Team:UT-Tokyo/Sudoku construct
From 2010.igem.org


Sudoku
Introduction System Lab note Result Reference
Construct
Now we'll explain how to solve Sudoku with E.coli.
To explain this, we divide several steps:
- 1. #4C3 leak switch
- 2. #Signal virus
- * transcription of MS2 phage & location sequence
- * infection of signal virus
- * selective translational suppression
- 3. #Visualization of results
In order to solve Sudoku by E. coli, each microbe should possess grid and number information, with the ability to transmit the information mutually. At the same time, it is also necessary to control which E. coli to receive the information, according to the grid information (e.g. the information of grid 1-1 can be received by E. coli in grid 1-2, but cannot be received by E. coli in 4-4 grid). We propose a new system for the information transformation by RNA virus; “signal virus”. In this system, we propose using four types of RNA phage each with number information from 1 to 4(namely, four types of messenger RNA of site-specific recombination enzyme (SSRE)).
The point is that the grid information is not spatial one. Instead, information transmission is replaced by mutual interaction between genomes. The practical solvation is not done in the plate with 4x4 cells. It is actually solved by E. coli in each tube correspond to each grid, and the answer is induced by genome interactions in the tube.
E. coli whose number is already determined (let's call them "uni-output state") transmit their number information using "signal virus" to those whose number is still not determined ("multi-output state"). Once those E. coli in “multi-output state” received number information from appropriate grids, they have to recognize and manage the information, and finally, output their apt number. For example, if E. coli in "multi-output state" received number information in the order like 1->2->3, then, it'll become "uni-output state" and output 4. If E. coli in "multi-output state" received number information in the order like 3->2->4, then, it'll output 1 and transmit the information to grids around it. Here, you have to be aware that in this system, we need to admit randomness in the order of number information E. coli receive, and detect the variation, not the frequency of the number information. Which means that, E. coli