isolated by gel run and gel extraction as described above.
- Digested <partinfo>BBa_K300982</partinfo> and <partinfo>BBa_I52002</partinfo> were ligated and transformed into competent DB3.1 E. coli strain (<partinfo>BBa_V1005</partinfo>), selected with Chloramphenicol at 34 ug/ml.
- Positive transformants were screened by restriction mapping with EcoRI-HindIII (Roche) and sequencing with VF2 (<partinfo>BBa_G00100</partinfo>) and VR (<partinfo>BBa_G00101</partinfo>) standard primers. The yielded plasmid was <partinfo>BBa_K300000</partinfo>.
All the DNA manipulations were performed according to manufacturer's protocols.
Integration protocol:
- Transform the <partinfo>BBa_J72008</partinfo> helper plasmid in the host strain (MC1061 or MG1655) and select transformants on Amp (50 ug/ml) plates under permissive conditions (30°C) overnight.
- Inoculate a single colony in selective LB and let the culture grow at 30°C, 220 rpm. When the culture reaches the OD600 of 0.4-0.6 prepare chemically competent cells.
- Transform the integrative vector with the desired insert in the BBa_J72008-containing strain and select co-transformants on Cm (34 ug/ml) plates under permissive conditions (30°C) overnight. At this temperature <partinfo>BBa_J72008</partinfo> can be replicated and so the pir protein product can be expressed in the cells. The pir product enables the propagation of the integrative vector by replicating the R6K origin.
- Inoculate a single colony in 5 ml of LB + Cm at 12.5 ug/ml and incubate the culture at 37°C, 220 rpm overnight. At this temperature the <partinfo>BBa_J72008</partinfo> helper cannot be replicated and the Phi80 integrase is expressed by the remaining copies of the helper. The bacteria that are able to grow in this selective medium should be correct integrants because the integrative vector cannot be replicated by the pir product anymore.
- Streak the culture on a Cm plate (at 12.5 ug/ml) and incubate it at 43°C overnight to ensure the loss of the helper plasmid. The bacteria that form colonies should be correct integrants without the <partinfo>BBa_J72008</partinfo> helper plasmid.
Validate the loss of the helper plasmid by inoculating colonies in Cm (at 12.5 ug/ml) media and counterselecting them in Amp (at 50 ug/ml) media. Validate the correct integration position by performing colony PCR with primers P1/P2, P3/P4, P1/P4, P2,P3 and VF2/VR. Validate the phenotype (when possible).
Expected amplicon length [bp] when the vector is integrated into the Phi80 locus:
|
| No integrant
| Single integrant
| Multiple tandem integrants (>1)
|
| VF2/VR
| none
| 280 + insert length
| 280 + insert length
|
| P1/P4
| 546
| 546 + insert length + 2171 (i.e. the BBa_K300000 length)
| 546 + insert length + 2171 (i.e. the BBa_K300000 length)
|
| P1/P2
| none
| 452
| 452
|
| P3/P4
| none
| 666
| 666
|
| P2/P3
| none
| none
| 572
|
Marker excision protocol:
- Inoculate an integrant in selective LB medium and let it grow to OD600=0.4-0.6. Prepare chemically competent cells.
- Transform the pCP20 helper plasmid in the competent strain and select transformants on Amp (100 ug/ml) plates under permissive conditions (30°C) overnight. At this temperature the pCP20 can be replicated. The pCP20 plasmid contains Amp and Cm resistance markers, a thermoinducible Flp recombinase expression system and a heat-sensitive replication origin. The permissive temperatures for the pCP20 propagation are the same as <partinfo>BBa_J72008</partinfo>.
- Inoculate a single colony in 5 ml of LB without antibiotic and incubate the culture at 37°C, 220 rpm overnight. At this temperature the pCP20 helper cannot be replicated and the Flp recombinase is expressed by the remaining copies of the helper. The bacteria should loose the R6K origin and the Cm resistance upon FRT sites recombination, mediated by Flp.
- Streak the culture on a LB plate and incubate it at 43°C overnight to ensure the loss of the helper plasmid. The bacteria that form colonies should be without the pCP20 helper plasmid.
Validate the loss of the helper plasmid by inoculating colonies in Amp (at 100 ug/ml) media and validate the loss of the Cm resistance from the genome by inoculating colonies in Cm (at 12.5 ug/ml) media. Validate the correct length of the integrated part without Cm resistance and R6K origin by performing colony PCR with primers P1/P4 (which amplify the entire Phi80 locus) and VF2/VR (which amplify the integrated part). Validate the phenotype (when possible).
Colony PCR: a single colony or 1 ul of culture was added to the Invitrogen Platinum Taq reaction mix and was heated at 94°C for 10 min. Then it was assayed with this cycle (X 35): 94°C 30 sec, 60°C (for VF2/VR) or 63°C (for the other primers) 30 sec, 72°C according to the amplicon expected length (1Kb/min). Then the reaction was kept at 72°C for 10 min and it was run on a 1% agarose gel with the GeneRuler 1Kb Plus DNA ladder (Fermentas).
Fluorescence assays: integrants were inoculated in 1 ml of M9 + Cm (12.5 ug/ml) and grown at 37°C, 220 rpm overnight. The cultures were diluted 1:100 in 2 ml of selective M9 and let grow for about 4-6 hours under the same conditions as before. Three 200 ul aliquots for each culture were transferred to a 96-well microplate and assayed in the Infinite F200 microplate reader (Tecan) for about 20 hours with the following kinetic cycle: 37°C, 5 min sampling time, linear shaking 15 sec (amplitude=3), wait 5 sec, measure absorbance at 600nm, measure fluorescence with the proper filter (EX:nm/EM:540nm for GFP or EX:535nm/EM:620nm for RFP) with gain=70. The same protocol was followed for the MC1061 and the MG1655 non-integrant strains, which were grown in M9 without antibiotic.
Data analysis: the absorbance measurements were normalized by subtracting the absorbance of the M9, while the fluorescence measurements were normalized by subtracting the fluorescence of the non-integrant strains over time. For each well, the Scell signal (proportional to the reporter protein synthesis rate per cell) was computed as (1/OD600*dXFP/dt), where OD600 is the normalized absorbance and XFP is the normalized fluorescence. The Scell signal was then averaged over time to obtain a single value for each well. Results are presented as the average Scell with their 95% confidence intervals of the mean.
Results
Validation of pir strains to propagate the R6K replication origin
The BW25141 (pir+) and BW23474 (pir-116) E. coli strains were chosen to propagate the vectors with the R6K replication origin at medium copy (~15 molecules per cell) and high copy (~250) respectively.
The results about their capability to propagate R6K plasmids are shown here:
| Strain
| Efficiency with no DNA
| Efficiency with pSB*** (positive control)
| Efficiency with the self-ligated <partinfo>BBa_K300008</partinfo> (R6K plasmid)
|
| BW25141 (<partinfo>BBa_K300084</partinfo>, pir+)
| 0
| 10^5
| 10^5
|
| BW23474 (<partinfo>BBa_K300085</partinfo>, pir-116)
| 0
| 10^6
| 10^6
|
| DH5alpha (<partinfo>BBa_V1001</partinfo>, neg control)
| 0
| 10^8
| 0
|
| MC1061 (<partinfo>BBa_K300078</partinfo>, neg control)
| 0
| 10^6
| 0
|
| MG1655 (<partinfo>BBa_V1000</partinfo>, neg control)
| 0
| 10^5
| six small colonies from 2 ng of DNA
|
These results show that the R6K conditional replication origin can be only propagated in pir+ and pir-116 strains (<partinfo>BBa_K300084</partinfo> and <partinfo>BBa_K300085</partinfo>), while the transformation of other strains with the R6K plasmid yielded no colonies after transformation. The exception was MG1655, for which six small unwanted colonies appeared on the selective plate. Plasmid DNA was purified/HindIII-digested/gel-run for at least one colony for BW25141-<partinfo>BBa_K300008</partinfo> and BW23474-<partinfo>BBa_K300008</partinfo>, while all the six colonies were analyzed for MG1655-<partinfo>BBa_K300008</partinfo> plate. The electrophoresis showed the expected length of the transformed DNA for all the clones except for the MG1655 six colonies, for which a smear was present in the lane (data not shown). Probably, Cm-resistant contaminants were present in the MG1655 culture during the preparation of competent cells.
