Team:SDU-Denmark/labnotes8
From 2010.igem.org
Lab notes (8/30 - 9/5)
Contents |
Flagella
Since the previous FlhDCmut was wrongly mutated due to incorrect mutation primers, we are now back to square one with new correct primers. The next weeks the flagella-group are working according to the following plan:
1) Miniprep of plasmids with "wrong" FlhDCmut
2) Two-step PCR to get mutatet FlhDC
3) Cut and Ligate into pSB1C3 and pSB1AK3 and transform into TOP10 cells
4) Send to sequencing
5) Characterize biobrick
Miniprep of "wrong" FlhDCmut
Done by: Louise
Date: September 3rd
Protocol: [MP1.2]
Notes:
No changes of protocol were made.
Results:
Nanodrop after sample was dried down:
Sample 1: Concentration: 192ng/ul Purity: 1.96/1.90
Sample 2: Concentration: 200ng/ul Purity: 1.84/1.88
The samples were run on a 1.5% gel with a 10kb ladder. The bands are positioned between 1500bp and 1200bp. FlhDCmut in pSB1C3 is 1248bp.
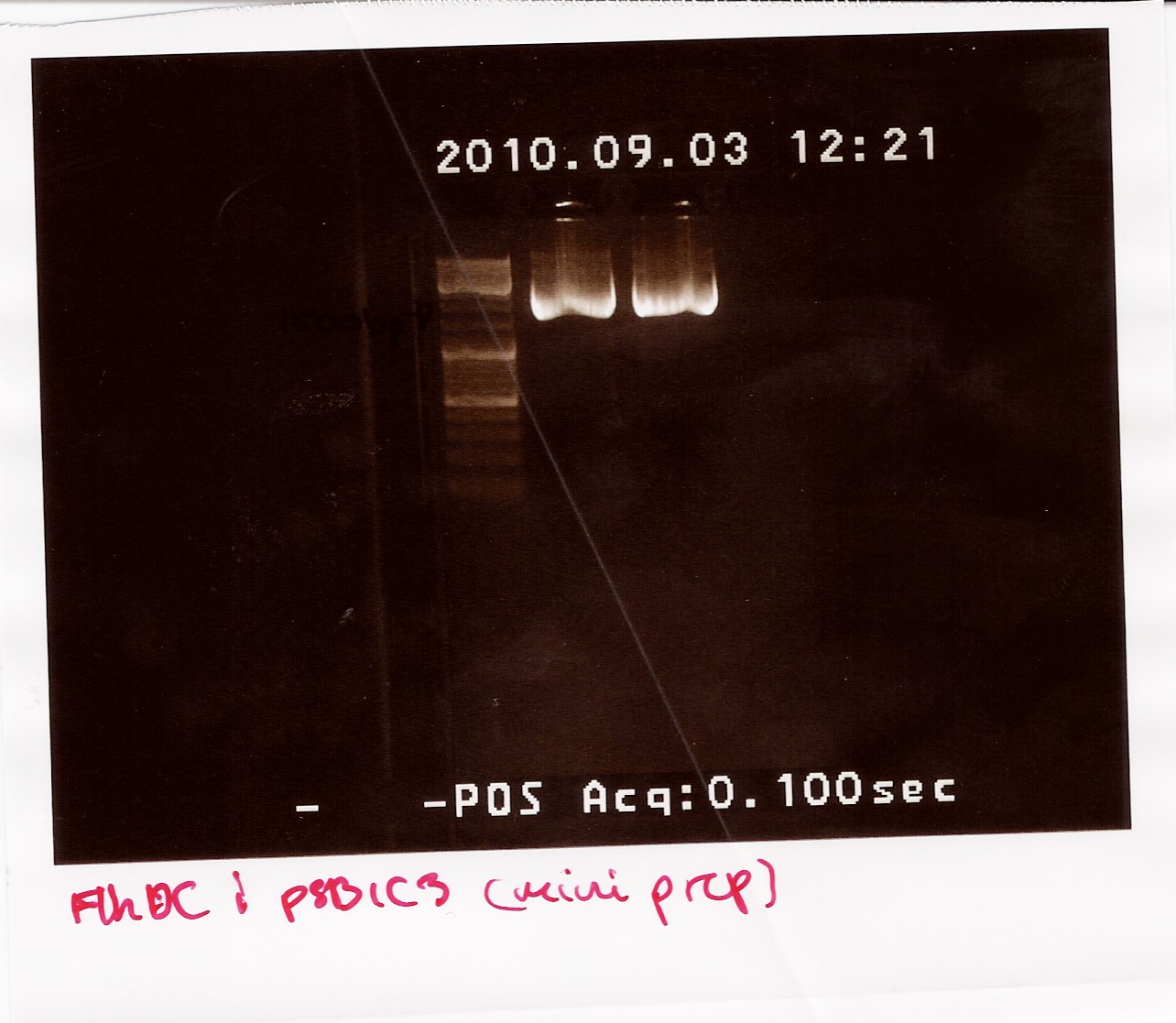
Photosensor
PCR on pKJ606 with PSfw and VR primers
Date: 31/8
https://2010.igem.org/wiki/index.php?title=Team:SDU-Denmark/labnotes8&action=edit§ion=3
Done by: LC
Methods: PCR
Protocols: CP1.3[1]
Notes:
Premix:
7,5 µl 10xTAQ Buffer
3 µl MgCl2
3 µl VF2
3 µl VR
1,5 µl dNTP
55,5 µl H2O
3 µl template (PS1 miniprep)
3/8 µl TAQ Polymerase
PCR Program:
| Start |
94 C |
2 min |
| Denaturing |
94 C |
1 min |
| Annealing |
55 C |
1 min |
| Elongation |
72 C |
3 min |
| Goto2 |
rep |
29x |
| End |
72 C |
3 min |
| Hold |
4 C |
Results: The bands that showed up were around 2500 BP as expected from the sequencing results. This means that the designed primers have a high possibility of working, so that they will be ordered.