in "multi-output state" have to output 4 whether it received number information in the order 1->2->3 or 3->1->2, and have to remain in "multi-output state" whether it recieved information 2, or 1->2, or 1->2->2->2->1 in series. That is to say, E. coli should turn into "uni-output state" only when it recieved 3 different types of the number in any order, and output the left number information. This is what our "4C3 leak switch" realizes.
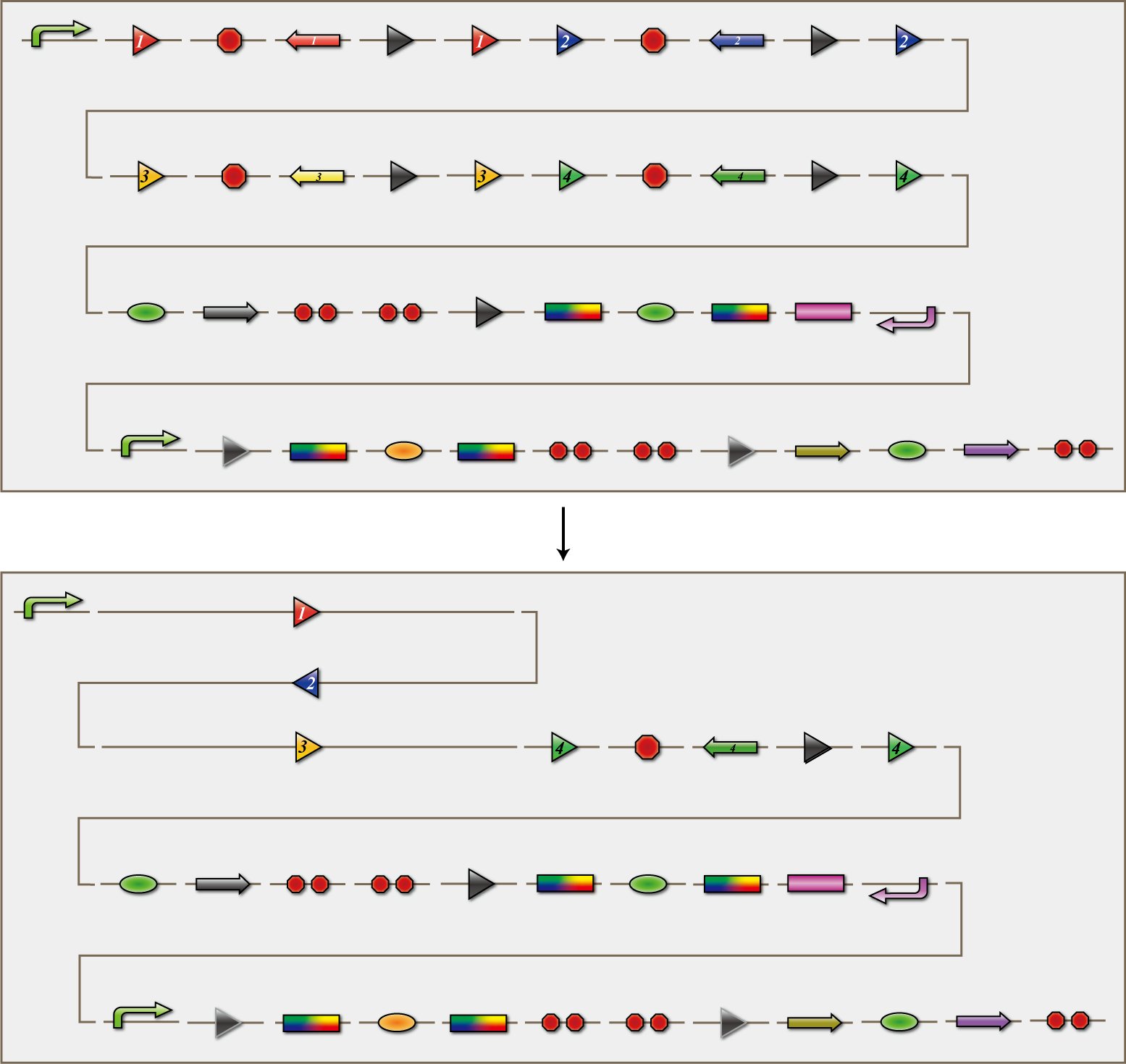
4C3 leak switch
The 4C3 leak-switch is a switch which receives three number signals and outputs number signal that it did not receive, regardless of the order of reception. In fact, number signals are mRNA encoding homologous recombinases. When three types of mRNA encoidng the recombinases enter E.coli, the switch turns on and transcription of the recombinase that it did not receive begins. 4C3 leak-switch is composed of a promoter, the cre protein coding region and four terminators in between. Each terminator is flanked on both sides by the recognition sequence of a homologous recombinase, and the terminator is excised following expression of the recombinase. Therefore when the E.coli do not receive a number signal, they have four terminators, when they receive one number signal, they have three terminators, and so on. Importantly, bacteria are able to differentiate only after they receive three number signals. 4C3 leak-switch, as the name implies, is a switch that makes use of transcriptional leakage. We assembled the sequence so that when there is more than one terminator, cre protein is not expressed but it is when there is only one. In addition, a recombinase coding sequence flanking the terminator is excised at the same time. Therefore, when the recombinase A coding sequence is excised, terminator A is also excised. As a result, after a cell receives the number signals 1, 2 and 3, only recombinase 4 coding sequence remains. Each cre protein expressed in this way irreversibility excises the sequence between the lox sites independently of the other species of cre proteins. In the example above, mRNA coding recombinase 4 corresponding to the answer of this particular puzzle, is transcribed. In conclusion, The 4C3 leak-switch begins transcription of the recombinase that it did not receive begins after receiving three signals.
