Team:Michigan/Pili August September
From 2010.igem.org
(New page: {{Michigan Header}} {|style="color:#1c2bf2;background-color:#fafa19;font-size:9pt;text-align:center" cellpadding="5" cellspacing="0" border="1" bordercolor="#fff" width="62%" |- ! |Sunday...) |
|||
| Line 19: | Line 19: | ||
| - | | - | ||
| - | | - | ||
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#8/7/2010|8/7/2010]] |
|- | |- | ||
!Week 7 | !Week 7 | ||
| - | | - | ||
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#8/9/2010|8/9/2010]] |
| - | | - | ||
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#8/11/2010|8/11/2010]] |
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#8/12/2010|8/12/2010]] |
| - | | - | ||
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#8/14/2010|8/14/2010]] |
|- | |- | ||
!Week 8 | !Week 8 | ||
| - | | - | ||
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#8/16/2010|8/16/2010]] |
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#8/17/2010|8/17/2010]] |
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#8/18/2010|8/18/2010]] |
| - | | - | ||
| - | | - | ||
| Line 40: | Line 40: | ||
|- | |- | ||
!Week 9 | !Week 9 | ||
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#8/22/2010|8/22/2010]] |
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#8/23/2010|8/23/2010]] |
| - | | - | ||
| - | | - | ||
| Line 53: | Line 53: | ||
| - | | - | ||
| - | | - | ||
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#9/2/2010|9/2/2010]] |
| - | | - | ||
| - | | - | ||
| Line 60: | Line 60: | ||
| - | | - | ||
| - | | - | ||
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#9/7/2010|9/7/2010]] |
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#9/8/2010|9/8/2010]] |
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#9/9/2010|9/9/2010]] |
| - | | [[Team:Michigan/ | + | | [[Team:Michigan/Pili_August_September#9/10/2010|9/10/2010]] |
| - | | - | ||
|- | |- | ||
| Line 85: | Line 85: | ||
|- | |- | ||
|} | |} | ||
| + | |||
| + | ==8/7/2010== | ||
| + | ''Kevin, Marc, Alena'' | ||
| + | |||
| + | PCR #1 | ||
| + | Used a gradient from 40C to 60C for the first 3 cycles to find the optimum anneling temperature. | ||
| + | |||
| + | All of the annealing temperatures gave a good result according to the gel. | ||
| + | |||
| + | [[media:Lab10.pdf|8/7/10 notes]] | ||
| + | |||
| + | ==8/9/2010== | ||
| + | |||
| + | ''Kevin, Marc, Alena'' | ||
| + | |||
| + | Used a 57C degree annealing temperature to get enough DNA for the digest and ligation. | ||
| + | |||
| + | 4 out of the 5 PCR reactions worked well according to the gel. | ||
| + | |||
| + | The 5th well could have been a loading problem or there wasn't enough DNA. | ||
| + | |||
| + | [[media:Lab11.pdf|8/9/10 notes]] | ||
| + | |||
| + | ==8/11/2010== | ||
| + | |||
| + | ''Kevin, Marc'' | ||
| + | |||
| + | Met with Chris, received advice for updating digest and ligation protocols. | ||
| + | |||
| + | ==8/12/2010== | ||
| + | |||
| + | ''Kevin, Marc, Alena'' | ||
| + | |||
| + | Met and discussed protocols for digestion and ligation of FIMB into pBAD. | ||
| + | |||
| + | 1. Added 5 mL of LB broth each to 2 50 mL falcon tubes in the ERB lab using sterile technique. | ||
| + | |||
