Team:Toronto/Design
From 2010.igem.org
| Home | Project | Design | Protocols | Notebook | Results | Parts Submitted to the Registry | Modeling | Software Used | Human Practices | Safety | Team | Official Team Profile | FAQ | Acknowledgments |
|---|
Project Design
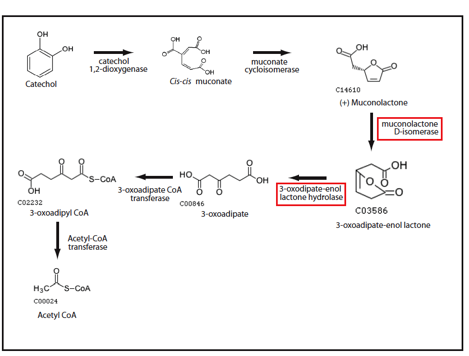
In order to identify enzymes in the catechol degradation pathway (ortho cleavage) that benefit from the introduction of metabolic channelling, we performed in silico analyses with software developed in our lab (Sanford et al.). Preliminary results from our simulations suggests that the 3rd (muconolactone D-isomerase) and 4th (3-oxodipate-enol lactone hydrolase) enzymes in the pathway would benefit most from channelling (Figure 1). Muconolactone D-isomerase catalyzes an energetically unfavourable reaction, while 3-oxodipate-enol lactone hydrolase catalyzes a very energetically favourable reaction. By introducing channeling between these two enzymes, the energetically unfavourable product of the mucanolactone D-isomerase-catalyzed reaction would be converted to the product of 3-oxodipate-enol lactone hydrolase before mucanolactone D-isomerase converts the energetically unfavourable product back into the energetically favourable substrate. The net effect would be an increased flux through the catechol degradation pathway.
Figure 1: Ortho degradation of catechol (1,2-Benzenediol) proceeds in six steps. Only the last enzyme is present in E. coli. Enzyme candidates with the largest predicted channeling effect as predicted by Cell++ are boxed in red.
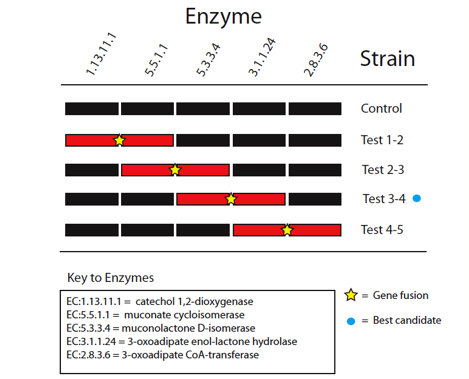
The five enzymes in the catechol degradation pathway not naturally present in E. coli will be introduced into E. coli as BioBrick parts. The parts will then be used to construct five recombinant strains of E. coli (Figure 2). These include a full reconstruction of the pathway (control strain) and several partial reconstructions (test strains). The latter will contain pathway 'holes' corresponding to all combinations of sequential pairs of enzymes, with complementary fusion pairs. The test strain with the fusion of enzymes 3 and 4 will be of prime interest, while the other test strains will be used validate the predictions made by Cell++.
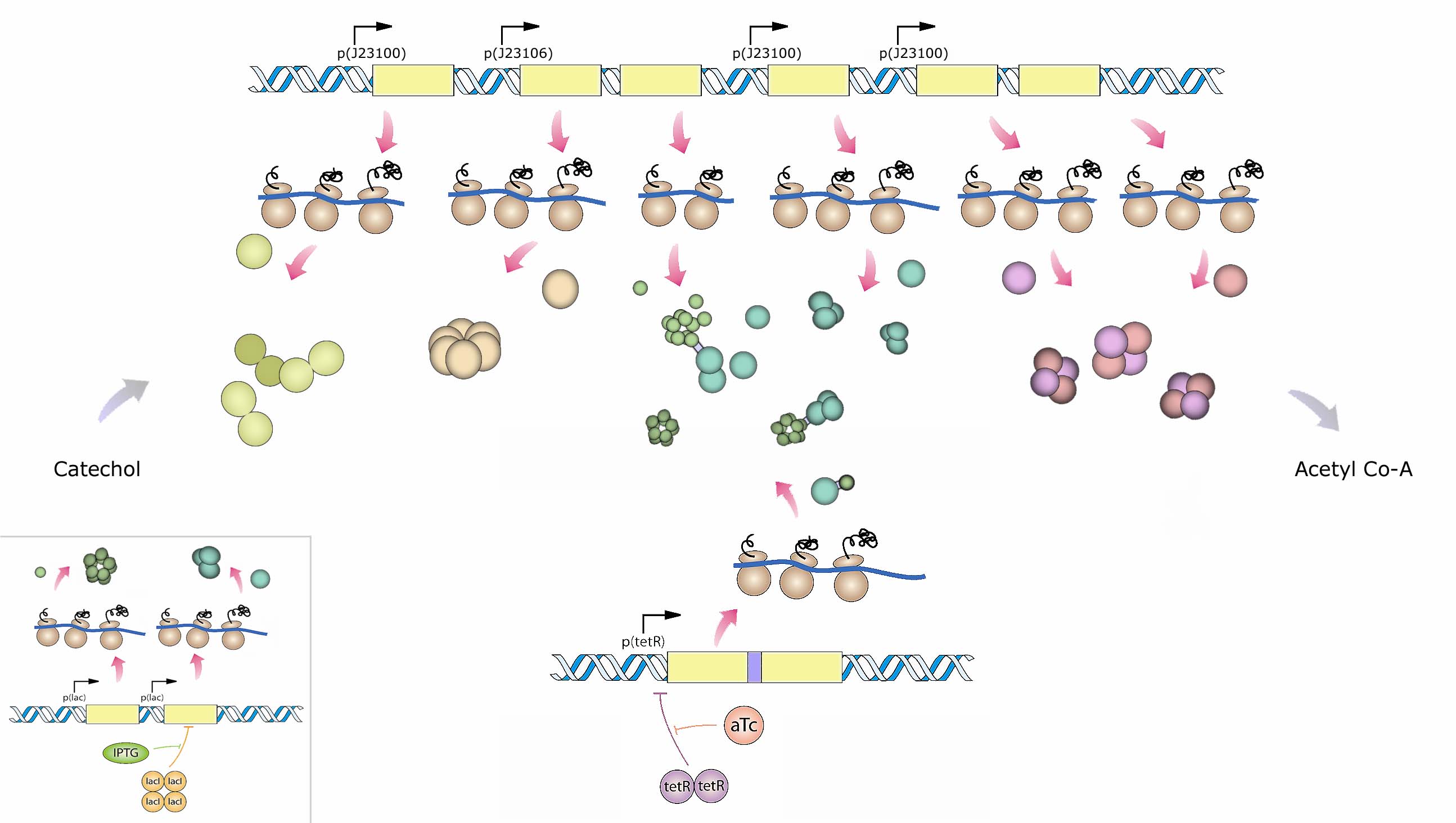