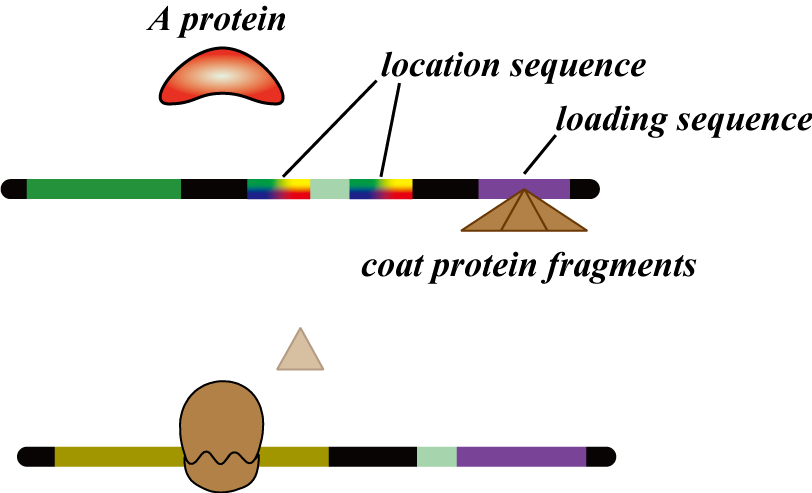
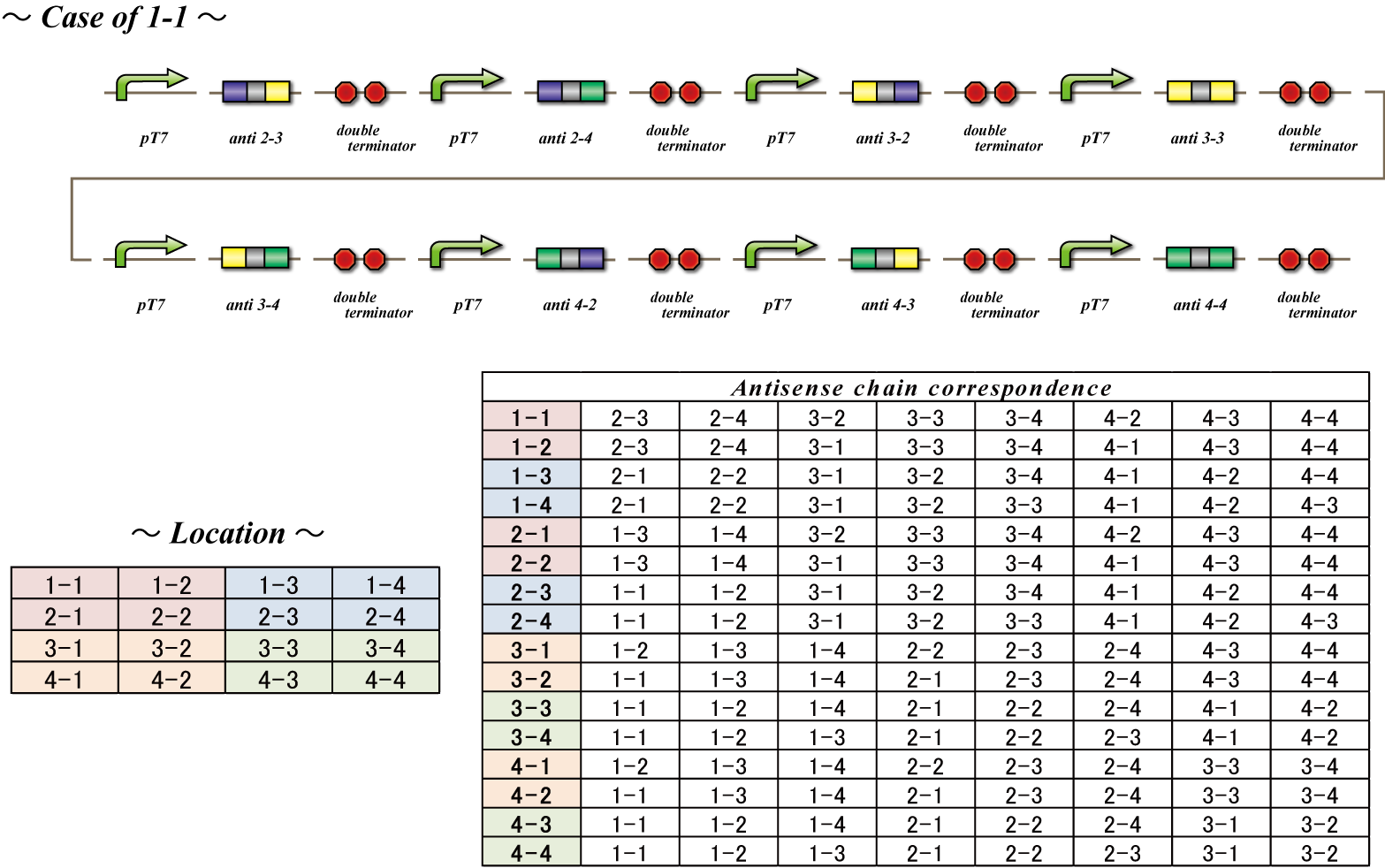
Genotype Specific Signal Transmitting Virus
The mRNA coding the homologous recombinases has two additional sequences. First, they contain a Loading sequence to the MS2 phage which enables the mRNA to be attached to the coat protein of MS2 phage. This MS2 phage is the carrier that transfers the number signal to other bacteria. These phage are only synthesized after recombination takes place, therefore preventing undifferentiated bacteria from emitting a number signal precociously. Secondly, they contain a location sequence which assigns a cell number to the bacterium. We will explain this in detail later. In conclusion, 4C3 leak-switch turns on → transcription of mRNA coding homologous recombinase → production of MS2 phage protein → phage with the number signal and location sequence attached is produced. In this way, differentiated E.coli produces phage with a number signal and location sequence and conveys this information to other bacteria.