| + | 2. Added 5 microliters of Kanamycin to each of the 50 mL tubes in step 1. | ||
| + | |||
| + | Went to the budget committee meeting for 1 hour with the tubes. | ||
| + | |||
| + | 3. Obtained the cryostock of pBAD from the Lin -80C freezer (iGEM box cell #73) | ||
| + | |||
| + | 4. Stabbed cryostock using a sterile 200 microliter pipette tip and pipetted into media from step 2. | ||
| + | |||
| + | 5. At 8:05PM placed the two falcon tubes from step 4 into the incubator/shaker at 30C. | ||
| + | |||
| + | ==8/14/2010== | ||
| + | ''Kevin, Marc, Alena'' | ||
| + | |||
| + | '''Miniprep pBAD plasmid''' | ||
| + | #inoculate 5.0mL LB in 50mL conical tubes w/ 100ug/mL of ampilicin | ||
| + | ##the cultures (2 of them) grew for ~12hrs (8am to 8pm) | ||
| + | #centrifuge (using the 50mL conical tubes) for 5000rpm for 10min at 4C | ||
| + | #carefully discard the supernatant and resuspend the pellet with 250uL of P1 buffer (kept in the 4C fridge) | ||
| + | #transfer into a labelled 1.5mL eppendorf tube (set pipetman to 500uL just in case) | ||
| + | #add 250uL P2 Buffer--> invert 4-6times (should turn blue)--> add 350uL N3 buffer--> invert 4-6times (should become vicious or clumpy) | ||
| + | ##centrifuge for 13,000rpm for 10mins | ||
| + | #pipet out supernatant into a QIA spin column--> centrifuge for 60s--> discard flow through | ||
| + | ##did not pipet out all of the supernatant | ||
| + | #add 500uL PB buffer --> centrifuge 60s--> discard flow through | ||
| + | #add 750uL PE buffer--> centrifuge 60s--> discard--> centrifuge 60s again | ||
| + | #transfer into a labeled 1.5mL eppendorf tube--> add 50uL EB (elution buffer) | ||
| + | ##allow it to sit for 1 min (it helps to release the DNA from the column) | ||
| + | |||
| + | '''fimB PCR product Purification''' | ||
| + | #Used sample A and B of PCR product (save C and D for later) | ||
| + | ##total volume of PCR product = 81 uL (40.5uL separately) | ||
| + | #add 5 volumes of PB buffer (202.5uL) to 1 volume of PCR product (40.5uL); invert | ||
| + | #transfer into a QIAquick spin column (provided in the Qiagen kit) | ||
| + | ##set pipetman to 260uL to be sure to get all of the mixture | ||
| + | #centrifuge spin column at 13,000rpm for 60s | ||
| + | #discard the flow through (in the collection tube) and add 750uL PE buffer (to wash the DNA) | ||
| + | repeat step 4 again | ||
| + | #discard flow through and centrifuge again to get the remain buffers out | ||
| + | #place the column into a labeled 1.5mL eppendorf tube | ||
| + | #add 50uL EB buffer (to elute out the DNA) directly to the white inner circle of the column (avoid touching the pipet tip to the column) | ||
| + | ##allow the mix to sit in the column for 1 min, then centrifuge for 1 min (13,000rpm) | ||
| + | ##remember to point the cap of the eppendorf tube in the opposite direction of the centrifuge machine | ||
| + | |||
| + | '''fimB Digest''' | ||
| + | #pre-programmed PCR machine to Digest (DIG1) | ||
| + | #use 5 PCR reaction tubes for each; 5 for fimB and 5 for pBAD | ||
| + | #add the following amounts in that order (total volume of 20uL) | ||
| + | *16 uL of the DNA (fimB and pBAD to their respective tubes) | ||
| + | *2 uL of NEB 2 buffer | ||
| + | *1 uL NcoI | ||
| + | *1uL HindIII | ||