For this reason, competent cells were prepared again for MG1655 and the transformation procedure was repeated for this strain, yielding no colonies in the <partinfo>BBa_K300008</partinfo> plate as expected.
Miniprep of the BW25141 and BW23474 strains transformed with <partinfo>BBa_K300008</partinfo> yielded a DNA concentration of ~20 ng/ul (qualitatively comparable with medium copy number plasmids) and ~90-100 ng/ul (qualitatively comparable with high copy number plasmids).
The results shown in the table above also show that the R6K plasmid in pir+ and pir-116 strains was transformed with the same efficiency as the pSB*** positive control plasmid, demonstrating that the R6K origin doesn't give any handicap in plasmid transformation.
So, the BW25141 and BW23474 strains can be successfully used to propagate the integrative vector after the excision of the pUC19-derived high copy replication origin, present in the default insert <partinfo>BBa_I52002</partinfo>.
Integration of the desired BioBrick part into the Phi80 genome locus
MC1061 and MG1655 were chosen as host strains for integration. <partinfo>BBa_K173001</partinfo> (constitutive strong promoter with GFPmut3) and the EcoRI-PstI fragment of <partinfo>BBa_J61002</partinfo>-<partinfo>BBa_J23101</partinfo> (here called PconRFP - constitutive strong promoter with RFP) were chosen as two proof of concept BioBrick parts to test the integration capability of the <partinfo>BBa_K300000</partinfo> vector in the Phi80 genome locus of these strains. For this reason, <partinfo>BBa_K173001</partinfo> and PconRFP were ligated in <partinfo>BBa_K300000</partinfo> (digested with EcoRI-PstI) and propagated using BW23474.
The integration protocol was performed as described in the Materials and Methods section for 4 different combination:
| Integrant name
| Strain
| Insert of <partinfo>BBa_K300000</partinfo>
|
| MC-GFP
| MC1061
| <partinfo>BBa_K173001</partinfo>
|
| MC-RFP
| MC1061
| PconRFP
|
| MG-GFP
| MG1655
| <partinfo>BBa_K173001</partinfo>
|
| MG-RFP
| MG1655
| PconRFP
|
Three colonies grown after the overnight incubation at 43°C (step 5 of integration protocol) were analyzed for each plate. These 12 clones were called: MC-GFP-A,B,C , MC-RFP-A,B,C , MG-GFP-A,B,C and MG-RFP-A,B,C.
Validation of the loss of BBa_J72008: all the picked colonies did not grow in Amp (50 ug/ml) media, thus validating that <partinfo>BBa_J72008</partinfo> Amp-resistant helper had been actually cured from the cells. However, one of these 12 clones (MG-GFP-A) also failed to grow in Cm (12.5 ug/ml) liquid media, probably because of a mistake in its inoculation. We decided not to consider this clone and to continue with 11 clones.
Validation of the actual integration site: colony PCR was performed for all the 11 clones, using MC1061 and MG1655 without integrants as negative controls. Primer pairs P1/P2 and P3/P4 were used to validate the presence of the integrative vector in the Phi80 genomic locus, while the primer pair P2/P3 was used to validate the presence of multiple tandem integrants (see Fig.1 in Materials and Methods).
 Figure 2: colony PCR with P1/P2 on all the 11 integrant clones. The blank is the reaction mix without bacteria. Expected amplicon for correct integrants: 452 bp. |
 Figure 3: colony PCR with P3/P4 on all the 11 integrant clones. The blank is the reaction mix without bacteria. Expected amplicon for correct integrants: 666 bp. |
 Figure 4: colony PCR with P2/P3 on all the 11 integrant clones. The blank is the reaction mix without bacteria. The lanes with the amplicon were expected to come from bacteria with multiple tandem integrants. Expected amplicon for multiple integrants: 572 bp. |
PCR results with primers P1/P2 and P3/P4 showed that each clone had the correct integrant in the correct genomic position (see Materials and Methods for a list of the expected amplicon lengths). Negative controls showed no amplicons with primers P1/P2 as expected, but showed an unexpected band with P3/P4. The reason of the presence of this band was not further investigated and the results with this primer pair cannot be a useful tool for future analysis. Anyway, the P1/P2 primer pair can be sufficient to successfully validate the presence of the DNA of interest in the Phi80 genomic locus.
PCR results with primers P2/P3 showed that two clones (MC-GFP-B and MC-GFP-C) were single integrants, while all the other clones were multiple tandem integrants (i.e. the Phi80 locus contained more than one copy of the DNA of interest). Negative controls showed no amplicons, as expected.
Validation of the integrants phenotype: all the 11 clones were assayed as described in the Materials and Methods section. Unfortunately, the green fluorescent clones (MC-GFP-A,B,C and MG-GFP-B,C) did not show appreciable differences when compared to negative controls, most probably because the autofluorescence of the cells was too high and hid the GFP signal. For this reason, GFP clones were not considered for further analysis. Other instruments should be used to detect the GFP signal.
On the other hand, RFP clones (MC-RFP-A,B,C and MG-RFP-A,B,C) all showed a higher fluorescence than the negative controls (see Fig.5). As Fig.5 show, the fluorescence of the three MG-RFP had a higher variability between clones when compared to the three MC-RFP. However, the clones were not necessarily expected to behave in the same way because all of them were multiple tandem integrants and the copy number of the PconRFP construct could be arbitrary.
 Figure 5: relative GFP synthesis rate for all the RFP expressing clones. |
Chloramphenicol resistance marker excision
The marker excision was performed on two of the previously validated integrant strains: MC-RFP-A and MG-RFP-A (even if they were multiple tandem integrants).
The marker excision protocol was performed as described in the Materials and Methods section for both strains, here named:
| Original name
| Name after marker excision
|
| MC-RFP
| MC-RFPflip
|
| MG-RFP
| MG-RFPflip
|
Three colonies grown after the overnight incubation at 43°C (step 4 of marker excision protocol) were analyzed for each plate. These 6 clones were called MC-RFPflip-A,B,C and MG-FRPflip-A,B,C.
Validation of the loss of pCP20 and the resistance marker: all the 6 picked colonies failed to grow on both Amp (100 ug/ml) media and Cm (12.5 ug/ml) media. They could only grow in LB without antibiotics, thus validating that the pCP20 helper had been actually cured and the R6K-CmR DNA containing the Chloramphenicol selection marker had been actually eliminated.
Validation of the length of the integrated part: colony PCR was performed for all the 6 clones, using MC1061 and MG1655 without integrants as negative controls. Primer pairs VF2/VR and P1/P4 were used to validate if the passenger of interest was still present in the genome and the length of the entire Phi80 locus respectively after the marker excision.