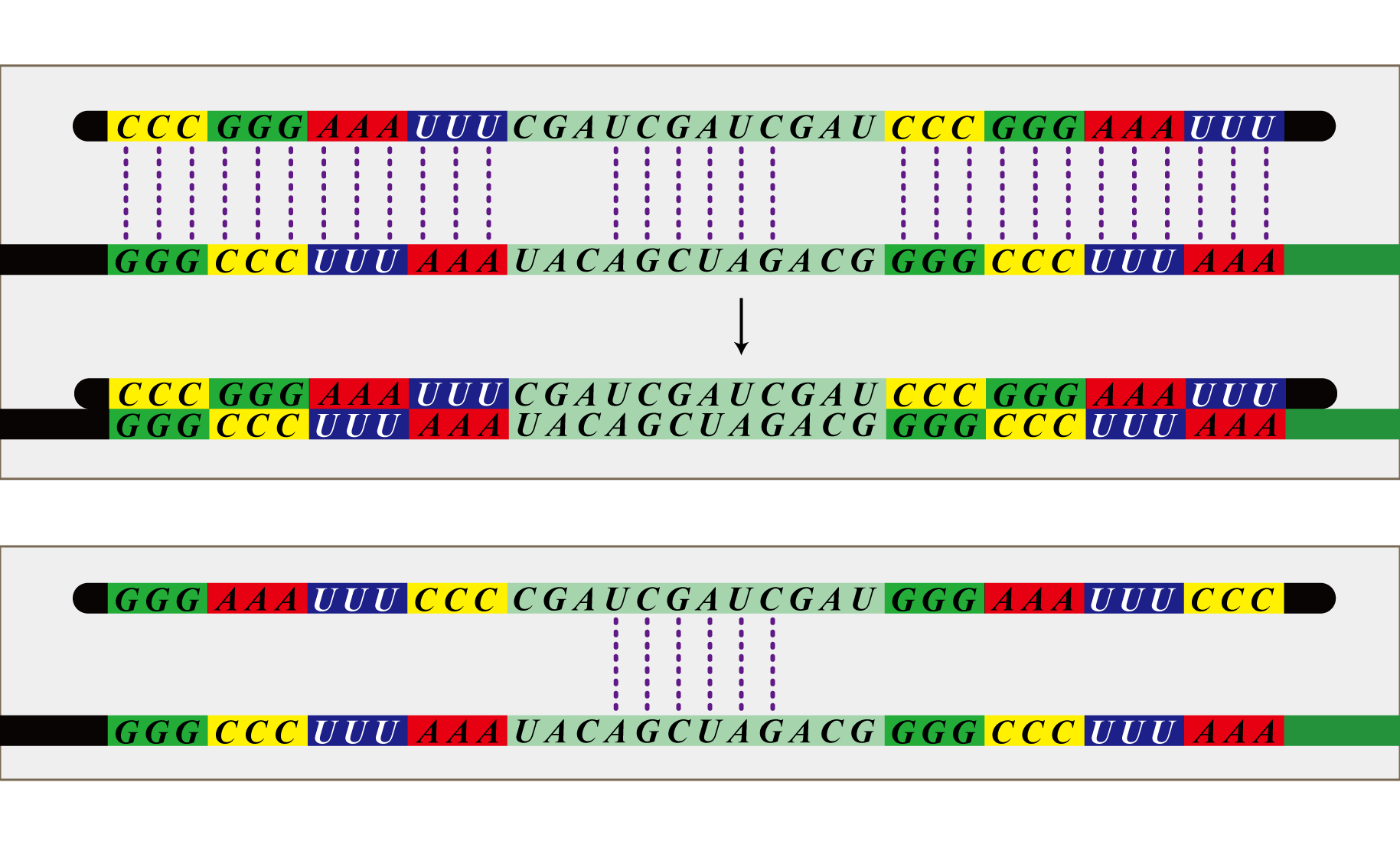
Antisense RNA
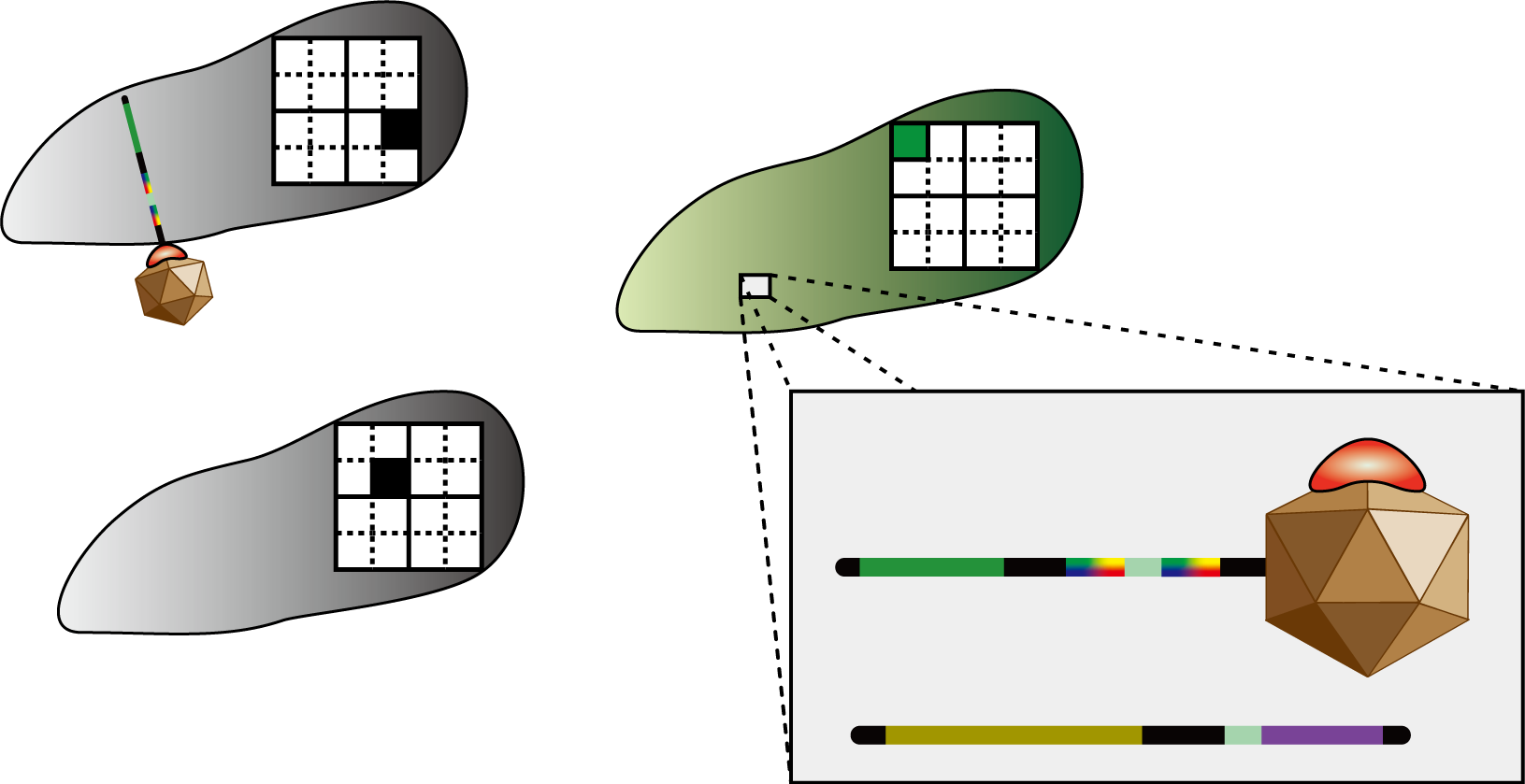
Infection of signal virus
Infection of signal virus.
Infection of signal virus.
Infection of signal virus.
Infection of signal virus.
Infection of signal virus.
Infection of signal virus.
