Existing Parts from the Registry: list
<partinfo>BBa_R0010</partinfo>, <partinfo>BBa_R0011</partinfo> - Wild type and hybrid lac promoters
<partinfo>BBa_R0011</partinfo> hybrid lac promoter and the <partinfo>BBa_R0010</partinfo> wild type lac promoter were characterized at different copy number in TOP10 E. coli strain. This strain contains a lacI expression system in the genome.
Induction static transfer function (computed in Relative Promoter Units), dynamics and metabolic burden were evaluated as a function of different IPTG concentrations in M9 supplemented with glycerol growth medium.
A RFP generator (<partinfo>BBa_I13507</partinfo>) was used as a reporter gene. In particular, these measurement systems were used:
- <partinfo>pSB1A2</partinfo>-<partinfo>BBa_J107010</partinfo>
- <partinfo>pSB1A3</partinfo>-<partinfo>BBa_J04450</partinfo>
- <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J107010</partinfo>
- <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J04450</partinfo>
At first, <partinfo>BBa_J107010</partinfo> and <partinfo>BBa_J04450</partinfo> inducibility was tested in a high copy number vector (<partinfo>pSB1A2</partinfo> or <partinfo>pSB1A3</partinfo>). The results are shown here as the relative RFP synthesis rate per cell.
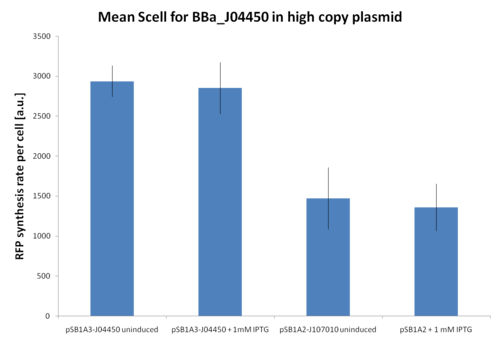 Relative RFP synthesis rate per cell in <partinfo>pSB1A2</partinfo>-<partinfo>BBa_J107010</partinfo> and <partinfo>pSB1A3</partinfo>-<partinfo>BBa_J04450</partinfo>. The error bars represent the standard errors of three independent measurements. |
Results show that in this condition <partinfo>BBa_R0010</partinfo> is about 2-fold stronger than <partinfo>BBa_R0011</partinfo>, but induced and uninduced cultures did not show differences in the RFP signal.
This result is expected because the vectors are propagated at about 200 copies per cell, while the lacI repressor is present at single copy in the genome and thus it is not able to repress the lac promoters in such high copy.
The doubling times and their standard errors estimated from data are reported below for <partinfo>pSB1A2</partinfo>-<partinfo>BBa_J107010</partinfo> and <partinfo>pSB1A3</partinfo>-<partinfo>BBa_J04450</partinfo> with and without 1mM of IPTG.
| Cultures | Mean doubling times [minutes] | standard errors over 3 independent experiment [minutes]
|
| <partinfo>pSB1A2</partinfo>-<partinfo>BBa_J107010</partinfo> | 77,7 | 3,1
|
| <partinfo>pSB1A2</partinfo>-<partinfo>BBa_J107010</partinfo> + 1mM IPTG | 76,5 | 2,2
|
| <partinfo>pSB1A3</partinfo>-<partinfo>BBa_J04450</partinfo> | 107,8 | 0,3
|
| <partinfo>pSB1A3</partinfo>-<partinfo>BBa_J04450</partinfo> + 1mM IPTG | 101,7 | 4,8
|
These results demonstrate that cells growth is not significantly affected by the presence of IPTG, even at the high of 1 mM.
<partinfo>BBa_J107010</partinfo> and <partinfo>BBa_J04450</partinfo> were then tested in the low copy (~5 copies per cell) vector <partinfo>pSB4C5</partinfo> in order to test their inducibility. The results are shown here as the RPU values at the steady state (constant RFP sysnthesis rate per cell) at different IPTG concentrations.