 Figure 6: colony PCR with VF2/VR on all the 6 flipped clones. The blank is the reaction mix without bacteria. Expected amplicon for correct insert: 1.2 Kb. |
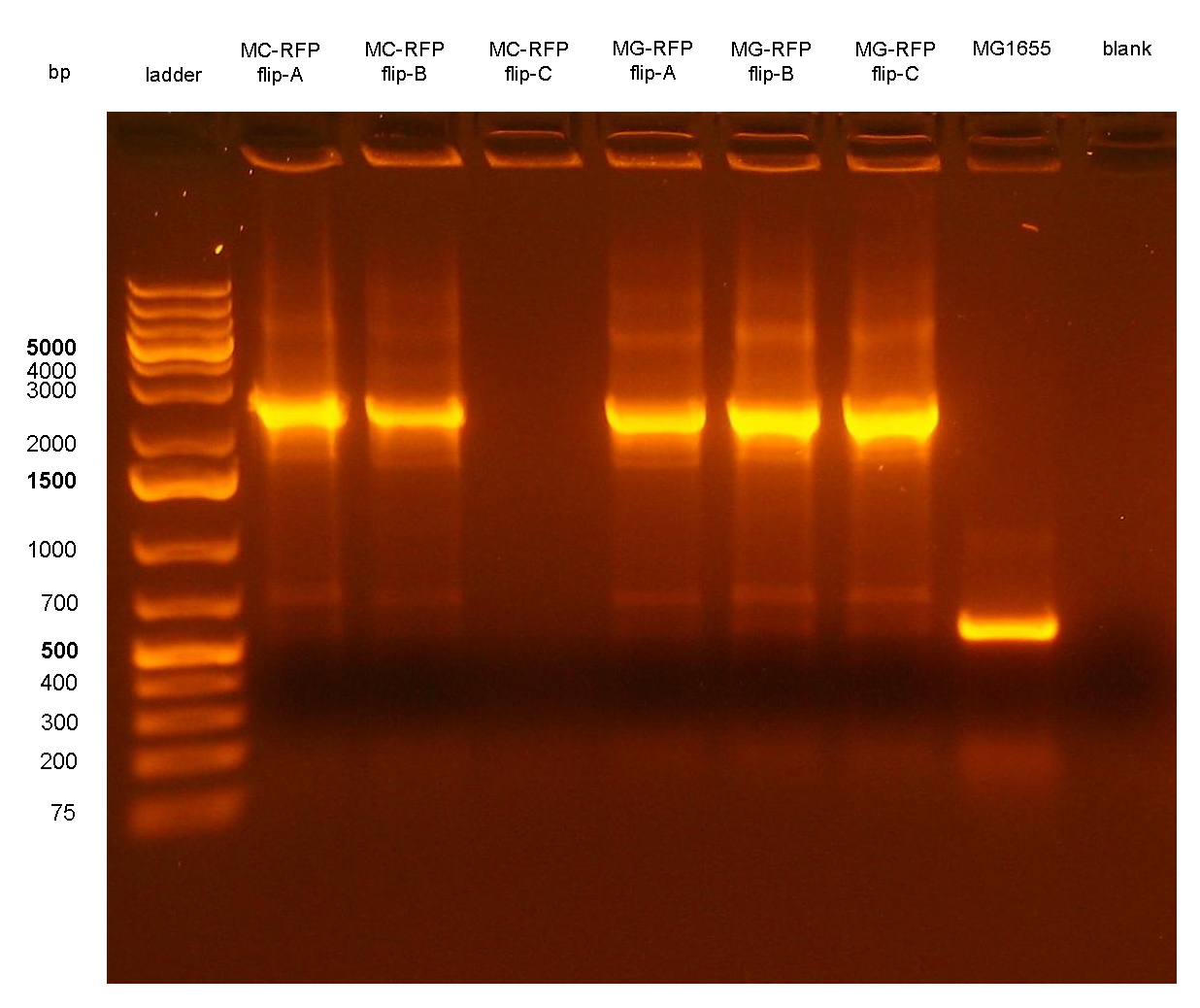 Figure 7: colony PCR with P1/P4 on all the 6 flipped clones. The blank is the reaction mix without bacteria. Expected amplicon for correct construct in the correct position: 2.3 Kb. Expected amplicon for the non-integrant strain MG1655: 546 bp. |
PCR results with primers VF2/VR showed that all the 6 clones still contain the passenger of interest, i.e. PconRFP, in the genome after the marker excision. The reaction blank, the MG1655 strain (neg control) and also the other samples showed some extra bands, but the ~1.2Kb amplicons of MC-RFP-A,B,C and MG-RFP-A,B,C had the correct length and was much brighter than the other bands.
PCR results with primers P1/P4 (Fig.7) showed that an amplicon of ~2.3Kb was present in all but one screened clones, while the MG1655 negative control showed the expected 546bp length for a non-integrant. MC-RFPflip-C did not show the P1-P4 amplicon because the reaction failed: the tube was damaged and the reaction mix was completely evaporated at the end of the PCR program. For this reason, a PCR was performed again on this clone (Fig.8).
The ~2.3Kb amplicon was consistent with a single integrant of <partinfo>BBa_K300000</partinfo>-PconRFP without the R6K-CmR DNA fragment, thus validating the successful excision of the FRT-flanked DNA fragment containing R6K-CmR and confirming that PconRFP was still present in the correct locus in single copy.
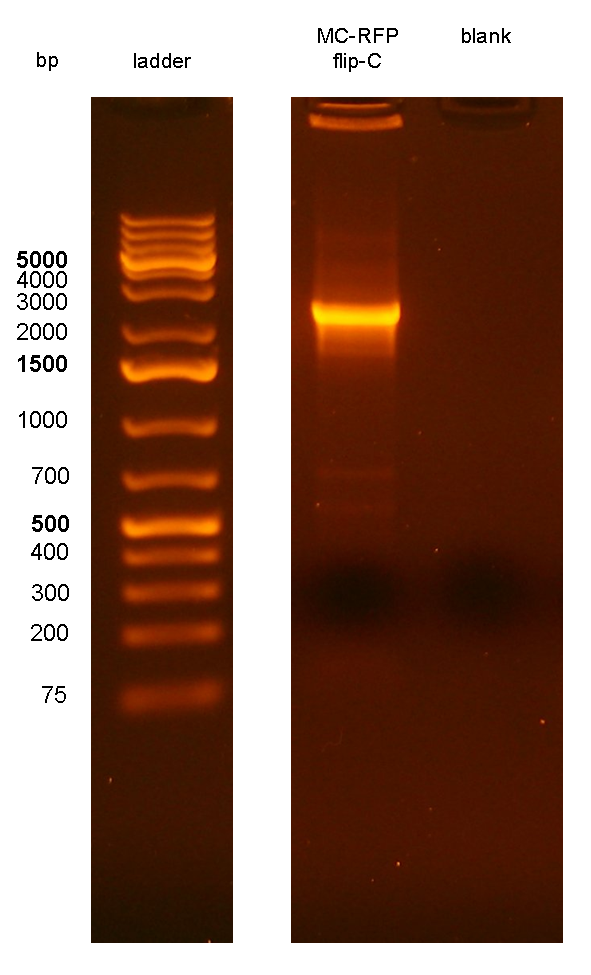 Figure 8: colony PCR with P1/P4 on MC-RFPflip-C clone. The blank is the reaction mix without bacteria. Expected amplicon for correct construct in the correct position: 2.3 Kb. |
These results showed that, even if the clones were multiple tandem integrants, they became single integrants after marker excision. This is because the Flp recombinase mediated the recombination of all the FRT sites of the multiple integrants until only a single FRT site was present in the Phi80 locus, thus leaving only the single integrant of interest without the selection marker in the genome.
Validation of the marker-less phenotype:all the 6 clones were assayed as described in the Materials and Methods section. They all showed a low variability and their fluorescence was lower than their two parents, i.e. MC-RFP-A for the MC1061 strains and MG-RFP-A for the MG1655 strains (see Fig.9). This result is consistent with the copy number of the PconRFP construct in the clones, in fact both MC-RFP-A and MG-RFP-A were multiple tandem integrants, while MC-RFPflip-A,B,C and MG-RFPflip-A,B,C were single integrants, as described above.
All the MG-RFPflip showed a very low relative RFP synthesis rate when compared to the other strains, but the signal is systematically grater than the fluorescence of the negative control, thus validating the phenotype for the MG1655 strain. MC-RFPflip-A,B,C showed a higher fluorescence than MG-RFPflip-A,B,C.
In conclusion, it has been demonstrated that, even after the marker excision process, the phenotype of the engineered cells is maintained.
