Team:ETHZ Basel/Modeling
From 2010.igem.org
(→Support for wet laboratory) |
(→Support for wet laboratory) |
||
| Line 25: | Line 25: | ||
== Support for wet laboratory == | == Support for wet laboratory == | ||
To create the biological implementation of E. lemming, the parts of the core components had to be chosen in an order to improve chances to result in a functioning ensemble. By using the combined molecular models for ''in silico'' evaluation of the best possible parts, it was possible to reduce the amount of different combinations to be tested. | To create the biological implementation of E. lemming, the parts of the core components had to be chosen in an order to improve chances to result in a functioning ensemble. By using the combined molecular models for ''in silico'' evaluation of the best possible parts, it was possible to reduce the amount of different combinations to be tested. | ||
| + | |||
| + | [[Team:ETHZ_Basel/Modeling/Evaluation '''Evaluation results''']] have showed, that molecular modeling and experimental biology can collaborate to gain new insight for both perceptions of the problem. | ||
== Test bench for information processing == | == Test bench for information processing == | ||
Revision as of 13:06, 22 September 2010
Molecular Modeling Overview
In order to support wet laboratory experiments and to create a test bench for the information processing part, a molecular model of E. lemming was created. This goal was achieved by implementing and combining deterministic molecular models of the individual parts.
Implementation of molecular models
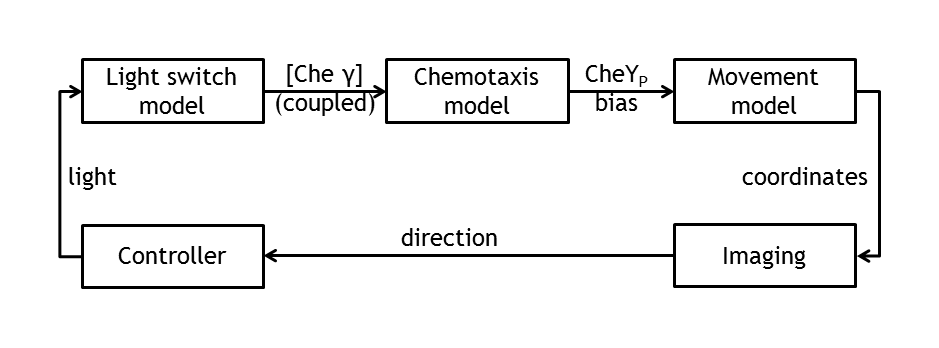
The core component of E. lemming is the fusion of one light-sensitive protein (LSP?) to a protein of the chemotaxis pathway (Che?). Upon change of wavelength of light pulses, this component will dimerize with the corresponding light-sensitive protein (LSP?'), which is linked to an anchor protein, bound to an anchor (plasmid). The result is a change of the spatial localization of Che? and perturbation of the chemotaxis pathway, which ultimately leads to a different tumbling/directed flagellar movement state ratio.
Individual molecular models
In a first step, we implemented individual deterministic molecular models of subdevices.
- Light switch: based upon the light-sensitive dimerizing Arabidopsis proteins PhyB and PIF3.
- Chemotaxis: two similar models of the chemotaxis receptor pathway.
- Movement: a statistical model of E. coli movement, determined by distribution of input bias.
Combined molecular models
The next step, combination of the individual molecular models to a comprehensive model of E. lemming was achieved in two substeps:
- Light switch - Chemotaxis: used to provide support for wet laboratory.
- Light switch - Chemotaxis - Movement: complete molecular model.
Support for wet laboratory
To create the biological implementation of E. lemming, the parts of the core components had to be chosen in an order to improve chances to result in a functioning ensemble. By using the combined molecular models for in silico evaluation of the best possible parts, it was possible to reduce the amount of different combinations to be tested.
[[Team:ETHZ_Basel/Modeling/Evaluation Evaluation results]] have showed, that molecular modeling and experimental biology can collaborate to gain new insight for both perceptions of the problem.
 "
"