Many of the enzymes in the system are multimeric. For this reason we propose expressing the fusion proteins alongside their native (un-fused) counterparts to promote linkage of the enzymes while allowing for assembly of unlinked monomers onto the fusions which we refer to as 'seeds'. The relative expression of free monomers and fusion seeds is predicted to be important since too many seeds relative to free monomers will lead to tangles whereas too many free monomers relative to seeds will ablate any channeling effects. The assembly dynamics and desired relative expression ratios of seeds and free monomers are being explored using in silico modeling as described in the modeling section. Below is a diagramatic representation of the proposed expression system (Figure 3). The exact choice of promoters will depend upon predictions from our modeling results.
Figure 2: Diagramatic representation of proposed gene fusion experiments. Pathway reconstructions are shown in black for the control and test strains with pathway holes for the partial reconstructions shown in red. Yellow stars mark proposed gene fusions to be tested to validate computational predictions. The blue circle marks the best candidate for enzyme channeling based on in silico experiments
In order to demonstrate the breakdown of catechol by these recombinant strains, catechol will be added into the growth media of these strains. A colorimetric assay, previously developed to quantify the amount of catechol in solution, will be utilized to assay the rate of catechol degradation by the different strains (Tao et al.). In addition, a survival assay will be used to determine the threshold for catechol exposure for non recombinant E. coli and to evaluate the effect that the recombinantly-expressed enzymes in the catechol pathway will have on E. coli survival.
Figure 3: An overview of the recombinant fusion construct and experimental system. Muconolactone D-isomerase (enzyme 3; green) and 3-oxodipate-enol lactone hydrolase (enzyme 4; sky blue) subunits will be linked via an alpha-helical linker. The linker is expected to serves a dual purpose: 1) The fusion product seeds the formation of multimeric proteins required for enzymatic function, and 2) The linker spatially localizes the two enzymes to aid channeling. The fusion construct (terR) along with a redundant pair of non-fused enzymes 3 and 4 (inset; lac) are placed under different inducible promoters. This allows for the assessment of potential channeling effects by selectively inducing the fusion protein relative to the induction of redundant copies of the same un-fused enzymes.
 "
"