Team:Stockholm/11 August 2010
From 2010.igem.org
Contents |
Hassan
- Started making animation for what's happening in our project.
- TODO for tomorrow: complete the interaction map, and starting summarizing texts for the map, hope to finish it by sunday...
Nina
Digestion of Tyrosinase
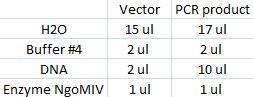
I cut the miniprepped site directed mutagenesis and PCR product tyrosinase with the restriction enzyme NgoMIV.
Digestion:
Incubated in 37 °C for 2 hours.
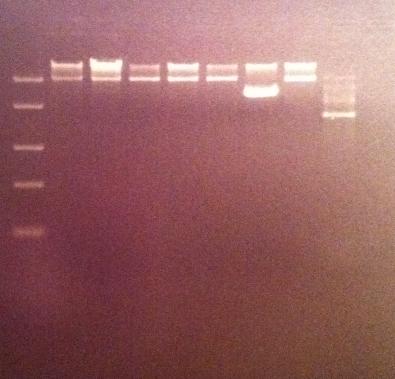
I ran the digested products in an agarose 1 % gel 80 V.
DNA Ladder: FastRuler™ Middle Range, ready-to-use, 100-5000 bp Fermentas
Arrangement on gel:
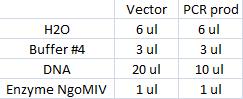
I ran a new digestion since the first one did not indicate wether or not the digest had worked or not. The second digest I altered the amounts in the mixture into those values that I have used previously when digesting.
Second digest:
Incubated in 37 °C for 1 hour (PCR prod) and 3 hours (vector).
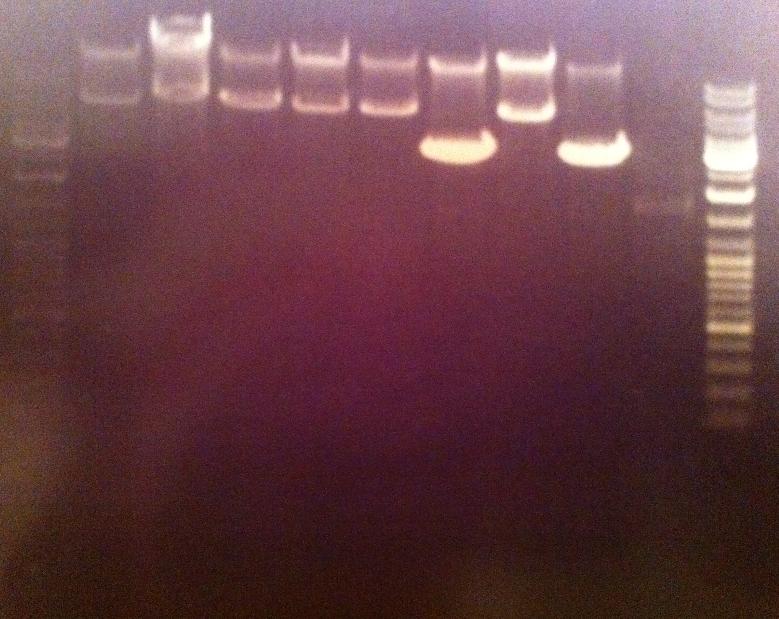
I ran the digested products in an agarose 1 % gel 80 V.
Ladder: GeneRuler™ DNA Ladder Mix, ready-to-use, 100-10,000 bp Fermentas
Arrangement on gel:
I still don't get reliable results that the enzyme has cut, therefore I will run a gel tomorrow with an uncut vector to compare with the cut vector sample, this should show if the enzyme can cut and from there I know that the site directed mutagenesis tyrosinase samples have been succesful in altering the restiction site for NgoMIV since they won't be cut.
Andreas
Cloning of IgG protease
Continued from 10/8
Colony PCR
Four clones picked from each of the two plates (Q for quick transformation; S for standard transformation).
- QA, QB, QC, QD
- SA, SB, SC, SD
- Pos. control: pSB1A3.BBa_J04450 (plasmid)
Procedures
As described in colony PCR protocol. Elongation time: 1:40.
Gel verification
1 %, 80 V, 45 min
Expected: 1262 bp (BBa_J04450: 1386 bp)
Results
Very strangely sized bands again! None of the bands corresponding to the 1262 bp expected. Control corresponding well to expected 1386 bp.
Digested vector and insert will be tested on gel tomorrow.
 "
"