Team:Michigan/Modeling
From 2010.igem.org
Mathematical models are very important in the design of an iGEM project. The Michigan modeling team is composed of Josh, Kevin, and Jennifer. We are currently researching different models that have been previously done that can help us in our own modeling endeavors. To construct our models, we have taken advantage of MATLAB's Simbiology toolset.
Pili
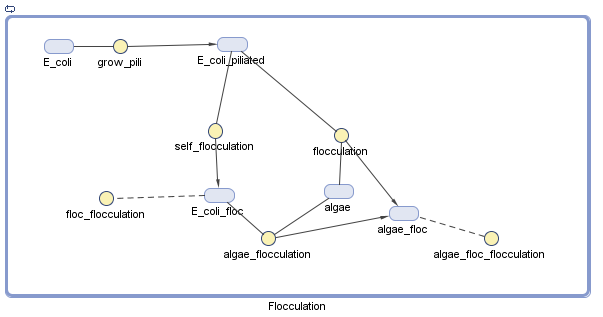
We have decided to model the growth of a biofilm using pili. By modeling the flocculation of the algae, we hope to be able to predict how our reactions will proceed and determine quantities such as the optimal initial concentration of E. coli.
The pili neural network has been characterized in several papers [1][2]. Essentially, the two recombinases FimB and FimE control an invertible DNA element that acts as a switch, known as FimS. When FimS is in the "on" position, the cell becomes fimbriated. It has been previously determined that the level of piliation depends on the ratio [FimE]/[FimB]. The goal of the modeling team is to determine the optimal ratio to promote flocculation.
Quorum Sensing
Quorum sensing has been modeled by other iGEM teams for previous competitions, including: [http://parts.mit.edu/igem07/index.php/Bangalore Bangalore] and Singapore
Surface Display
References
1. Kuwahara, H., Myers, C., Samoilov, M., Abstracted Stochastic Analysis of Type 1 Pili Expression in E. Coli.
2. Wolf, D., and Arkin, A., Fifteen Minutes of fim: Control of Type 1 Pili Expression in E. Coli. OMICS 6 2002
 "
"