 RPU of <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J107010</partinfo> and <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J04450</partinfo> as a function of IPTG concentration. The error bars represent the standard errors of three independent measurements. |
Results show that in this condition both <partinfo>BBa_R0010</partinfo> and <partinfo>BBa_R0011</partinfo> produce different amounts of RFP as a function of the IPTG concentration. The amplitude of the two curves show that the promoters are very strong when induced with IPTG >= 10 uM. Although the experiments were carried out in the same conditions, the variability between experiments was high, especially for <partinfo>BBa_R0010</partinfo> (mean coefficient of variaton of about 37% for <partinfo>BBa_R0010</partinfo> and 15% for <partinfo>BBa_R0011</partinfo>), while the RPU variability between three wells in the same experiment is much lower (mean coefficient of variaton of bout 3.5% for both promoters).
The above figure shows that <partinfo>BBa_R0011</partinfo> is stronger than the <partinfo>BBa_R0010</partinfo> wild type promoter in low copy plasmid. This result is unexpected because the same promoters in high copy vectors behaved differently (<partinfo>BBa_R0010</partinfo> was stronger than the <partinfo>BBa_R0011</partinfo>, see above).
In the uninduced state, <partinfo>BBa_R0011</partinfo> has about the same strength as the <partinfo>BBa_J23101</partinfo> reference standard promoter.
This static characteristic shows that the promoters are both leaky and a very low IPTG concentration (10 uM) is sufficient to trigger gene expression at *very* high levels.
These results demonstrate that the genomic lacI is partially able to repress the two promoters, but very low IPTG concentrations are sufficient to bind the repressor and trigger the promoters transcription.
Doubling times were also estimated for these cultures. Their values are reported below for uninduced and 1 mM IPTG-induced cultures.
| Cultures | Mean doubling times [minutes] | standard errors over 3 independent experiments [minutes]
|
| <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J107010</partinfo> | 113,5 | 10,8
|
| <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J107010</partinfo> + 1mM IPTG | 106,8 | 5,5
|
| <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J04450</partinfo> | 85 | 5
|
| <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J04450</partinfo> + 1mM IPTG | 90 | 4,5
|
As obtained for the cultures with high copy plasmids, the growth rate of TOP10 harbouring low copy vectors with the measurement parts is not affected by IPTG presence.
Dynamic characterization in low copy vector: The figure below shows a typical relative RFP synthesis rate per cell time series for <partinfo>BBa_J107010</partinfo> and <partinfo>BBa_J04450</partinfo> induced with 1 mM of IPTG and uninduced. These time series show that the full induction can be reached after about 50 min from the induction.
 Mean Scell signal as a function of time for <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J107010</partinfo> and <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J04450</partinfo>. Induced (with 1 mM of IPTG) and uninduced cultures are shown. Induction occurs at t=0. The shown graph is relative to one of the three experiments performed in different days. |
Conclusion: the characterization of two IPTG-inducible promoters has been performed and the performance of these two promoters have been compared in terms of transcriptional strength. The reported results are easily sharable in different laboratories thanks to the used standard RPU approach.
Methods:
- A of long term storage glycerol stock was streaked on a LB plate with suitable antibiotic. Tha plate was incubated overnight at 37°C.
- A single colony was inoculated in 1 ml of M9 + suitable antibiotic in a 15 ml tube and incubated at 37°C, 220 rpm for about 16 hours.
- The grown cultures were then diluted 1:100 in 2-5 ml of M9 supplemented medium and incubated in the same conditions as before for about 4-5 hours.
- For each desired IPTG concentration to be tested, three 200 ul aliquots of the cultures were aliquoted in a flat-bottom 96-well microplate, avoiding to perform dynamic experiments in the microplate frame (in order to prevent evaporation effects in the frame).
- 2 ul of properly diluted IPTG (Sigma Aldrich) were added to the three wells for each desired concentration.
- The microplate was incubated in the Tecan Infinite F200 microplate reader and fluorescence and absorbance were measured with this automatic protocol:
- 37°C constant for all the experiment;
- sampling time of 5 minutes;
- fluorescence gain of 50 or 80;
- O.D. filter at 600 nm;
- RFP filters at 535nm (ex) / 620nm (em);
- 15 seconds of linear shaking (3mm amplitude) followed by 10 seconds of waiting before the measurements in order to make a homogeneous culture.
- Experiment duration time: about 6 hours.
- This experiment was performed three times in different days.