 Figure 9: relative RFP synthesis rate for all the RFP-expressing clones after marker excision. In this figure, the bars corresponding to the fluorescence of the clones before marker excision is also reported to facilitate the comparison between them. Note that all the three flip clones are derived from MC-RFP-A for the MC1061 clones and from MG-RFP-A for the MG1655 clones. |
Discussion
A novel integrative vector for E. coli has been successfully designed, constructed and used to integrate two proof of concept protein expression systems in two commonly used E. coli strains.
The results showed that the vector is fully functional and can integrate into the correct targeted locus of the host chromosome through the Phi80 site-specific recombination system by using <partinfo>BBa_J72008</partinfo>, an existing BioBrick helper plasmid from the Registry. In most cases, the integration occurs in tandem copies, probably because of the too high Chloramphenicol concentration used during the selection of integrants, which forces multiple integration of Cm-resistant constructs. This concentration was the same used during the pSC101 low copy plasmid (~5 copies per cell) selection. In some cases, it is desirable to have a single copy of the desired BioBrick in the genome, for example when the gene dosage is important. In [Haldimann A and Wanner BL, 2001] the usage of Chloramphenicol at 6 ug/ml yielded a very high percentage of single integrants. However, when tested in our lab, the MG1655 strain could survive on LB plates with Cm at 6 ug/ml and also at 8 ug/ml. For this reason a higher concentration of Cm was chosen for selection. Further studies should investigate the optimal antibiotic concentration to yield the highest single integrants percentage as possible.
The Flp/FRT mediated excision of the R6K and, most importantly, of the Cm resistance marker also worked by using the pCP20 helper plasmid. The estimated efficiency of this process was 100%. In addition, multiple tandem integrants became single integrants after the marker excision. This is because the Flp recombinase mediated the recombination of all the FRT sites of the multiple integrants until only a single FRT site was present in the Phi80 locus. The marker excision is a powerful tool to engineer microbial strains for industrial protein manufacturing because the engineered organism should not carry unsafe antibiotic resistances that may be diffused in the environment.
The fluorescence phenotype confirmed the correct integration into the E. coli chromosome. As expected, in general multiple integrants showed a higher fluorescence than the single integrants.
The BioBrick compatibility and the vector modularity give the possibility to the scientific community to stably engineer novel biological functions in E. coli with a very easy and user friendly methodology. A user’s handbook about the vector usage is shared in the Registry, as well as the users experiences and the compatibility information.

Integrative standard vector for yeast
Materials and Methods
Strain: the S. cerevisiae S288C strain (<partinfo>BBa_K300979</partinfo>, genotype: MATα ρ° trp1-0) was purchased from Open Biosystems.
Construction of BBa_K300001:
- <partinfo>BBa_K300980</partinfo>, provided by Mr Gene DNA synthesis service (www.mrgene.com), was excised from its original shipping vector (pMA) through digestion with MfeI (Fermentas) and NsiI (Fermentas) restriction enzymes. It was then isolated through a 1% agarose gel electrophoresis and gel-extracted (Macherey-Nagel NucleoSpin Extract II).
- <partinfo>BBa_I763007</partinfo>, available in the Registry, was digested with EcoRI-PstI (Roche), ran on agarose gel and its <partinfo>pSB1A2</partinfo> vector was gel-extracted as described above.
- Digested <partinfo>BBa_K300980</partinfo> and <partinfo>pSB1A2</partinfo> have compatible ends (EcoRI-MfeI and PstI-NsiI). They were ligated with T4 Ligase (Roche) and transformed into competent TOP10 E. coli strain (<partinfo>BBa_V1009</partinfo>) which were then plated on Ampicillin (100 ug/ml) plates.
- Positive transformants were identified by colony PCR with VF2 (<partinfo>BBa_G00100</partinfo>) and VR (<partinfo>BBa_G00101</partinfo>) standard primers and by restriction mapping with EcoRI (Roche) or NsiI (Fermentas). The yielded plasmid had <partinfo>BBa_B0033</partinfo> flanked by EcoRI and SpeI.
- The yielded plasmid was then digested with EcoRI-SpeI (Roche) and <partinfo>BBa_K300007</partinfo> (digested with EcoRI-SpeI as well) was assembled in the vector, thus yielding <partinfo>BBa_K300001</partinfo>-<partinfo>BBa_K300007</partinfo> (i.e. there is the <partinfo>BBa_K300007</partinfo> part in the cloning site).
Yeast transformation:
- S288C strain was inoculated in 5 ml of YPD from a long term 15% glycerol stock and grown for 24h (30°C, 200rpm).
- The culture was diluted 1:10 in 50 ml of pre-warmed YPD in a 250 ml flask and was grown for additional 4 hours under the same conditions as before.
- Cells were pelleted (4000 rpm, 5 min) and resuspended in 25 ml of deionized water.
- Cells were pelleted (4000 rpm, 5 min), the supernatant was discarded and the pellet was resuspended in 1 ml of deionized water and transferred into a 1.5 ml tube.
- Cells were pelleted (4000 rpm, 30 sec), the supernatant was discarded and the pellet was resuspended in deionized water to a final volume of 1 ml (vortex mix vigorously).
- Three 100 ul aliquots were transferred into 1.5 ml tubes, while the remaining 600 ul of cells were not used in this protocol.
- The three tubes were centrifuged (4000 rpm, 30 sec) and the supernatant discarded.
- Each of the three pellets were resuspended (vortex mix vigorously) in 360 ul of transformation mix (240 ul of PEG 3350 50% w/v, 36 ul of LiAc 1.0 M, boiled salmon sperm DNA, 34 ul of linearized plasmid DNA plus water). The salmon sperm DNA was boiled for 5 min and pre-chilled before adding it in the transformation mix. The plasmid DNA was previously digested with SbfI (Fermentas), purified with the NucleoSpin Extract II kit (MN) and quantified with the NanoDrop in order to add 1 ug of DNA to the transformation mix.
- The tubes were heated at 42°C for 40 min.
- Cells were pelleted (4000 rpm, 30 sec), the supernatant was removed by pipetting and the pellet was gently resuspended in 1 ml of deionized water.
- Cells were pelleted (4000 rpm, 30 sec), the supernatant was discarded, the pellet was resuspended in 1 ml of YPD and incubated at 30°C, 200 rpm for 3 hours.
- Cells were pelleted (4000 rpm, 30 sec), resuspended in 200 ul of YPD and plated on a YPD agar plate with G418 antibiotic at 200 ug/ml.
- The plates were incubated at 30°C for about 3 days until colonies appeared.
The integration efficiency was estimated as the colony forming units (CFUs) yielded for each ug of DNA.
Protocol references:
[1] http://openwetware.org/wiki/High_Efficiency_Transformation
[2] Guldener U, Heck S, Fiedler T, Beinhauer J, Hegemann JH (1996), A new efficient gene disruption cassette for repeated use in budding yeast. Nucleic Acids Research, Vol. 24, No. 13 2519–2524.
Results
The transformed inserts and their integration efficiency in S288C are listed here:
| SbfI-digested plasmid
| ug of transformed DNA
| # of colonies
| Estimated integration efficiency [CFU/ug]
|
| <partinfo>BBa_K300001</partinfo>-<partinfo>BBa_K300006</partinfo>
| 1
| 6500
| 6.5*10^3
|
| <partinfo>BBa_K300001</partinfo>-<partinfo>BBa_K300007</partinfo>
| 1
| 1700
| 1.7*10^3
|
| no DNA
| 0
| 0
| 0
|
The colony count was quite difficult (see pictures), but it is useful to have an estimation of the integration efficiency.
