Team:Newcastle/12 August 2010
From 2010.igem.org

| |||||||||||||
| |||||||||||||
Contents |
rocF Gibson Cloning
Discussion
Today we have colonies from our transformation yesterday. Overnight cultures will be done for DNA extraction tomorrow.
Subtilin Immunity BioBrick
Aims
The aims for today are to perform gel electrophoresis again, and then gel extraction.
Materials and protocol
Please refer to the gel electrophoresis and the gel extraction protocols.
Results
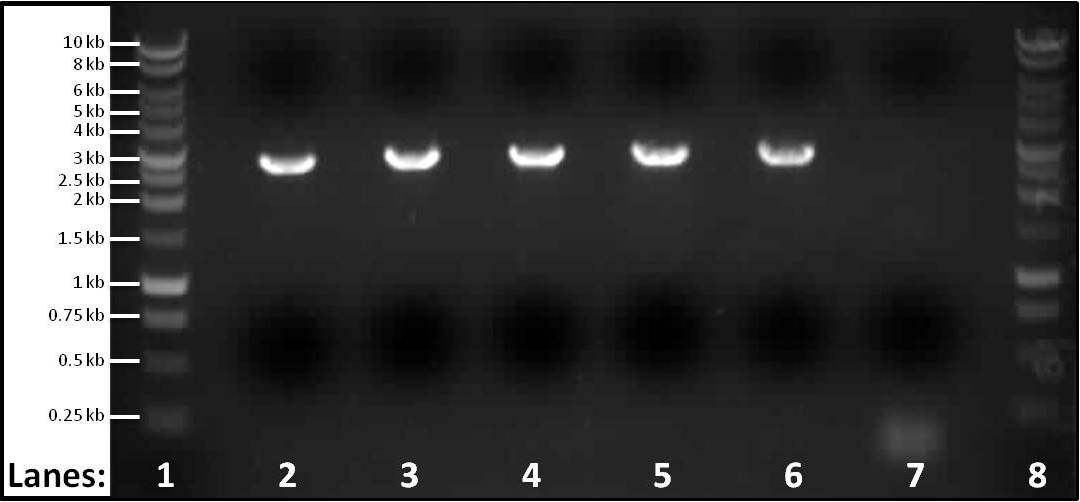
Figure #: Gel electrophoresis of the PCR products of part 3 spaIFEG gene cluster required for the subtilin immunity BioBrick. The PCR reactions were performed at 5 different temperatures, Tms, and a control PCR reaction was also carried out. Please see below for the specific temperatures and extension time of each tube. The primers used for lanes 2-6 were the primer 1-S1for(B) and primer 2-S1rev(B). For the control the ara forward and reverse primers were used.
- Lane 1: 1 kb DNA ladder
- Lane 2: spaIFEG Gene Cluster - Tm: 46°C, extension time: 110 seconds
- Lane 3: spaIFEG Gene Cluster - Tm: 51°C, extension time: 110 seconds
- Lane 4: spaIFEG Gene Cluster - Tm: 56°C, extension time: 110 seconds
- Lane 5: spaIFEG Gene Cluster - Tm: 61°C, extension time: 110 seconds
- Lane 6: spaIFEG Gene Cluster - Tm: 66°C, extension time: 110 seconds
- Lane 7: Control - Tm of 59°C, extension time 15 seconds
- Lane 8: 1 kb DNA ladder
Discussion
We found that the PCR tube containing the spaIFEG gene (part 3) did not have any band, even though another gel electrophoresis was performed (at 63°C) again. We decided to perform five different PCR reactions at different temperatures: 46°C, 51°C, 56°C, 61°C and 66°C - this is to check whether the missing band was due to the incorrect Tm. We also performed a sixth PCR tube as a control, which contained the ara forward and reverse primers (we previously had extracted from Bacillus subtilis ATCC 6633 on 7th July 2010) - the Tm for this was 59°C, with the extention time of fifteen seconds (specific for ara primers). ara primers were put into the tube because they were the primers that confirmed that the chromosomal DNA extraction from Bacillus subtilis ATCC 6633 had worked.
We have cut the Plasmid Vector (part 1), Promoter & RBS (part 2) and Double terminator (part 4) gel bands following the gel extraction protocol. We will wait until the spaIFEG gene (part 3) has been cut from the gel. This will be carried out tommorrow. After all four parts have been cut from gels, the gel extraction protocol will be completed and the concentration of the DNA will be tested using the NanoDrop Spectrophotometer. Once all four parts have been successfully extracted, a single PCR tube will be prepared for the final step for the Gibson cloning.
Conclusion
We have successfully extracted part 3 (spa gene) by seeing all the bands in the gel electrophoresis. The bands were all the same, even though the Tms were different (ranged from 46°C - 66°C. (Link)
Go back to our main Lab book page
 
|
 "
"