Data analysis: Relative Promoter Units (RPUs) were estimated as described by [Kelly JR et al. (2009), J Biol Eng 3:4].
Briefly:
- Absorbance and fluorescence time series were normalized by subtracting the absorbance of the media and the fluorescence of a negative control (a non fluorescent TOP10 culture) respectively, thus yielding O.D.600 and RFP time series.
- RFP synthesis rate per cell (called Scell) was computed as (1/O.D.600)*dGFP/dt. (this signal is not actually the RFP synthesis rate, but is proportional to it).
- The RFP synthesis rate per cell was averaged at the steady state during the exponential growth phase (validated by identifying the linear region of the ln(O.D.600)).
- The RPU of the promoter of interest in a specific condition was computed as mean_Scell,phi/mean_Scell,J23101 where phi is the promoter of interest, J23101 is the reference standard and mean_Scell is the mean Scell signal value, computed as explained above.
<partinfo>BBa_K300009</partinfo>/<partinfo>BBa_I4102</partinfo> - PoPS->3OC6HSL sender device
<partinfo>BBa_F2620</partinfo> - 3OC6HSL -> PoPS Receiver
<partinfo>BBa_J61001</partinfo> - R6K Origin of replication
<partinfo>BBa_K300008</partinfo> was used as a validation construct for this conditional replication origin, in order to test its capability to be propagated in pir+ or pir-116 strain and its inability to propagate in the other E. coli strains.
In particular, <partinfo>BBa_K300008</partinfo> was cut with XbaI-SpeI and the insert was isolated and purified from a 1% agarose gel. Then, it was self-ligated to generate a Cm-resistant R6K plasmid).
BW25141 (<partinfo>BBa_K300984</partinfo>) and BW23474 (<partinfo>BBa_K300985</partinfo>) were chosen as pir+ and pir-116 strains respectively, while DH5alpha (<partinfo>BBa_V1001</partinfo>), MC1061 (<partinfo>BBa_K300078</partinfo>) and MG1655 (<partinfo>BBa_V1000</partinfo>) were chosen as pir- strains.
All these strains were made competent following the commonly used CaCl2 method [Sambrook J, Fritsch EF, and Maniatis T (1989), Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.]. Then, a vial of 100 ul of competent cells was transformed with 2-4 ng of:
- no DNA (negative control);
- a pSB*** series vector (positive control);
- self-ligated <partinfo>BBa_K300008</partinfo>.
and plated on LB+Cm at 34 ug/ml for high-copy plasmids, Cm at 12.5 ug/ml for medium/low copy plasmids and for the negative control strains transformed with the R6K plasmids.
The colonies were counted in each plate and the transformation efficiency was estimated in [CFU/ug of DNA] as:
efficiency [CFU/ug of DNA]= # CFU * 1000 ng of DNA / amount of transformed DNA [ng]
The results are shown here:
| Strain
| Efficiency with no DNA
| Efficiency with pSB*** (positive control)
| Efficiency with the self-ligated <partinfo>BBa_K300008</partinfo> (R6K plasmid)
|
| <partinfo>BBa_K300084</partinfo>
| 0
| 10^5
| 10^5
|
| <partinfo>BBa_K300085</partinfo>
| 0
| 10^6
| 10^6
|
| <partinfo>BBa_V1001</partinfo>
| 0
| 10^8
| 0
|
| <partinfo>BBa_K300078</partinfo>
| 0
| 10^6
| 0
|
| <partinfo>BBa_V1000</partinfo>
| 0
| 10^5
| 0
|
These results show that <partinfo>BBa_J61001</partinfo> replication origin can be only propagated in pir+ and pir-116 strains (<partinfo>BBa_K300084</partinfo> and <partinfo>BBa_K300085</partinfo>), while the transformation of other strains with the R6K plasmid yielded no colonies after transformation.
Moreover, these results show that the R6K plasmid in pir+ and pir-116 strains was transformed with the same efficiency as the pSB*** positive control plasmid, demonstrating that the R6K origin doesn't give any handicap in plasmid transformation.