 S288C transformed with no DNA (negative control). | 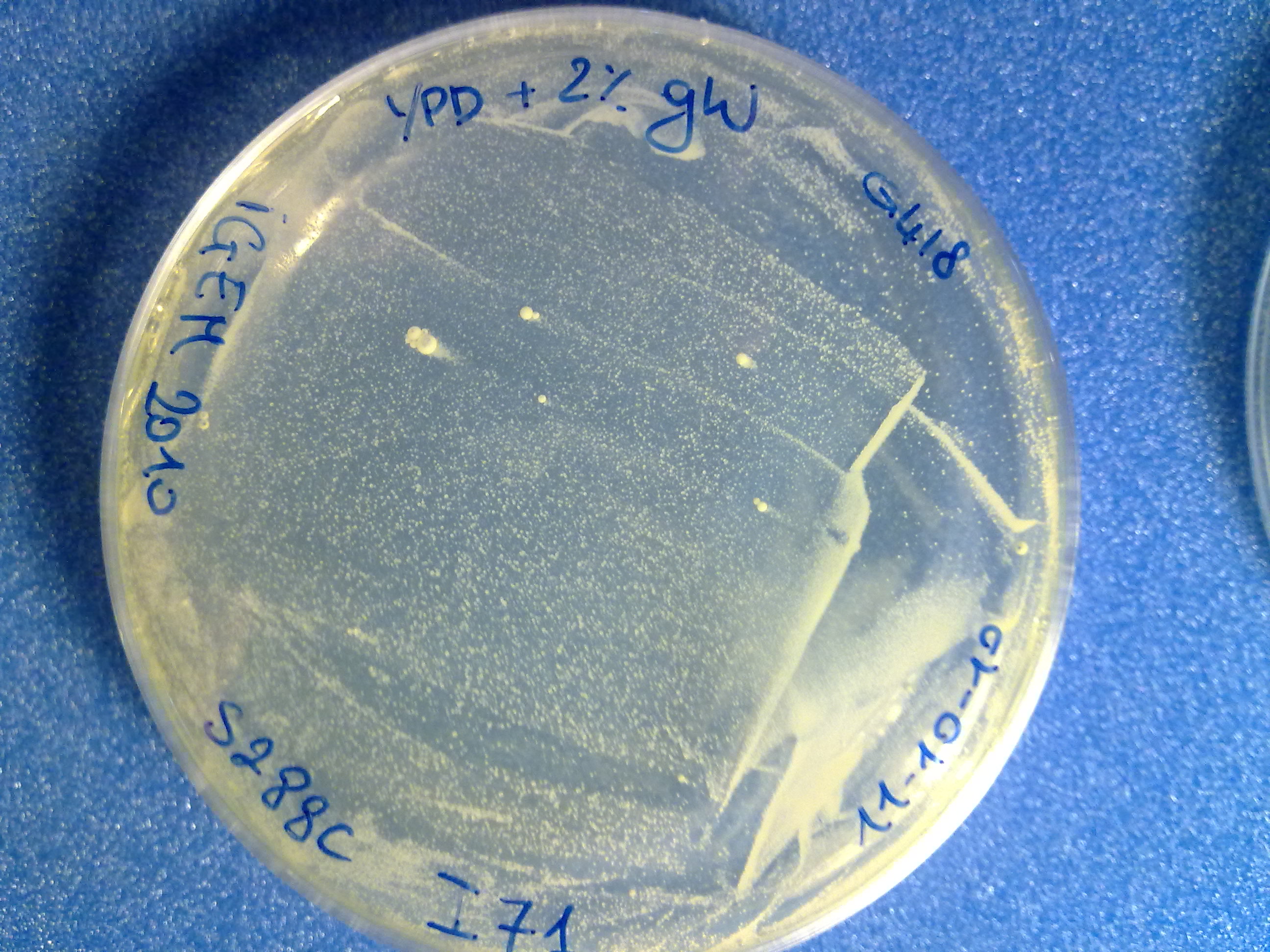 S288C transformed with linearized <partinfo>BBa_K300001</partinfo>-<partinfo>BBa_K300006</partinfo>. |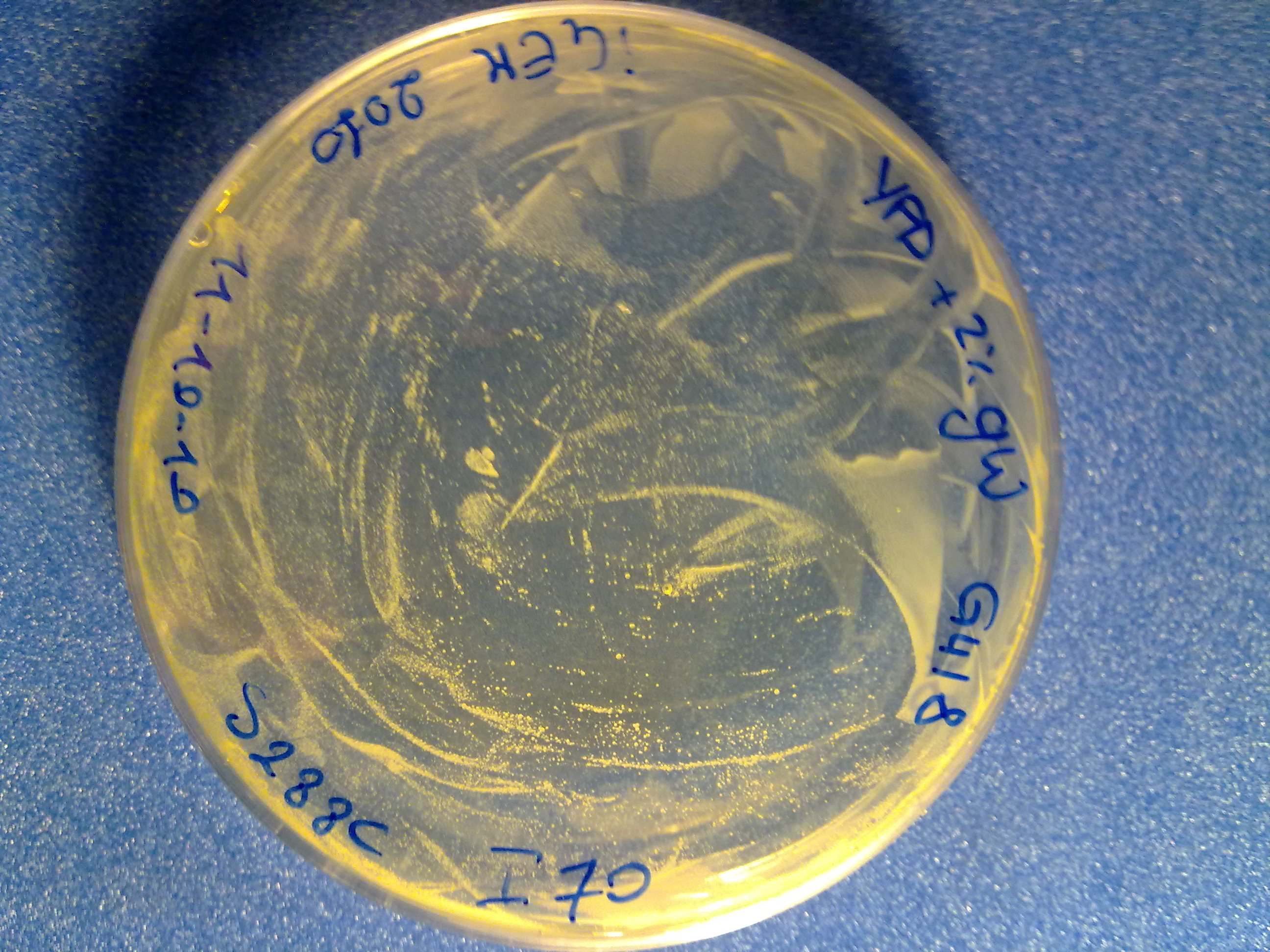  S288C transformed with linearized <partinfo>BBa_K300001</partinfo>-<partinfo>BBa_K300007</partinfo>. |
Discussion
The obtained results suggest that the integrative vector actually works and that the selection marker is highly specific (no colonies appeared on the "no DNA" plate).
