Team:ETHZ Basel/Modeling
From 2010.igem.org
(→Molecular Modeling Overview) |
(→Implementation of molecular models) |
||
| Line 6: | Line 6: | ||
== Implementation of molecular models == | == Implementation of molecular models == | ||
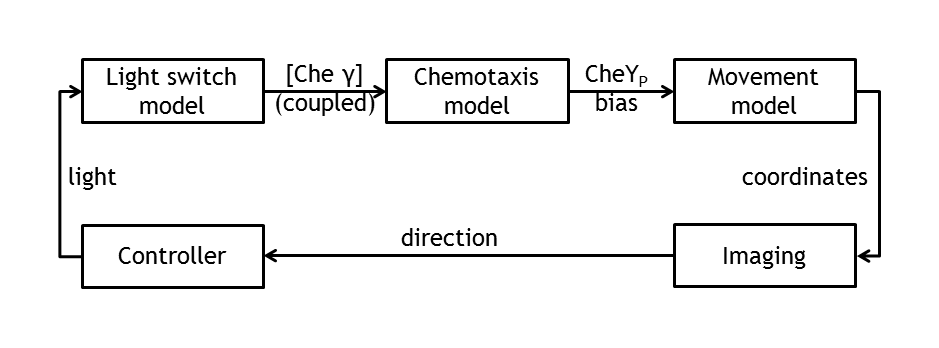
| - | [[Image:Modeling_overview.png|thumb|400px|'''Combined models''' Coupled individual models for the simulation of the whole process and their interfaces.]] | + | [[Image:Modeling_overview.png|thumb|400px|'''Combined models.''' Coupled individual models for the simulation of the whole process and their interfaces.]] |
E. lemming, aims to modify the chemotaxis property of E.coli such that, instead of response to a chemical attractant or repellent, the bacterium responds to a light stimulus. Furthermore, this light sensitivity is used to control E.coli’s movement by deciding, at any given time, which type of motion the bacterium will adopt (tumbling or straight run). This leads to a controllable E.coli, which can move to a defined direction, as a result of the combination of tumbling and straight run. The bacteria in the experiment are imaged and, by image processing, the position of a single tracked cell is inferred. By activating a light switch, the user decides whether the bacterium should continue running or should change direction. | E. lemming, aims to modify the chemotaxis property of E.coli such that, instead of response to a chemical attractant or repellent, the bacterium responds to a light stimulus. Furthermore, this light sensitivity is used to control E.coli’s movement by deciding, at any given time, which type of motion the bacterium will adopt (tumbling or straight run). This leads to a controllable E.coli, which can move to a defined direction, as a result of the combination of tumbling and straight run. The bacteria in the experiment are imaged and, by image processing, the position of a single tracked cell is inferred. By activating a light switch, the user decides whether the bacterium should continue running or should change direction. | ||
Revision as of 11:27, 22 September 2010
Molecular Modeling Overview
In order to support wet laboratory experiments and to create a test bench for the information processing part, a molecular model of E. lemming was created. This goal was achieved by implementing and combining deterministic molecular models of the individual parts.
Implementation of molecular models
E. lemming, aims to modify the chemotaxis property of E.coli such that, instead of response to a chemical attractant or repellent, the bacterium responds to a light stimulus. Furthermore, this light sensitivity is used to control E.coli’s movement by deciding, at any given time, which type of motion the bacterium will adopt (tumbling or straight run). This leads to a controllable E.coli, which can move to a defined direction, as a result of the combination of tumbling and straight run. The bacteria in the experiment are imaged and, by image processing, the position of a single tracked cell is inferred. By activating a light switch, the user decides whether the bacterium should continue running or should change direction.
In the theory world, the steps we are following in mindlessly driving E.coli to our pre - defined target are the following:
• deterministic (ODE) & stochastic models of the chemotaxis pathway (documented from the literature)
• model of the movement of E.coli (built on the information of the pathway derived from the molecular models)
• control algorithms ( built on the user’s desire to play around with E.coli)
• image tracking & image processing algorithms
• java applications/movies of the E.Lemming (the fun part)
 "
"