<partinfo>BBa_J23100</partinfo>, <partinfo>BBa_J23101</partinfo>, <partinfo>BBa_J23105</partinfo>, <partinfo>BBa_J23106</partinfo>, <partinfo>BBa_J23110</partinfo>, <partinfo>BBa_J23114</partinfo>, <partinfo>BBa_J23116</partinfo>, <partinfo>BBa_J23118</partinfo> - constitutive promoters from Anderson's collection
The <partinfo>BBa_J23100</partinfo>, <partinfo>BBa_J23101</partinfo>, <partinfo>BBa_J23105</partinfo>, <partinfo>BBa_J23106</partinfo>, <partinfo>BBa_J23110</partinfo>, <partinfo>BBa_J23114</partinfo>, <partinfo>BBa_J23116</partinfo>, <partinfo>BBa_J23118</partinfo> were charcterized in LB and M9 supplemented with glycerol (0.4%) growth media in high copy and low copy vectors in E. coli TOP10 (<partinfo>BBa_V1009</partinfo>).
RPU and doubling time were characterized for all of them, according to the protocols reported in this section.
The following measurement systems were used for high copy plasmids:
- <partinfo>BBa_J61002</partinfo>-<partinfo>BBa_J23100</partinfo>
- <partinfo>BBa_J61002</partinfo>-<partinfo>BBa_J23101</partinfo>
- <partinfo>BBa_J61002</partinfo>-<partinfo>BBa_J23105</partinfo>
- <partinfo>BBa_J61002</partinfo>-<partinfo>BBa_J23106</partinfo>
- <partinfo>BBa_J61002</partinfo>-<partinfo>BBa_J23110</partinfo>
- <partinfo>BBa_J61002</partinfo>-<partinfo>BBa_J23114</partinfo>
- <partinfo>BBa_J61002</partinfo>-<partinfo>BBa_J23116</partinfo>
- <partinfo>BBa_J61002</partinfo>-<partinfo>BBa_J23118</partinfo>
In order to build low copy plasmid measurement systems, the EcoRI-PstI fragment (J231xx-RFP) of each <partinfo>BBa_J61002</partinfo>-BBa_J231xx was assembled into <partinfo>pSB4C5</partinfo> vector. This fragment contains the constitutive promoter of interest upstream a RBS-RFP-TT expression system.
The following measurement parts were used for low copy plasmids:
- <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J23100</partinfo>
- <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J23101</partinfo>
- <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J23105</partinfo>
- <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J23106</partinfo>
- <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J23110</partinfo>
- <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J23114</partinfo>
- <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J23116</partinfo>
- <partinfo>pSB4C5</partinfo>-<partinfo>BBa_J23118</partinfo>