Although this integrative vector has already given promising integration results, a lot of work still remains to do:
- the correct phenotype of the S288C bearing these parts has to be validated (by mOrange fluorescence measurement for the <partinfo>BBa_K300007</partinfo> part);
- the actual integration position has to be confirmed by PCR with ad hoc-designed primers;
- the KanMX selection marker excision has to be validated.
The compatibility with the BioBrick standard (RFC10, RFC12 and RFC23) enables the engineering the chromosomes of S. cerevisiae by integrating the desired biological functions following a standard procedure. Moreover, the modularity of the designed integrative vector enables the modification of the integration target site, in fact the default Homologous Regions can only target the Gal genomic system and users may need to target different DNA sequences.
As reported for the integrative vector for E. coli, the main information about the usage of this integrative vector for yeast is shared in the Registry. We hope to have positively contributed to the diffusion of standard biological parts for yeast, a very interesting chassis for synthetic biology for which only a limited number of parts are available in the Registry and only a few of them has been exaustively characterized.

Self-cleaving affinity tags to easily purify proteins
PHB production
Methods
Preparation of samples for BioPlastic (PHB) screening:
- Inoculum in LB+Amp of:
- DSMZ15372 E. coli strain
- <partinfo>BBa_B0032</partinfo>
- Cultures were left grow ON at +37°C, 220 rpm.
- Cultures of DSMZ15372 and <partinfo>BBa_B0032</partinfo> were diluted 1:100 in fresh LB+Amp and than prepared as follows:
- <partinfo>BBa_B0032</partinfo> with NOTHING added (negative control)
- DSMZ15372 with NOTHING added
- DSMZ15372 + 2% glycerol (a carbon source for bioplastic production)
- DSMZ15372 + 1mM IPTG (inducer for lac promoter, expressing bioplastic enzymes)
- DSMZ15372 + 2% glycerol + 1mM IPTG
After 5 and 30 hours' growth, Sudan Black staining protocol was performed on slides prepared for each culture. Slides were observed at the microscope.
Results
5 hours'
Sudan Black staining protocol was performed on 70ul samples and 5 microscope slides were prepared. The resulting images are shown here:
 <partinfo>BBa_B0032</partinfo> with nothing added in the culture, after 5 hours (negative control) |  <partinfo>BBa_B0032</partinfo> with nothing added in the culture, after 5 hours (negative control) |
 DSMZ15372 with nothing added in the culture, after 5 hours |  DSMZ15372 with nothing added in the culture, after 5 hours |
 DSMZ15372 with 2% glycerol added in the culture, after 5 hours |  DSMZ15372 with 2% glycerol added in the culture, after 5 hours |
 DSMZ15372 with 1mM IPTG added in the culture, after 5 hours |  DSMZ15372 with 1mM IPTG in the culture, after 5 hours |
 DSMZ15372 with 1mM IPTG and 2% glycerol added in the culture, after 5 hours |  DSMZ15372 with 1mM IPTG and 2% glycerol added in the culture, after 5 hours |
30 hours'
Sudan Black staining protocol was performed on 70ul samples and 5 microscope slides were prepared. The resulting images are shown here:
 <partinfo>BBa_B0032</partinfo> with nothing added in the culture, after 30 hours (negative control) |  <partinfo>BBa_B0032</partinfo> with nothing added in the culture, after 30 hours (negative control) |
 DSMZ15372 with nothing added in the culture, after 30 hours |  DSMZ15372 with nothing added in the culture, after 30 hours |
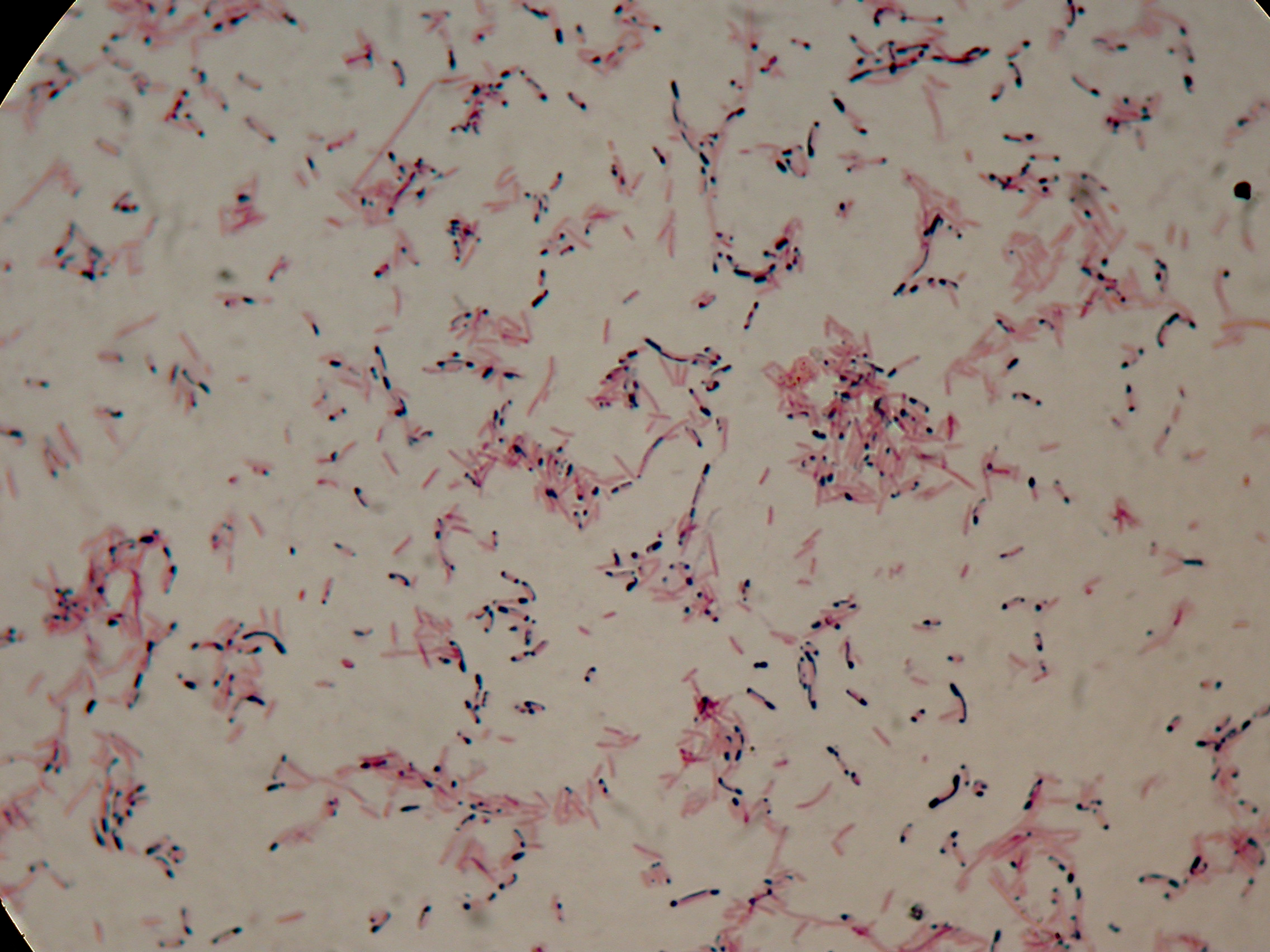 DSMZ15372 with 2% glycerol added in the culture, after 30 hours |  DSMZ15372 with 2% glycerol added in the culture, after 30 hours |
 DSMZ15372 with 1mM IPTG added in the culture, after 30 hours |  DSMZ15372 with 1mM IPTG in the culture, after 30 hours |
 DSMZ15372 with 1mM IPTG and 2% glycerol added in the culture, after 30 hours |  DSMZ15372 with 1mM IPTG and 2% glycerol added in the culture, after 30 hours |
Discussion
In the above images it is clearly possible to see that after 5 hours DSMZ15372 without any kind of addition is completely similar to negative control <partinfo>BBa_B0032</partinfo> and there is no trace of bioplastic granules production. In DSMZ15372 in which IPTG or glycerol were added to the media, it is possible to see very small dark spots that could be identified as PHB granules in a few bacteria. In samples with both IPTG and glycerol, bioplastic granules are clearly visible in many cells.