| + | #incubate for 37C overnight (12hrs) | ||
| + | ##place into 4C fridge the next day | ||
| + | |||
| + | ==8/16/2010== | ||
| + | ''Kevin, Marc, Alena'' | ||
| + | |||
| + | Alena order/picks up NEB Ligase (T4 DNA Ligase, 20,000U/mL) from MSRB II enzyme store | ||
| + | |||
| + | Attempted experiment: | ||
| + | *Added 1uL of CIP (Calf Intestine Phosphatase) to plasmid reactions only (pBAD) | ||
| + | **CIP will cut off the 3' phosphate to prevent the plasmid from folding back on itself | ||
| + | *incubate CIP-pBAD at 37C for 1 hr and then heat-shock (65C for 15 min) | ||
| + | |||
| + | '''We made a huge human error when setting up our thermal cycle program on the PCR machine. We did not realize we only set the thermal cycle for our digest on 8/14/2010>> [https://2010.igem.org/Team:Michigan/Pili_Expression#8.2F14.2F2010] for 12 minutes (12:00) rather than the intended time period of 12 hours (12:00:00). This explains the condensation present in the PCR machine (it stayed at 4C for too long). We discovered our mistake after running the incubation of CIP+pBAD for 37C for (1:00) 1 minute then heat-shock at 65C for 15 minute. It is VERY IMPORTANT to triple check one another when entering in any program.''' | ||
| + | |||
| + | Rest of the day: | ||
| + | *headed over to the ERB and made two cultures of 5mL LB-Amp (in 50mL conical tubes)--> placed in the Lin Lab 4C fridge | ||
| + | |||
| + | ==8/17/2010== | ||
| + | ''Kevin, Marc, Alena'' | ||
| + | |||
| + | Marc inoculated the two cultures made last night (8/16/2010) with pBAD and grew in 37C shaker (~9am to ~6p, 9hrs) | ||
| + | |||
| + | '''NOTE REGARDING LB CULTURE MADE ON 8/16/2010''' | ||
| + | *Marc noticed the 400mL LB media made in the ERB to be contaminated. | ||
| + | *Our two cultures could have been contaminated but pBAD grew pretty well (saturated) so we're not too worried | ||
| + | |||
| + | fimE and fimB knock out (K.O.) strains came in | ||
| + | *Marc adds 50uL of kanamycin onto each LB-agar plate and spread evenly with sterile glass beads | ||
| + | **LB-agar plate ~25mL | ||
| + | **Jeremy Minty's advice: more Kan is better than too little --> Kan 100 instead of Kan 50 | ||
| + | **allow the plate to absorb the kanamycin for 2 hours before applying the strains | ||
| + | *Alena follows Jeremy Minty's example of carefully taking out the small filter circle out of the foil with sterile techniques onto a LB-agar plate (previously labeled) | ||
| + | **add ~75uL of LB onto the filter circle | ||
| + | **streak the filter paper w/ inoculating streakers (used 3 of them) | ||
| + | *incubate plates w/ filter paper (still on the agar) upside (agar side up) at 37C overnight | ||
| + | |||
| + | Followed 8/14/2010 protocol (above) for: | ||
| + | *Miniprep for pBAD plasmid | ||
| + | **Last time we used the incorrect spin column (QIAquick spin column; purple; used for PCR purification). This time we used QIAprep spin column (blue; had no cap on the column) | ||
| + | *PCR purification for fimB | ||
| + | **used PCR products labeled C and D tubes | ||
| + | *Digest fimB and pBAD | ||
| + | **This time, made sure the time on the program was set to 12:00:00 | ||
| + | **5 rxns for each fimB and pBAD (10 total tubes, 20uL total volume in each tube) | ||