The RPU values and doubling times are here reported:
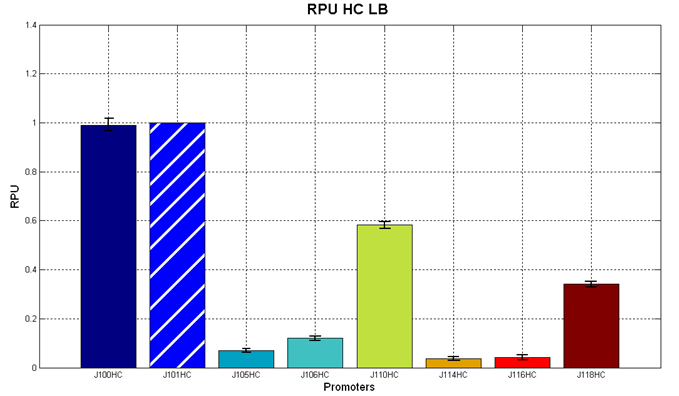 Figure 5 - R.P.U. of the studied promoters from Anderson promoters' collection, LB medium and high copy plasmid (<partinfo>BBa_J61002</partinfo>) | 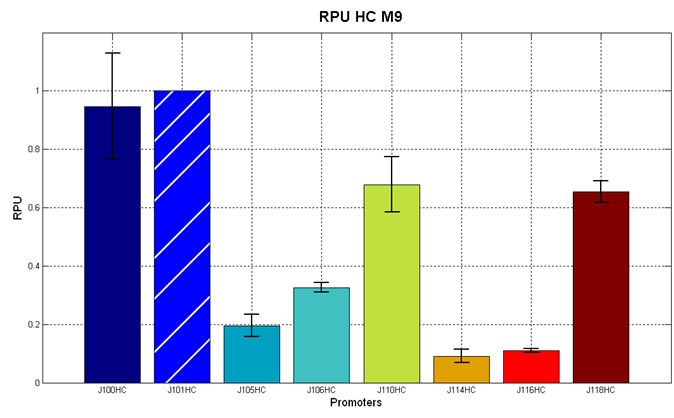 Figure 6 - R.P.U. of the studied promoters from Anderson promoters' collection, M9 medium and high copy plasmid (<partinfo>BBa_J61002</partinfo>) |
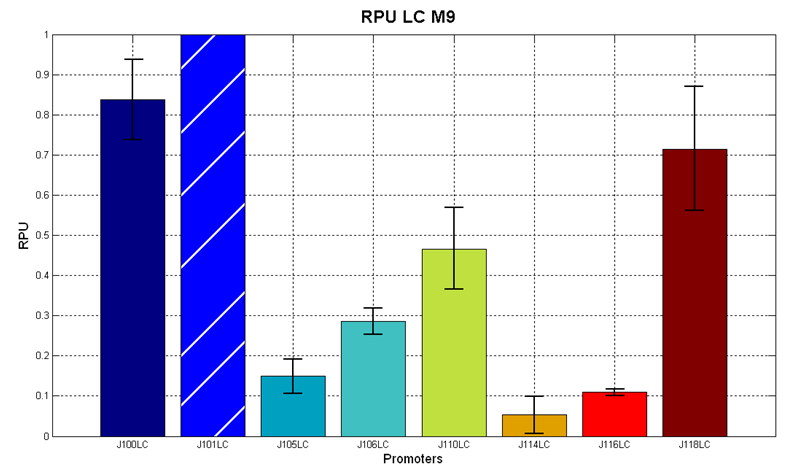 Figure 7 - R.P.U. of the studied promoters from Anderson promoters' collection, M9 medium and low copy plasmid (<partinfo>pSB4C5</partinfo>). These plasmids were constructed by assembling the EcoRI-PstI fragment of <partinfo>BBa_J61002</partinfo>-BBa_J231xx in <partinfo>pSB4C5</partinfo> vector, in order to transfer the promoter and the RBS-RFP-TT expression construct from <partinfo>BBa_J61002</partinfo> to <partinfo>pSB4C5</partinfo>. |
The error bars represent the standard deviation for three dfferent wells in the same experiment.
Doubling times were evaluated for the described cultures (HC stands for High Copy and LC stands for Low Copy):
| Promoter
| doubling time [minutes]
|
| LB in HC plasmid | M9 in HC plasmid | M9 in LC plasmid
|
| <partinfo>BBa_J23100</partinfo> | 33.75
±
1.34 | 82.53
±
2.45 | 86.11
±
4.45
|
| <partinfo>BBa_J23101</partinfo> | 35.93
±
0.62 | 82.68
±
1.84 | 86.42
±
1.91
|
| <partinfo>BBa_J23105</partinfo> | 29.86
±
0.33 | 63.09
±
7.08 | 85.00
±
5.13
|
| <partinfo>BBa_J23106</partinfo> | 29.17
±
0.96 | 68.11
±
4.25 | 88.71
±
0.90
|
| <partinfo>BBa_J23110</partinfo> | 31.28
±
0.42 | 67.52
±
5.87 | 76.15
±
2.16
|
| <partinfo>BBa_J23114</partinfo> | 28.97
±
0.49 | 59.44
±
5.20 | 80.12
±
0.95
|
| <partinfo>BBa_J23116</partinfo> | 28.14
±
0.25 | 72.74
±
0.37 | 81.68
±
3.08
|
| <partinfo>BBa_J23118</partinfo> | 32.84
±
0.31 | 73.64
±
2.41 | 89.86
±
2.93
|
It was not possible to evaluate promoters activities in low copy number plasmids in LB because the RFP activity was too weak and not distinguishable from the background.
Discussion: we observed that the ranking previously documented in the Registry is not valid in all the tested conditions, even if a general agreement can be observed. As an example, <partinfo>BBa_J23110</partinfo> in high copy plasmid is stronger than <partinfo>BBa_J23118</partinfo>, in contrast with the ranking reported in the Registry.