As expected after 30 hours negative control <partinfo>BBa_B0032</partinfo> does not show any trace of granules, while DSMZ15372 shows bioplastic granules in each experimental condition. This demonstrates that PHB can be produced without the presence of both IPTG and glycerol.
In conclusion, we validated the right culture conditions for PHB production using an existing engineered strain. This enables the implementation of the purification system designed in this project. Future work can explore the possibility of producing and optimizing the PHB production with BioBrick standard parts.
Fusion protein validation
The newly designed and constructed affinity tags <partinfo>BBa_K300002</partinfo>, <partinfo>BBa_K300093</partinfo>, <partinfo>BBa_K300094</partinfo>, <partinfo>BBa_K300097</partinfo> were assembled to a constitutive or inducible promoter with RBS upstream and to a Silver fusion-compatible GFP coding sequence with terminator (<partinfo>BBa_K300005</partinfo>). The resulting measurement parts were assayed to validate the bacterial growth and GFP synthesis rate of the following constructs in order to verify the right protein folding.
Costitutive promoter devices
Methods
Inoculum (into 5 ml LB+Amp) from glycerol stock of:
- <partinfo>BBa_K300086</partinfo>
- <partinfo>BBa_K300088</partinfo>
- <partinfo>BBa_K300090</partinfo>
- <partinfo>BBa_K300099</partinfo>
- <partinfo>BBa_K173000</partinfo> (positive control, J23100 constitutive promoter expressing GFP)
- <partinfo>BBa_B0031</partinfo> (negative control, a non-fluorescent culture)
Cultures were grown ON at 37°C, 220 rpm.
The following day cultures were diluted 1:100 and let grow again for about five hours at 37°C, 220 rpm.
The optical density (O.D.) of each culture was than measured with TECAN Infinte F200. Samples were diluted in order to obtain the same O.D. equal to 0.02.
Then we performed a 21-hour experiment with measurements of absorbance and green fluorescence every five minutes with TECAN Infinite F200; cultures were shaken for 15 seconds every five minutes. Acquired data were blanked by subtracting the media absorbance (for absorbance measurements) and the <partinfo>BBa_B0031</partinfo> fluorescence (for fluorescence measurements). Then, the relative GFP synthesis rate per cell was evaluated by computing (1/O.D.600)*dGFP/dt, where O.D.600 is the blanked absorbance of the culture of interest and GFP is its blanked fluorescence.
Each value shown below is the mean of three measurements in exponential phase and error bars represent the 95% confidence interval of the mean.
Results
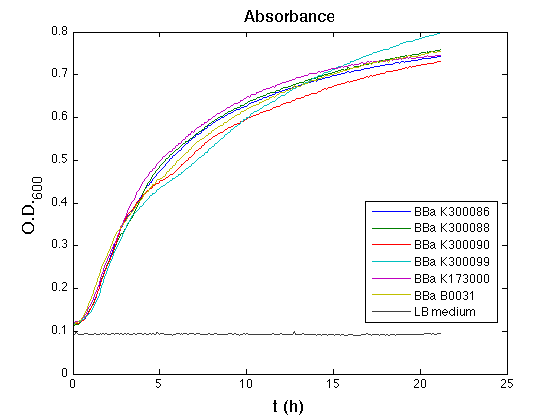
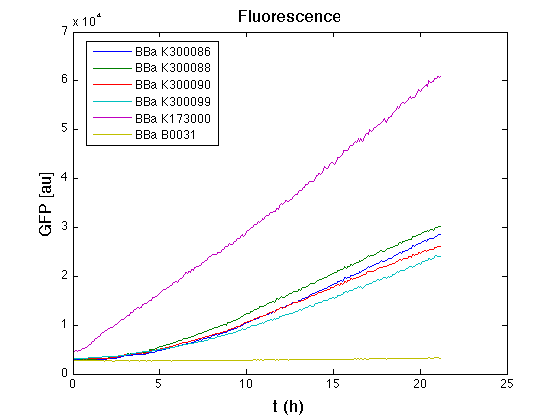
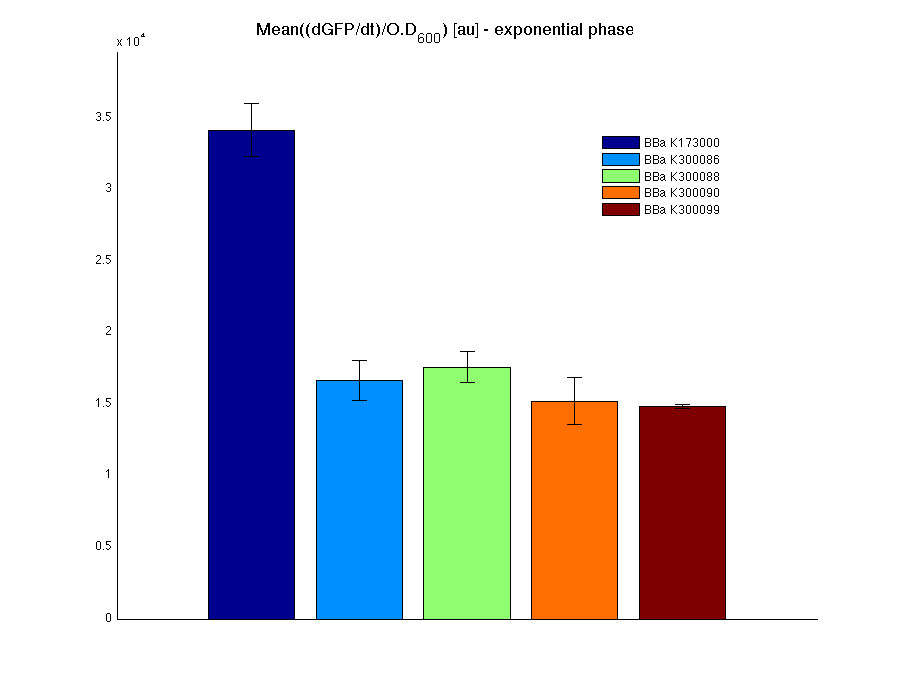 Mean (dGFP/dt)/O.D. over the exponential phase (under the hypothesis that GFP half-life in fusion contructs is similar to the original one -<partinfo>BBa_E0040</partinfo>) |
| Culture | Doubling time [min.] ± std error |
| <partinfo>BBa_K173000</partinfo> | 76.3336 ± 1.4362 |
| <partinfo>BBa_K300086</partinfo> | 73.6685 ± 1.6245 |
| <partinfo>BBa_K300088</partinfo> | 74.8806 ± 2.7699 |
| <partinfo>BBa_K300090</partinfo> | 75.9433 ± 3.6808 |
| <partinfo>BBa_K300099</partinfo> | 78.4634 ± 2.5622 |
| <partinfo>BBa_B0031</partinfo> | 70.8421 ± 2.2181 |
Discussion
All the cultures showed a similar growth curve; doubling time was computed as described here in order to obtain information about the metabolic burden due to the synthesis of the studied fusion proteins. It is possible to see that all doubling times are comparable; it is possible to assert that the expression of these BioBrick parts doesn't cause abnormal stress to the cells.
From GFP curve it is possible to appreciate that in <partinfo>BBa_K300086</partinfo>, <partinfo>BBa_K300088</partinfo>, <partinfo>BBa_K300090</partinfo>, <partinfo>BBa_K300099</partinfo> GFP accumulation is very similar and it is significantly different from the one of the negative control <partinfo>BBa_B0031</partinfo>. These results show that the green fluorescent protein assembled downstream of the genetic circuit is correctly folded.
