Team:Stanford/Research
From 2010.igem.org

| Home | Project | Applications | Modeling | Parts | Team | Notebook |
Contents |
Why Ratios?
Ratios rule the biological world, and controlling them will unlock a vast range of applications for synthetic biology.
In electrical and mechanical engineering, computers can be used to calculate a ratio from precise measurements, but biological engineering doesn't have that luxury. We need a hardwired device to compute and act on ratios independent of a controlling computer or human researcher.
The Stanford team's ratio sensors use biological machinery to eliminate the human component from ratio computation. Our output system is more efficient than two individual sensors and, most importantly, can pass its results on to other parts and devices. A ratio sensor performs a calculation that a user doesn't need to/won't be able to. A ratio sensor frees up a wavelength channel on the measurement device, and, more importantly for the future, allows the computed ratio to be passed on to another part. A useful ratio sensor that could be utilized in a broad regime of biological applications needs to be operable and accurate in micro-environments; the Stanford team's ratio sensor is engineered to function within E. Coli, which is fully operable over a range of environments.
Our Project: Two Designs for Ratio Detection
Our team decided to pursue two different systems for detecting ratios. After considering the scenarios in which biological engineers would want to use ratio sensors, having two specialized sensors seemed to allow greater flexibility of usage than a single design.
While both systems receive two input signals, the output they give is different, allowing them be applied in different situations. Here's a brief rundown of the two designs (you can see more by clicking through to the individual project pages).
The First Design:
Method: Small RNA Interference
Output: One of two possible proteins, depending on whether the ratio of input chemicals lies above or below a predetermined threshold
Useful: For reporting on a situation with a boolean output: disease detection, preterm labor warning, cancer metastasis warning
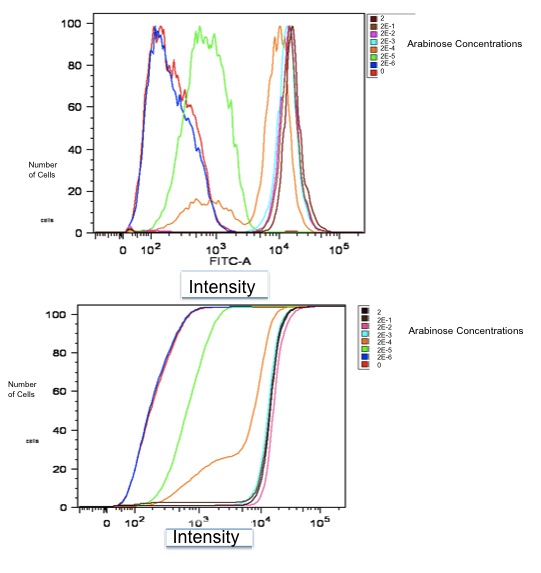
Data: Here is initial data gathered using FACS on part K353002:
Additionally, we fit a Hill Curve to this data and derived the following:
- Half-max induction: 143 +- 36 uM
- Hill coefficient: 1.4 +- .6
- Fold Induction: 860
The Second Design:
Method: Kinase/Phosphatase Regulation of a Transcription Factor
Output: One output protein whose concentration is linearly dependent on the ratio of the input chemicals
Useful: For acting in a situation requiring a graded response: drug delivery, metabolic flux control, detailed ratiometric reporting
Data: Please refer to the following pages for data:
- http://partsregistry.org/Part:BBa_K353005
- http://partsregistry.org/Part:BBa_K353006
- http://partsregistry.org/Part:BBa_K353007
- http://partsregistry.org/Part:BBa_K353008
Both sensors are:
- Modular: input and output molecules can be changed without affecting the interior mechanism of the device
- Orthogonal: device mechanisms are not found in E. coli, avoid crosstalk with host cell
Biosafety
1. Would any of your project ideas raise safety issues in terms of researcher safety, public safety, or environmental safety?
Answer: There are no major safety risks regarding our general system design. In its current form, our device is only meant for ratiometric analysis of compounds within E. coli. As with any other biological hazard, typical laboratory precaution should be taken when handling cells resistant against antibiotics.
2. Is there a local biosafety group, committee, or review board at your institution?
Answer: There is a local board at Stanford University called Health and Safety at the Stanford University School of Medicine.
3. What does your local biosafety group think about your project?
Answer: We have not communicated with our local biosafety group regarding the design or safety of our project.
4. Do any of the new BioBrick parts that you made this year raise any safety issues?
Answer: Although the kinases and phosphatases chosen for our project come from Mycobacterium tuberculosis and Streptomyces coelicolor A3(2), our team advisors and graduate mentors believe that these proteins do not pose any significant safety risks.
Research
Medal Requirements
Bronze Medal
- Register the team, have a great summer, and have fun attending the Jamboree.
- Done!
- Successfully complete and submit a Project Summary form.
- Done!
- Create and share a Description of the team's project via the iGEM wiki
- Present a Poster and Talk at the iGEM Jamboree
- Booked our plane tickets!
- Enter information detailing at least one new standard BioBrick Part or Device in the Registry of Parts
- [http://partsregistry.org/wiki/index.php?title=Part:BBa_K353002 Done!]
- Entered information for each new part or device should at least include primary nucleic acid sequence, description of function, authorship, any relevant safety notes, and an acknowledgement of sources and references.
- [http://partsregistry.org/wiki/index.php?title=Part:BBa_K353002 Done!]
- Submit DNA for at least one new BioBrick Part or Device to the Registry of Parts.
- Done!
Silver Medal
- Demonstrate that at least one new BioBrick Part or Device of your own design and construction works as expected.
- Characterize the operation of at least one new BioBrick Part or Device and enter this information on the Parts or Device page via the Registry of Parts
Gold Medal
- Characterize or improve an existing BioBrick Part or Device and enter this information back on the Registry.
- When sequencing one of our ligations, we noticed an unexpected sequence in our results. After more investigation, we determined that that sequence came from the Distribution part [http://partsregistry.org/wiki/index.php?title=Part:BBa_E1010 BBa_E1010], and that the part sequence listed on the Parts Registry was incomplete. We have notified the Registry and made a note of this discovery on the E1010 parts experience page.
- Help another iGEM team by, for example, characterizing a part, debugging a construct, or modeling or simulating their system.
- With our Twitter project, we hope to help all iGEM teams by facilitating collaboration between teams. Read more about it here!
 "
"