<partinfo>BBa_P1004</partinfo> - chloramphenicol resistance cassette
<partinfo>BBa_P1004</partinfo> has been successfully used in the assembly of <partinfo>BBa_K300000</partinfo> integrative base vector for E. coli. All the intermediate parts which contained <partinfo>BBa_P1004</partinfo> showed Chloramphenicol resistance (tested up to 34 ug/ml in LB media) when transformed in TOP10 (<partinfo>BBa_V1009</partinfo>), DH5alpha (<partinfo>BBa_V1001</partinfo>), MC1061 (<partinfo>BBa_K300078</partinfo>), MG1655 (<partinfo>BBa_V1000</partinfo>), BW23474 (<partinfo>BBa_K300985</partinfo>) and DB3.1 (<partinfo>BBa_V1005</partinfo>) strains in a high-copy plasmid.
<partinfo>BBa_K125500</partinfo> - GFP fusion brick
This part can be useful to construct fluorescent fusion proteins. It is composed by a tail domain (the GFP <partinfo>K125500</partinfo>) with a transcriptional terminator (<partinfo>BBa_B0015</partinfo>) downstream.
Other protein domains can be fused upstream of this part in order to create chimeric fluorescent proteins, or it can be ligated to tags useful for low-cost protein purification.
This part was used do design the following BioBrick measurement systems:
- <partinfo>BBa_K300086</partinfo>
- <partinfo>BBa_K300088</partinfo>
- <partinfo>BBa_K300090</partinfo>
- <partinfo>BBa_K300091</partinfo>
- <partinfo>BBa_K300092</partinfo>
- <partinfo>BBa_K300099</partinfo>
to test these contructs:
- <partinfo>BBa_K300002</partinfo>
- <partinfo>BBa_K300093</partinfo>
- <partinfo>BBa_K300094</partinfo>
- <partinfo>BBa_K300095</partinfo>
- <partinfo>BBa_K300084</partinfo>
- <partinfo>BBa_K300097</partinfo>
respectively. All of the tested parts are synthetic fusion tags whose activity could be measured by assembling a promoter with RBS upstream and a tail domain with terminator downstream, thus yielding the measurement systems. <partinfo>BBa_K300005</partinfo> was used as a tail domain with terminator downstream. It has been assembled to test the correct folding of the resulting fusion protein by measuring the GFP and to test the affinity tag performance with a proof of concept protein.
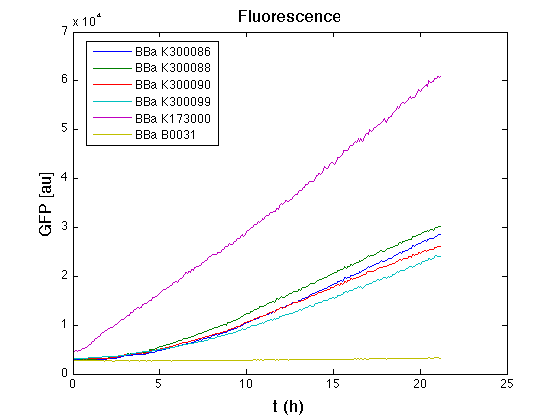
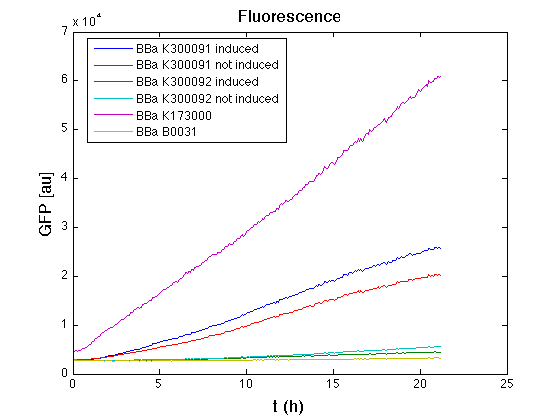
In all of the measurement parts, GFP could be successfully detected in bacteria harbouring the measurement parts in high copy plasmids. In this condition, GFP was detected by using an excitation filter at 485nm and an emission filter at 540nm in a Infinite F200 microplate reader (Tecan).