| + | |||
| + | ==8/18/2010== | ||
| + | ''Kevin, Marc, Alena'' | ||
| + | |||
| + | Met with Chris to discuss about yeast agglutination | ||
| + | *Chris sent us a possible paper with a decent protocol to look at for the assay [[Media:YeastAgglutinationPaper.pdf]] | ||
| + | |||
| + | '''NOTE ABOUT K.O. STRAINS''' | ||
| + | *fimE K.O. did not grow out (which contradicts the prediction that fimE K.O grows faster than fimB K.O.) | ||
| + | *try letting the plate grow at 37C for a longer period of time (one colony observed) | ||
| + | *remove the filter circle paper and put onto a new LB-agar plate | ||
| + | *suspect too much kanamycin on the plate | ||
| + | |||
| + | Lab work: | ||
| + | #add CIP (calf intestine phosphatase= cuts off the 3' phosphate to prevent plasmid from closing back on itself) to plasmid (pBAD) reaction tubes only--> incubate at 37C for 1 hour (using PCR machine) | ||
| + | #heat/inactivate pBAD and fimB reaction tubes for 15 min at 65C (again using PCR machine) | ||
| + | #perform DNA purification on the digests (follow 8/16/2010 procedure) | ||
| + | ##using 210uL and 200uL of PB buffer for pBAD and fimB respectively | ||
| + | #ran a gel on the digest | ||
| + | [[Image:100818_DigestGel.jpg| 500px]] | ||
| + | #ran nanodrop3.0.1 on the digest | ||
| + | ##use EB buffer as the blank | ||
| + | ##ALWAYS clean twice when done using the machine | ||
| + | |||
| + | {| style="color:#1c2bf2;background-color:#fafa19;font-size:9pt;text-align:center" cellpadding="5" cellspacing="0" border="1" bordercolor="#fff" width="33%" | ||
| + | |- | ||
| + | ! | ||
| + | ! FimB Digest | ||
| + | ! pBAD Digest | ||
| + | |- | ||
| + | | 260/280 | ||
| + | | 1.86 | ||
| + | | 1.90 | ||
| + | |- | ||
| + | | 260/230 | ||
| + | | 1.83 | ||
| + | | 2.20 | ||
| + | |- | ||
| + | | ng/uL | ||
| + | | 24.8 | ||
| + | | 59.7 | ||
| + | |} | ||
| + | |||
| + | ==8/22/2010== | ||
| + | ''Kevin, Marc'' | ||
| + | |||
| + | Ran Ligation of FimB and pBAD for 16 hrs O/N | ||
| + | |||
| + | Prepared culture of DH5a for electroporation tomorrow. | ||
| + | |||
| + | ==8/23/2010== | ||
| + | ''Kevin, Marc'' | ||
| + | |||
| + | Growing competent cells for electroporation. | ||
| + | |||
| + | Colony started growing at 2:00. | ||
| + | |||
| + | Measured ODs | ||
| + | {| style="color:#1c2bf2;background-color:#fafa19;font-size:9pt;text-align:center" cellpadding="5" cellspacing="0" border="1" bordercolor="#fff" width="33%" | ||
| + | |- | ||
| + | ! Time | ||
| + | ! OD600 | ||
| + | |- | ||
| + | | 3:30 | ||
| + | | .114 | ||
| + | |- | ||
| + | | 5:30 | ||
| + | | .206 | ||
| + | |- | ||
| + | | 6:30 | ||
| + | | .441 | ||
| + | |} | ||
| + | |||
| + | Precipitate Ligation product w/butanol | ||
| + | |||
| + | Add to eppendorf tube: | ||
| + | #50 ul ultrapure H2O | ||
| + | #Ligation product | ||
| + | #500 ul butanol | ||
| + | Spin at 4C, 13000 rpm for 20 min | ||
| + | |||
| + | Made ampicillin plates | ||
| + | *Add 25 ul Amp100 to LB plates | ||
| + | |||
| + | 6 plates: | ||
| + | *Cells only | ||
| + | *Plasmid only | ||
| + | *2x pBAD+FimB undiluted | ||
| + | *2x pBAD+FimB 1:100 | ||
| + | |||
| + | ==9/2/2010== | ||
| + | ''Kevin'' | ||
| + | |||
| + | Electroporation of pBAD+FimB into K12 and ΔFimB::kan cells. | ||