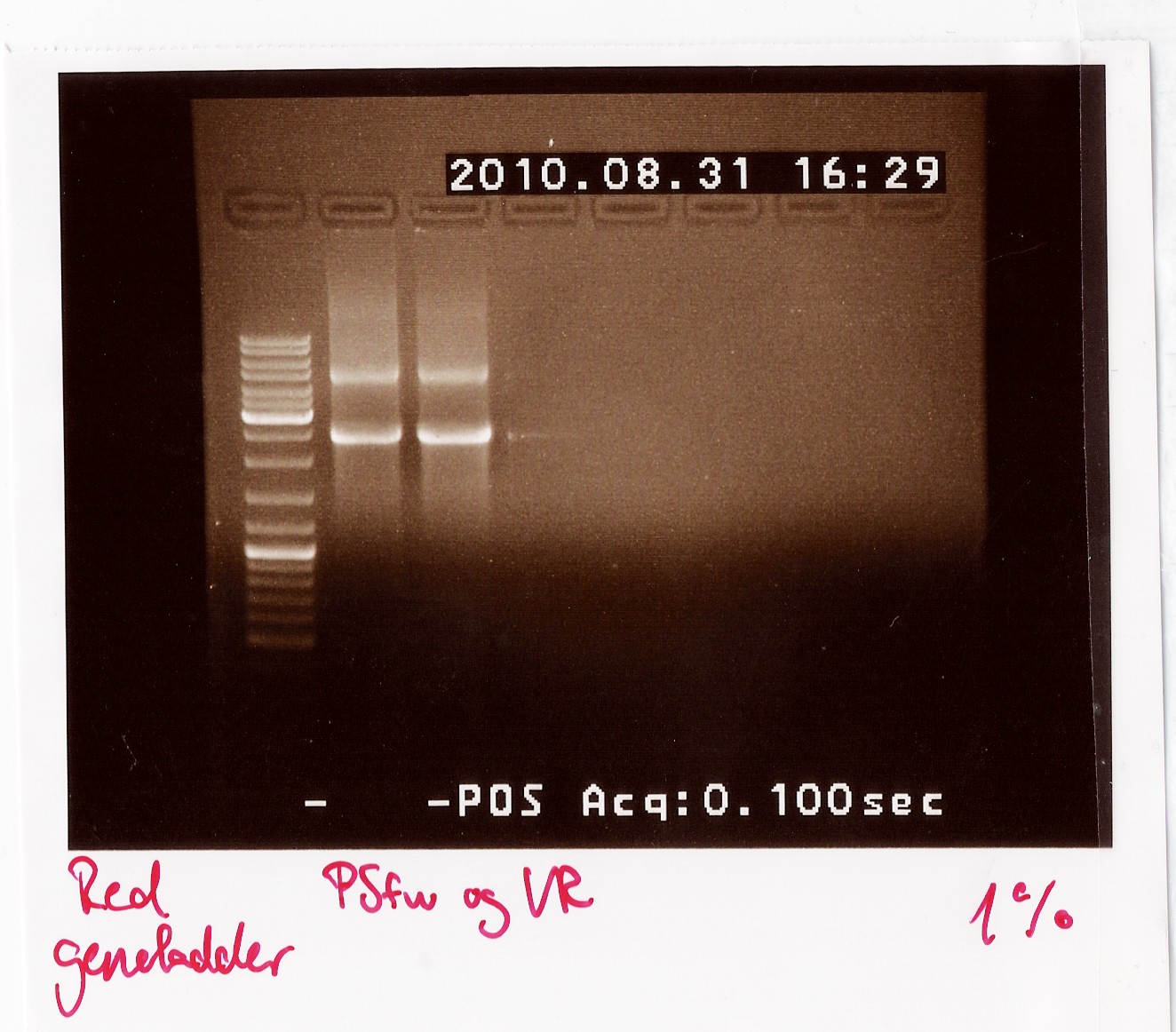
PCR on pKJ606 with PSfw and PSrv primers
Date: 02/09
Done by: LC
Methods: PCR
Protocols: CP1.3[2]
Notes:
Premix:
7,5 µl 10xTAQ Buffer
3 µl MgCl2
3 µl VF2
3 µl VR
1,5 µl dNTP
55,5 µl H2O
3 µl template (PS1 miniprep)
3/8 µl TAQ Polymerase
PCR Program:
| Start |
94 C |
2 min |
| Denaturing |
94 C |
1 min |
| Annealing |
55 C |
1 min |
| Elongation |
72 C |
2 min |
| Goto2 |
rep |
29x |
| End |
72 C |
3 min |
| Hold |
4 C |
Results: The bands that showed up were around 1750 BP, further confirming sequencing results. New primers for taking the whole gene out have been ordered.
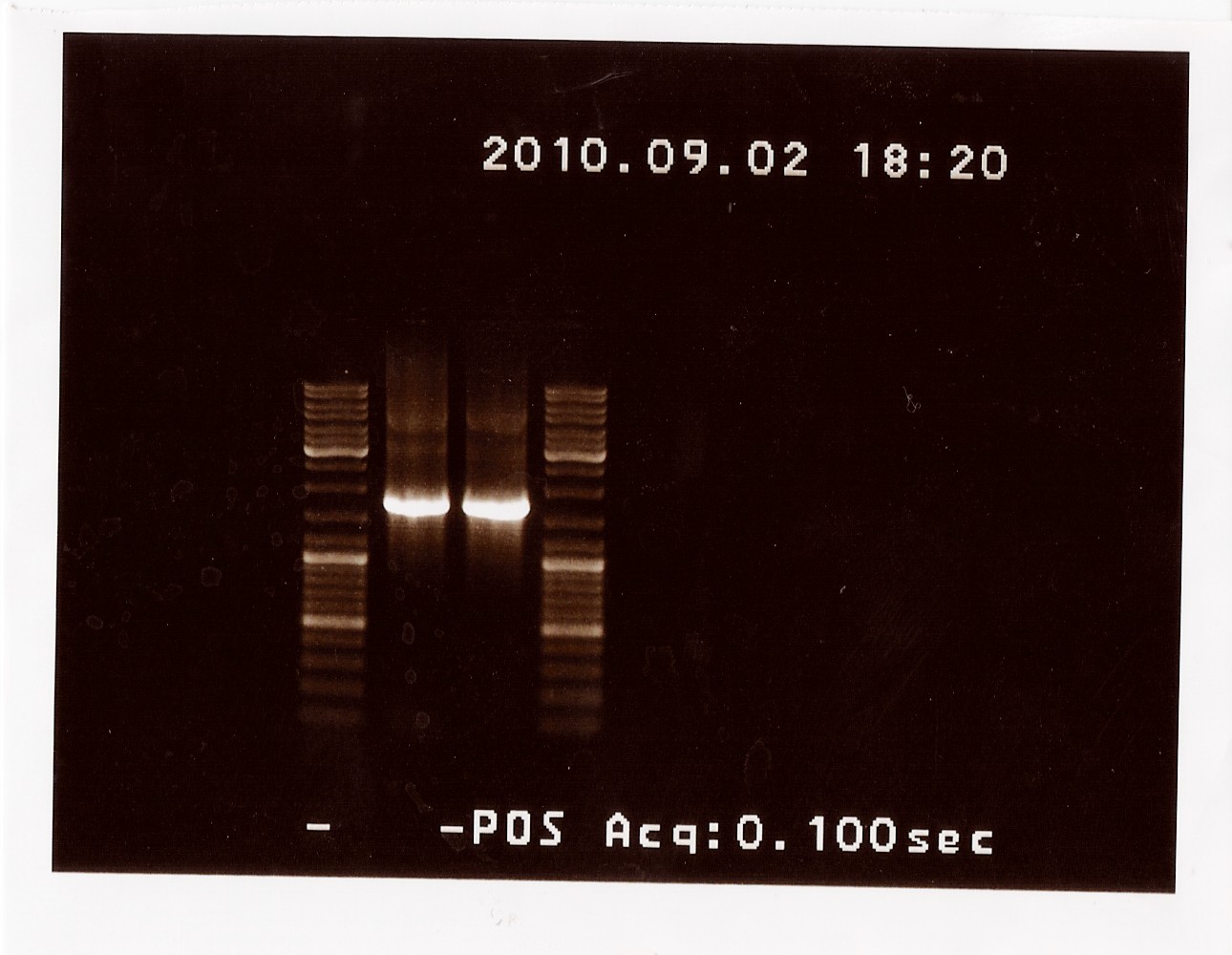
PCR on pKJ606 with fwPS2 and rvPS2 primers
Date: 04/09
Done by: LC
Methods: PCR
Protocols: CP1.3[3]
Notes:
Premix:
12,5 µl 10xTAQ Buffer
5 µl MgCl2
5 µl VF2
5 µl VR
2,5 µl dNTP
59,5 µl H2O
4 µl template (Consisting of 2 µl H2O and 2 µl of pKJ606 miniprep)
1/2 µl TAQ Polymerase
PCR Program:
| Start |
94 C |
2 min |
| Denaturing |
94 C |
1 min |
| Annealing |
55 C |
1 min |
| Elongation |
72 C |
2:30 min |
| Goto2 |
rep |
29x |
| End |
72 C |
3 min |
| Hold |
4 C |
Results: Primers were confirmed working, the next step will be PFU pcr.
Date: 30/08
Done by: Tommy & Marie
Methods: PCR
Protocols: CP1.3[4]
Notes:
To finde the optimal anneling temperatur a gradient PCR from 60 to 75 degrees
PCR Program:
| Start |
94 C |
2 min |
| Denaturing |
94 C |
1 min |
| Annealing |
Grad. C |
1 min |
| Elongation |
72 C |
4 min |
| Goto2 |
rep |
29x |
| End |
72 C |
5 min |
| Hold |
4 C |
Colonies 34, 37, 61 and 63 was selected for minipreps and freezing cultures.
 "
"