|
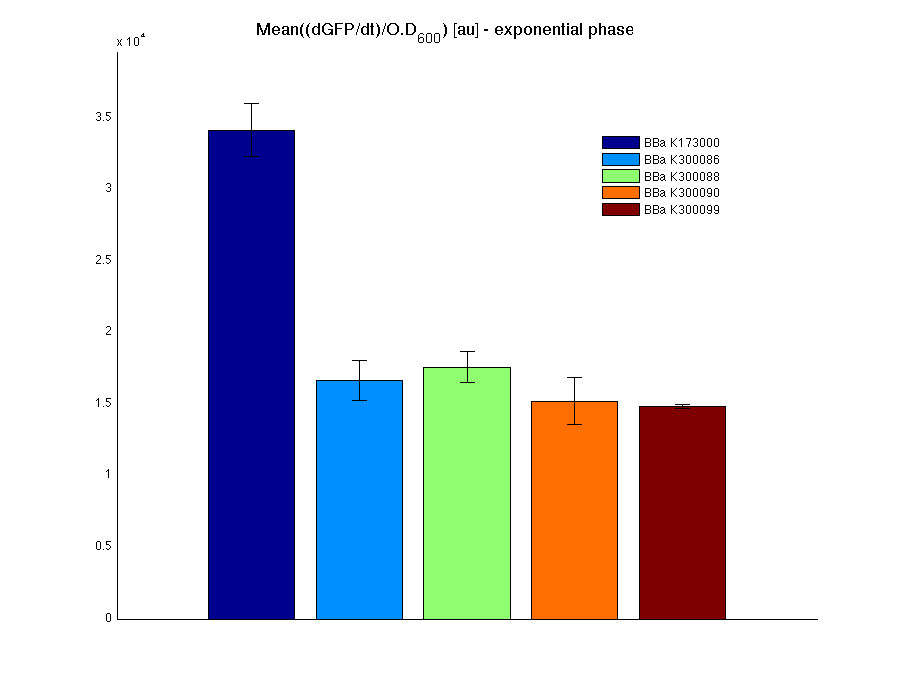 Mean (dGFP/dt)/O.D. over the exponential phase (under the hypothesis that GFP half-life in fusion contructs is similar to the original one - <partinfo>BBa_E0040</partinfo>) |
|
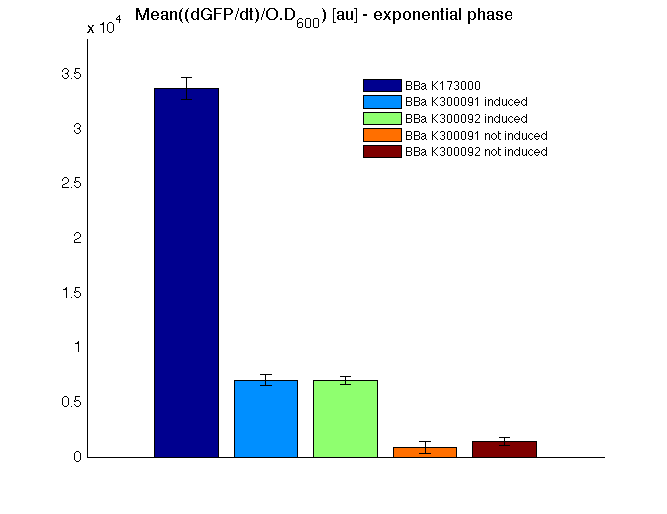 Mean (dGFP/dt)/O.D. over the exponential phase (under the hypothesis that GFP half-life in fusion contructs is similar to the original one - <partinfo>BBa_E0040</partinfo>) |
<partinfo>BBa_J72008</partinfo> - phi80 integration helper plasmid pInt80-649
<partinfo>BBa_J72008</partinfo> has been successfully used in the integration protocol of both MC1061 (<partinfo>BBa_K300078</partinfo>) and MG1655 (<partinfo>BBa_V1000</partinfo>) E. coli strains. See [http://partsregistry.org/Part:BBa_K300000:Experience BBa_K300000 Experience page] for details about how this plasmid was used.
It can be actually cured at 37-42°C, while it can be propagated at 30°C.
It actually enables the propagation of R6K (conditional replication origin) plasmids, thanks to its pir-116 gene.
|  "
"