| + | |||
| + | <u>15 plates</u> <br> | ||
| + | 1x K12 control (amp) <br> | ||
| + | 1x Plasmid control (amp + kan) <br> | ||
| + | 1x ΔFimB control (amp + kan) <br> | ||
| + | 6x ΔFimB (amp + kan) <br> | ||
| + | 6x K12 (amp) <br> | ||
| + | |||
| + | {| style="color:#1c2bf2;background-color:#fafa19;font-size:9pt;text-align:center" cellpadding="5" cellspacing="0" border="1" bordercolor="#fff" width="33%" | ||
| + | |- | ||
| + | ! Cell Type | ||
| + | ! Time Constant | ||
| + | |- | ||
| + | | Plasmid Control | ||
| + | | 5.6 | ||
| + | |- | ||
| + | | ΔFimB Control | ||
| + | | 5.4 | ||
| + | |- | ||
| + | | K12 Control | ||
| + | | 5.6 | ||
| + | |- | ||
| + | | ΔFimB A | ||
| + | | 5.4 | ||
| + | |- | ||
| + | | ΔFimB B | ||
| + | | 5.4 | ||
| + | |- | ||
| + | | K12 A | ||
| + | | 5.8 | ||
| + | |- | ||
| + | | K12 B | ||
| + | | 5.6 | ||
| + | |} | ||
| + | |||
| + | ==9/7/2010== | ||
| + | ''Kevin'' | ||
| + | |||
| + | Started overnight cultures of 1:1000 dilutions of K12 and ΔFim::kan, both with the pBAD+FimB plasmid. | ||
| + | |||
| + | ==9/8/2010== | ||
| + | ''Kevin'' | ||
| + | |||
| + | Created frozen stocks of K12 and ΔFim::kan w/the plasmid from overnight cultures. Stored them in box 1 in the ERB -20C freezer. | ||
| + | |||
| + | Ran miniprep of K12 and ΔFim::kan, using 5 ml of each culture. | ||
| + | Note: Centrifuge in ERB will only go up to 5000 rpm w/50 ml tubes, therefore we transferred the culture to several eppendorf tubes. This was probably not a good idea, because we were left with a lot of leftover supernatant and that could have diluted the buffers. In the future, we should centrifuge the culture 1 ml at a time, and add 1 ml after each cycle. | ||
| + | |||
| + | Stored the product from miniprep in box 2 in the ERB -20C freezer. | ||
| + | |||
| + | ==9/9/2010== | ||
| + | ''Kevin'' | ||
| + | |||
| + | Cryopreserved frozen stocks of K12 and ΔFimB::kan w/the plasmid. | ||
| + | |||
| + | Ran digest to determine whether the plasmid and FimB were actually in the cell. Used protocol from previous digest on 8/14. | ||
| + | |||
| + | ==9/10/2010== | ||
| + | ''Kevin'' | ||
| + | |||
| + | Ran gel of previous day's digest, unfortunately the gel was inconclusive. There is an undetermined error with the gel electrophoresis machine. | ||
Revision as of 05:46, 19 October 2010
| Sunday | Monday | Tuesday | Wednesday | Thursday | Friday | Saturday | |
| Week 6 | - | - | - | - | - | - | 8/7/2010 |
| Week 7 | - | 8/9/2010 | - | 8/11/2010 | 8/12/2010 | - | 8/14/2010 |
| Week 8 | - | 8/16/2010 | 8/17/2010 | 8/18/2010 | - | - | - |
| Week 9 | 8/22/2010 | 8/23/2010 | - | - | - | - | - |
| Week 10 | - | - | - | - | 9/2/2010 | - | - |
| Week 11 | - | - | 9/7/2010 | 9/8/2010 | 9/9/2010 | 9/10/2010 | - |
| Week 12 | - | - | - | - | - | - | - |
| Week 13 | - | - | - | - | - | - | - |
8/7/2010
Kevin, Marc, Alena
PCR #1 Used a gradient from 40C to 60C for the first 3 cycles to find the optimum anneling temperature.
All of the annealing temperatures gave a good result according to the gel.
8/9/2010
Kevin, Marc, Alena
Used a 57C degree annealing temperature to get enough DNA for the digest and ligation.
4 out of the 5 PCR reactions worked well according to the gel.
