Team:TU Delft/Modeling/interaction-mapping
From 2010.igem.org
(New page: __NOTOC__ ==Problems with introducing new genes== In our project, we are taking genes from various organisms and putting them as a biobrick plasmid into E. coli. This may cause various un...) |
|||
| Line 77: | Line 77: | ||
==Software development== | ==Software development== | ||
An application was developed that can run these queries on a PostgreSQL server running the STRING database. This database can be [http://string-db.org/newstring_cgi/show_download_page.pl downloaded] for free for academic use. Unfortunately, the STRING API does not provide enough functionality to do the homolog mapping, so a PostgreSQL database with the STRING data is always needed. | An application was developed that can run these queries on a PostgreSQL server running the STRING database. This database can be [http://string-db.org/newstring_cgi/show_download_page.pl downloaded] for free for academic use. Unfortunately, the STRING API does not provide enough functionality to do the homolog mapping, so a PostgreSQL database with the STRING data is always needed. | ||
| - | Source | + | |
| + | Source code is available in a [http://github.com/jcnossen/InteractionHomologMapping github repository], or by directly downloading the [http://github.com/jcnossen/InteractionHomologMapping/zipball/master code in zip format] | ||
==Result== | ==Result== | ||
| - | http://dl.dropbox.com/u/602630/cyweb2.htm | + | Work in progress. See http://dl.dropbox.com/u/602630/cyweb2.htm for preliminary results. |
Revision as of 15:35, 10 September 2010
Problems with introducing new genes
In our project, we are taking genes from various organisms and putting them as a biobrick plasmid into E. coli. This may cause various unexpected problems in the newly designed biological system:
- The protein in the original organism may need some other functional partners (proteins or RNAs) in order to behave correctly (I.A.W. a part of the imported system is missing)
- The introduced protein can interact with local proteins that are not ment to interact.
Searching for interactions
To get an idea of which interactions may occur, interactions within the original organisms where mapped to their E. coli homologs:
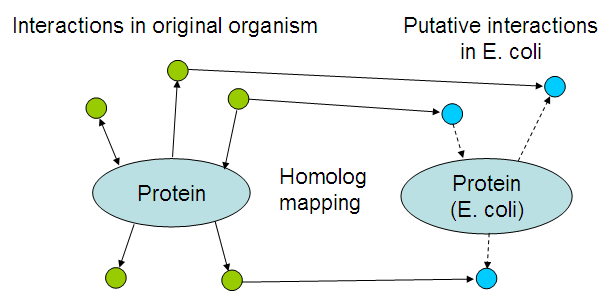
In more detail:
- The sequence of every introduced protein (See the coding biobricks) was blasted against the STRING protein database. This resulted in a number of proteins that are either the correct proteins, or homologs of the introduced proteins.
| Biobrick | Short name | STRING Database ID |
| BBa_K398000 | LadA | 420246.GTNG_3499 |
| BBa_K398001 | AlkB | 101510.RHA1_ro02534 |
| BBa_K398002 | RubA3 | 246196.MSMEG_1840 |
| BBa_K398003 | RubA4 | 350058.Mvan_1744 |
| BBa_K398004 | Rubredoxin reductase | 101510.RHA1_ro02537 |
| BBa_K398005 | Bt-ADH | 235909.GK0984 |
| BBa_K398006 | Bt-ALDH | 235909.GK2772 |
| BBa_K398100 | bbc1 | Dropped: Protein sequences blast only resulted in eukaryotic matches |
| BBa_K398200 | AlnA | 62977.ACIAD0697 |
| BBa_K398201 | OprG | 160488.PP_0504 |
| BBa_K398300 | AlkS | 399739.Pmen_0429 |
| BBa_K398400 | Prefoldin-alpha | 70601.PH0527 |
| BBa_K398401 | Prefoldin-beta | 70601.PH0532 |
- For each protein, the interactions stored in STRING where gathered.
- For each found interacting protein, E. coli homologs where gathered.
- This results in a combination of interactions with their E. coli homologs. The likelyness of this being a true interaction depends on the STRING interaction score (Data from publication text mining, microarrays and other sources) and the BLAST score (The homolog mapping step).
Software development
An application was developed that can run these queries on a PostgreSQL server running the STRING database. This database can be downloaded for free for academic use. Unfortunately, the STRING API does not provide enough functionality to do the homolog mapping, so a PostgreSQL database with the STRING data is always needed.
Source code is available in a github repository, or by directly downloading the code in zip format
Result
Work in progress. See http://dl.dropbox.com/u/602630/cyweb2.htm for preliminary results.
 "
"