The mean protein synthesis rate was also computed over the exponential growth phase, showing again an appreciable GFP production rate that is about half of the positive control GFP.
3OC6HSL inducible devices
Methods
Inoculum (into 5 ml LB+Amp) from glycerol stock of:
- <partinfo>BBa_K300091</partinfo>
- <partinfo>BBa_K300092</partinfo>
- <partinfo>BBa_K173000</partinfo> (positive control)
- <partinfo>BBa_B0031</partinfo> (negative control)
Cultures were grown ON at 37°C, 220 rpm.
The following day cultures were diluted 1:100 and let grow again for about five hours at 37°C, 220 rpm.
The optical density (O.D.) of each culture was than measured with TECAN Infinte F200. Samples were diluted in order to obtain the same O.D. equal to 0.02.
Then we performed a 21-hour experiment with measurements of absorbance and green fluorescence every five minutes using TECAN Infinite F200; cultures were shaken for 15 seconds every five minutes. <partinfo>BBa_K300091</partinfo> and <partinfo>BBa_K300092</partinfo> constructs were induced with 100nM of HSL directly in the 96-well microplate. Acquired data were blanked by subtracting the media absorbance (for absorbance measurements) and the <partinfo>BBa_B0031</partinfo> fluorescence (for fluorescence measurements). Then, the relative GFP synthesis rate per cell was evaluated by computing (1/O.D.600)*dGFP/dt, where O.D.600 is the blanked absorbance of the culture of interest and GFP is its blanked fluorescence.
Each value shown below is the mean of three measurements in exponential phase and error bars represent the 95% confidence interval of the mean.
Results
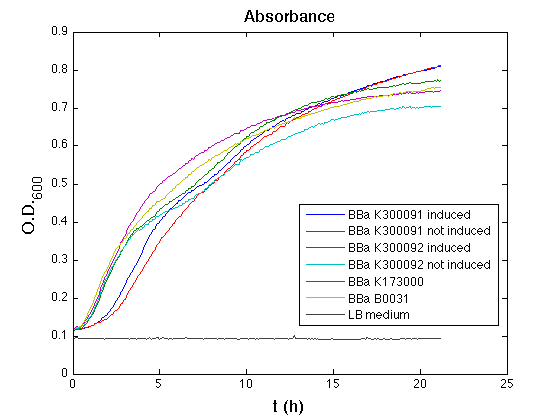
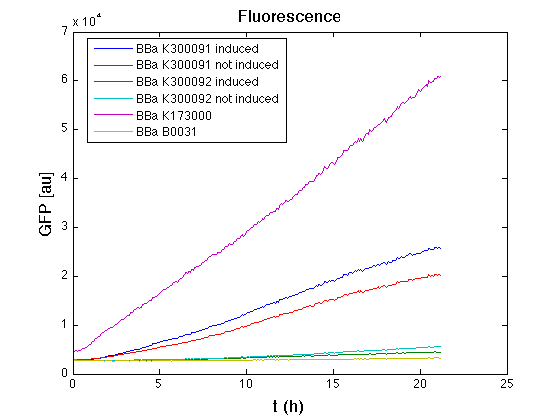
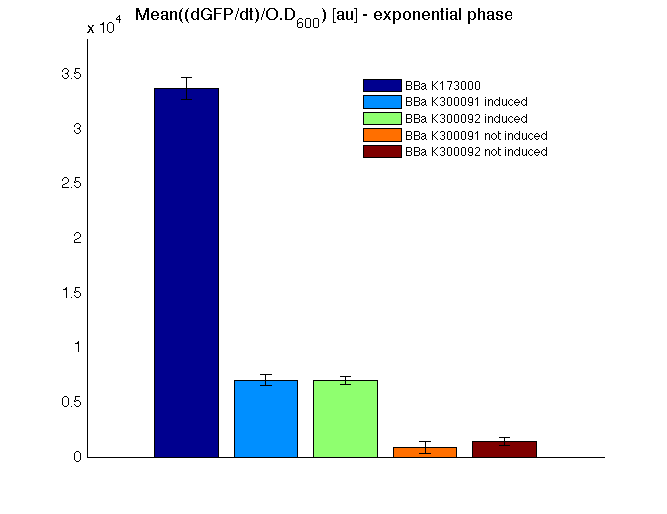 Mean (dGFP/dt)/O.D. over the exponential phase (under the hypothesis that GFP half-life in fusion contructs is similar to the original one -<partinfo>BBa_E0040</partinfo>) |
| Culture | Doubling time [min.] ± std error |
| <partinfo>BBa_K173000</partinfo> | 76.3336 ± 1.4362 |
<partinfo>BBa_K300091</partinfo>
induced | 121.1434 ± 7.0275 |
<partinfo>BBa_K300091</partinfo>
not induced | 74.4267 ± 1.3696 |
<partinfo>BBa_K300092</partinfo>
induced | 122.6088 ± 1.2785 |
<partinfo>BBa_K300092</partinfo>
not induced | 71.5105 ± 2.7113 |
| <partinfo>BBa_B0031</partinfo> | 70.8421 ± 2.2181 |
Discussion
All the cultures showed a similar growth curve; doubling time was computed as described here in order to obtain information about the burden due to the synthesis of such fusion proteins. It is possible to see that all doubling times are very similar except for induced cultures. In this case doubling time is much higher than both positive control and non-induced cultures; for this reason it is possible to assert that induction gives a high metabolic burden.
From GFP curve and mean protein synthesis rate it is possible to appreciate that induced <partinfo>BBa_K300091</partinfo> and <partinfo>BBa_K300092</partinfo> GFP accumulation profiles are comparable and they significantly differ from the GFP raw time series of the negative control <partinfo>BBa_B0031</partinfo>. On the other hand not induced <partinfo>BBa_K300091</partinfo> and <partinfo>BBa_K300092</partinfo> show a profile that is very similar to the negative control. These results show that the green fluorescent protein assembled downstream of the construct is correctly folded and that the inducible system works as expected.
Not induced <partinfo>BBa_K300091</partinfo> and <partinfo>BBa_K300092</partinfo> show a low GFP synthesis rate maybe due to 3OC6HSL inducible circuit leakage activity.
Submitted fusion bricks for self-cleaving affinity tag construction
The results shown above have demonstrated that the right folding can occur in a proof of concept fusion protein assembled with the newly designed synthetic affinity tags. However, the Phasin affinity with PHB granules and the self-cleavage behaviour of the Intein have not been tested yet.
Several BioBrick basic and composite parts have been submitted to the Registry to enable the construction of the desired composite affinity tags for protein purification using PHB. All these parts are compatible with the Silver fusion assembly to allow in-frame protein coding sequence assembly. A list of the designed and submitted affinity tags or self-cleavable affinity tags is reported below:
- <partinfo>BBa_K300002</partinfo> (DNA Sent)
- <partinfo>BBa_K300003</partinfo> (DNA Sent)
- <partinfo>BBa_K300004</partinfo> (DNA Sent)
- <partinfo>BBa_K300079</partinfo> (DNA Sent)
- <partinfo>BBa_K300080</partinfo> (DNA Sent)
- <partinfo>BBa_K300081</partinfo> (DNA Sent)
- <partinfo>BBa_K300083</partinfo> (DNA Sent)
- <partinfo>BBa_K300084</partinfo> (DNA Sent)
- <partinfo>BBa_K300093</partinfo>
- <partinfo>BBa_K300094</partinfo>
- <partinfo>BBa_K300095</partinfo>
- <partinfo>BBa_K300097</partinfo>
Registry users are welcome to use them to create and optimize protein purification through the binding of the target protein to PHB granules, thus engineering a simple and cheap purification system.
|  "
"