The 5th well could have been a loading problem or there wasn't enough DNA.
8/11/2010
Kevin, Marc
Met with Chris, received advice for updating digest and ligation protocols.
8/12/2010
Kevin, Marc, Alena
Met and discussed protocols for digestion and ligation of FIMB into pBAD.
1. Added 5 mL of LB broth each to 2 50 mL falcon tubes in the ERB lab using sterile technique.
2. Added 5 microliters of Kanamycin to each of the 50 mL tubes in step 1.
Went to the budget committee meeting for 1 hour with the tubes.
3. Obtained the cryostock of pBAD from the Lin -80C freezer (iGEM box cell #73)
4. Stabbed cryostock using a sterile 200 microliter pipette tip and pipetted into media from step 2.
5. At 8:05PM placed the two falcon tubes from step 4 into the incubator/shaker at 30C.
8/14/2010
Kevin, Marc, Alena
Miniprep pBAD plasmid
- inoculate 5.0mL LB in 50mL conical tubes w/ 100ug/mL of ampilicin
- the cultures (2 of them) grew for ~12hrs (8am to 8pm)
- centrifuge (using the 50mL conical tubes) for 5000rpm for 10min at 4C
- carefully discard the supernatant and resuspend the pellet with 250uL of P1 buffer (kept in the 4C fridge)
- transfer into a labelled 1.5mL eppendorf tube (set pipetman to 500uL just in case)
- add 250uL P2 Buffer--> invert 4-6times (should turn blue)--> add 350uL N3 buffer--> invert 4-6times (should become vicious or clumpy)
- centrifuge for 13,000rpm for 10mins
- pipet out supernatant into a QIA spin column--> centrifuge for 60s--> discard flow through
- did not pipet out all of the supernatant
- add 500uL PB buffer --> centrifuge 60s--> discard flow through
- add 750uL PE buffer--> centrifuge 60s--> discard--> centrifuge 60s again
- transfer into a labeled 1.5mL eppendorf tube--> add 50uL EB (elution buffer)
- allow it to sit for 1 min (it helps to release the DNA from the column)
fimB PCR product Purification
- Used sample A and B of PCR product (save C and D for later)
- total volume of PCR product = 81 uL (40.5uL separately)
- add 5 volumes of PB buffer (202.5uL) to 1 volume of PCR product (40.5uL); invert
- transfer into a QIAquick spin column (provided in the Qiagen kit)
- set pipetman to 260uL to be sure to get all of the mixture
- centrifuge spin column at 13,000rpm for 60s
- discard the flow through (in the collection tube) and add 750uL PE buffer (to wash the DNA)
repeat step 4 again
- discard flow through and centrifuge again to get the remain buffers out
- place the column into a labeled 1.5mL eppendorf tube
- add 50uL EB buffer (to elute out the DNA) directly to the white inner circle of the column (avoid touching the pipet tip to the column)
- allow the mix to sit in the column for 1 min, then centrifuge for 1 min (13,000rpm)
- remember to point the cap of the eppendorf tube in the opposite direction of the centrifuge machine
fimB Digest
- pre-programmed PCR machine to Digest (DIG1)
- use 5 PCR reaction tubes for each; 5 for fimB and 5 for pBAD
- add the following amounts in that order (total volume of 20uL)
- 16 uL of the DNA (fimB and pBAD to their respective tubes)
- 2 uL of NEB 2 buffer
- 1 uL NcoI
- 1uL HindIII
- incubate for 37C overnight (12hrs)
- place into 4C fridge the next day
8/16/2010
Kevin, Marc, Alena
Alena order/picks up NEB Ligase (T4 DNA Ligase, 20,000U/mL) from MSRB II enzyme store
Attempted experiment:
- Added 1uL of CIP (Calf Intestine Phosphatase) to plasmid reactions only (pBAD)
- CIP will cut off the 3' phosphate to prevent the plasmid from folding back on itself
- incubate CIP-pBAD at 37C for 1 hr and then heat-shock (65C for 15 min)
We made a huge human error when setting up our thermal cycle program on the PCR machine. We did not realize we only set the thermal cycle for our digest on 8/14/2010>> [1] for 12 minutes (12:00) rather than the intended time period of 12 hours (12:00:00). This explains the condensation present in the PCR machine (it stayed at 4C for too long). We discovered our mistake after running the incubation of CIP+pBAD for 37C for (1:00) 1 minute then heat-shock at 65C for 15 minute. It is VERY IMPORTANT to triple check one another when entering in any program.
Rest of the day:
- headed over to the ERB and made two cultures of 5mL LB-Amp (in 50mL conical tubes)--> placed in the Lin Lab 4C fridge
8/17/2010
Kevin, Marc, Alena
Marc inoculated the two cultures made last night (8/16/2010) with pBAD and grew in 37C shaker (~9am to ~6p, 9hrs)
NOTE REGARDING LB CULTURE MADE ON 8/16/2010
- Marc noticed the 400mL LB media made in the ERB to be contaminated.
- Our two cultures could have been contaminated but pBAD grew pretty well (saturated) so we're not too worried
fimE and fimB knock out (K.O.) strains came in
- Marc adds 50uL of kanamycin onto each LB-agar plate and spread evenly with sterile glass beads
- LB-agar plate ~25mL
- Jeremy Minty's advice: more Kan is better than too little --> Kan 100 instead of Kan 50
- allow the plate to absorb the kanamycin for 2 hours before applying the strains
- Alena follows Jeremy Minty's example of carefully taking out the small filter circle out of the foil with sterile techniques onto a LB-agar plate (previously labeled)
- add ~75uL of LB onto the filter circle
- streak the filter paper w/ inoculating streakers (used 3 of them)
- incubate plates w/ filter paper (still on the agar) upside (agar side up) at 37C overnight
Followed 8/14/2010 protocol (above) for:
- Miniprep for pBAD plasmid
- Last time we used the incorrect spin column (QIAquick spin column; purple; used for PCR purification). This time we used QIAprep spin column (blue; had no cap on the column)
- PCR purification for fimB
- used PCR products labeled C and D tubes
- Digest fimB and pBAD
- This time, made sure the time on the program was set to 12:00:00
- 5 rxns for each fimB and pBAD (10 total tubes, 20uL total volume in each tube)
8/18/2010
Kevin, Marc, Alena
Met with Chris to discuss about yeast agglutination
- Chris sent us a possible paper with a decent protocol to look at for the assay Media:YeastAgglutinationPaper.pdf
NOTE ABOUT K.O. STRAINS
- fimE K.O. did not grow out (which contradicts the prediction that fimE K.O grows faster than fimB K.O.)
- try letting the plate grow at 37C for a longer period of time (one colony observed)
- remove the filter circle paper and put onto a new LB-agar plate
- suspect too much kanamycin on the plate
Lab work:
- add CIP (calf intestine phosphatase= cuts off the 3' phosphate to prevent plasmid from closing back on itself) to plasmid (pBAD) reaction tubes only--> incubate at 37C for 1 hour (using PCR machine)
- heat/inactivate pBAD and fimB reaction tubes for 15 min at 65C (again using PCR machine)
- perform DNA purification on the digests (follow 8/16/2010 procedure)
- using 210uL and 200uL of PB buffer for pBAD and fimB respectively
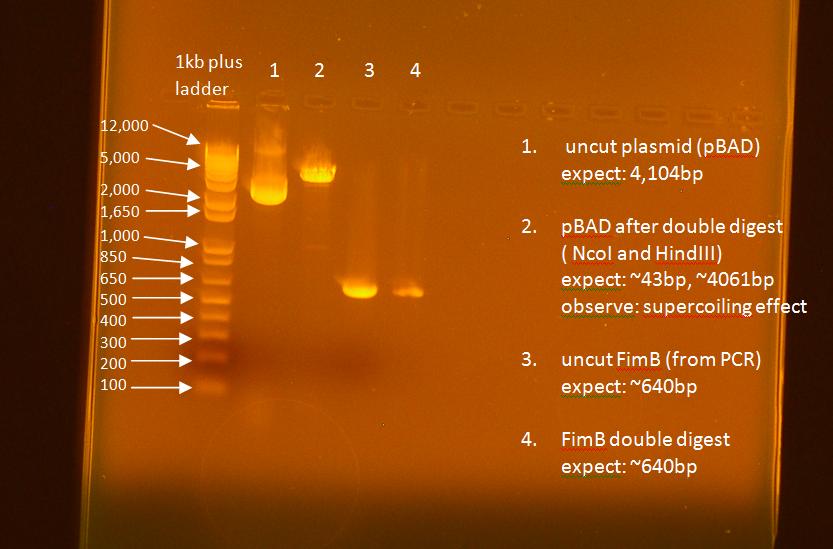
- ran a gel on the digest
- ran nanodrop3.0.1 on the digest
- use EB buffer as the blank
- ALWAYS clean twice when done using the machine
| FimB Digest | pBAD Digest | |
|---|---|---|
| 260/280 | 1.86 | 1.90 |
| 260/230 | 1.83 | 2.20 |
| ng/uL | 24.8 | 59.7 |
8/22/2010
Kevin, Marc
Ran Ligation of FimB and pBAD for 16 hrs O/N
Prepared culture of DH5a for electroporation tomorrow.
8/23/2010
Kevin, Marc
Growing competent cells for electroporation.
Colony started growing at 2:00.
Measured ODs
| Time | OD600 |
|---|---|
| 3:30 | .114 |
| 5:30 | .206 |
| 6:30 | .441 |
Precipitate Ligation product w/butanol
Add to eppendorf tube:
- 50 ul ultrapure H2O
- Ligation product
- 500 ul butanol
Spin at 4C, 13000 rpm for 20 min
Made ampicillin plates
- Add 25 ul Amp100 to LB plates
6 plates:
- Cells only
- Plasmid only
- 2x pBAD+FimB undiluted
- 2x pBAD+FimB 1:100
9/2/2010
Kevin
Electroporation of pBAD+FimB into K12 and ΔFimB::kan cells.
15 plates
1x K12 control (amp)
1x Plasmid control (amp + kan)
1x ΔFimB control (amp + kan)
6x ΔFimB (amp + kan)
6x K12 (amp)
| Cell Type | Time Constant |
|---|---|
| Plasmid Control | 5.6 |
| ΔFimB Control | 5.4 |
| K12 Control | 5.6 |
| ΔFimB A | 5.4 |
| ΔFimB B | 5.4 |
| K12 A | 5.8 |
| K12 B | 5.6 |
9/7/2010
Kevin
Started overnight cultures of 1:1000 dilutions of K12 and ΔFim::kan, both with the pBAD+FimB plasmid.
9/8/2010
Kevin
Created frozen stocks of K12 and ΔFim::kan w/the plasmid from overnight cultures. Stored them in box 1 in the ERB -20C freezer.
Ran miniprep of K12 and ΔFim::kan, using 5 ml of each culture. Note: Centrifuge in ERB will only go up to 5000 rpm w/50 ml tubes, therefore we transferred the culture to several eppendorf tubes. This was probably not a good idea, because we were left with a lot of leftover supernatant and that could have diluted the buffers. In the future, we should centrifuge the culture 1 ml at a time, and add 1 ml after each cycle.
Stored the product from miniprep in box 2 in the ERB -20C freezer.
9/9/2010
Kevin
Cryopreserved frozen stocks of K12 and ΔFimB::kan w/the plasmid.
Ran digest to determine whether the plasmid and FimB were actually in the cell. Used protocol from previous digest on 8/14.
9/10/2010
Kevin
Ran gel of previous day's digest, unfortunately the gel was inconclusive. There is an undetermined error with the gel electrophoresis machine.
 "
"