Infection of signal virus.
Infection of signal virus.
Infection of signal virus.
Infection of signal virus.
Selective translational suppression
Selective translational suppression.
Selective translational suppression.
Selective translational suppression.
Selective translational suppression.
Selective translational suppression.
Selective translational suppression.
Selective translational suppression.
Selective translational suppression.
Selective translational suppression.
Selective translational suppression.
Visualization of results
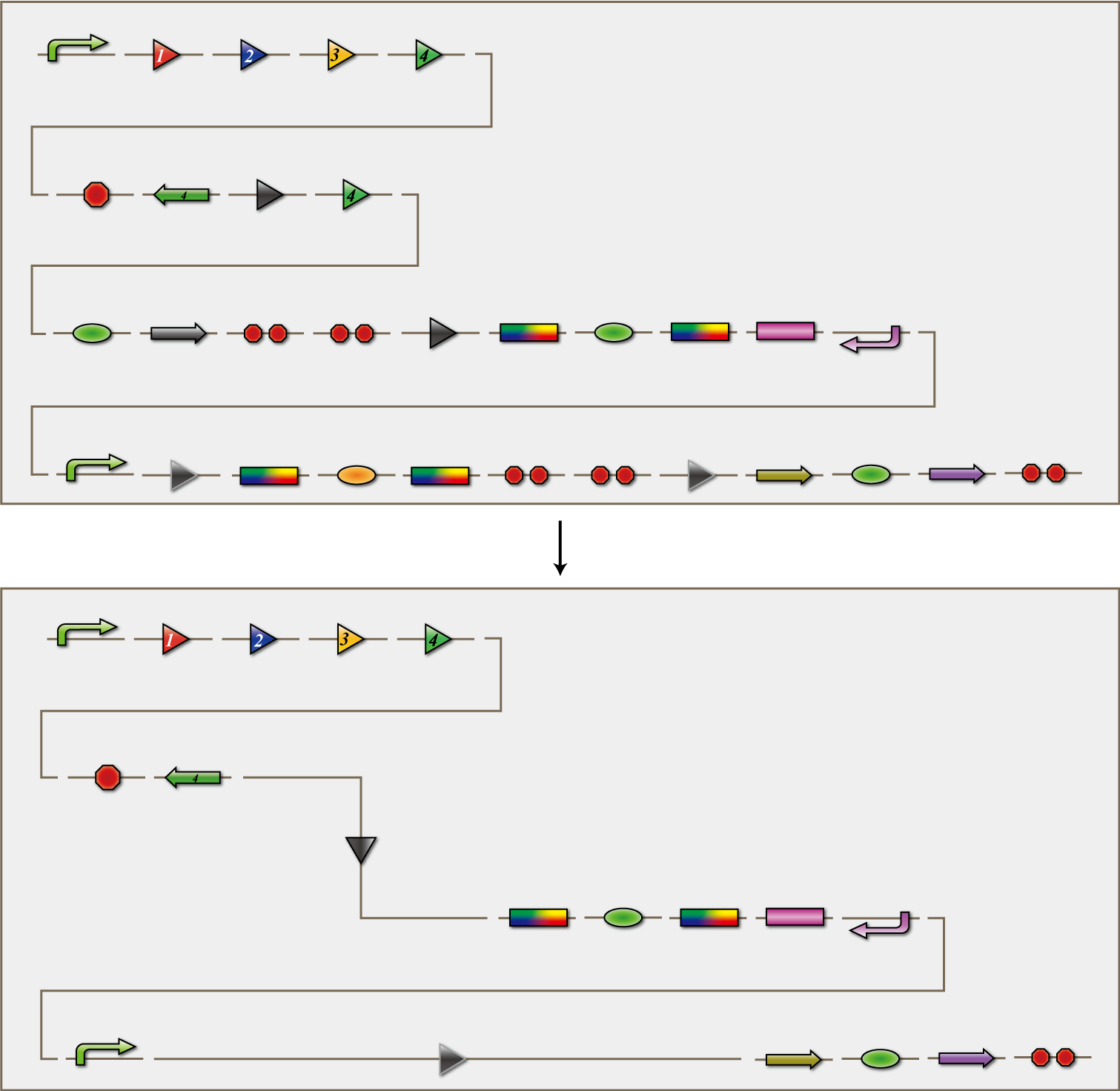
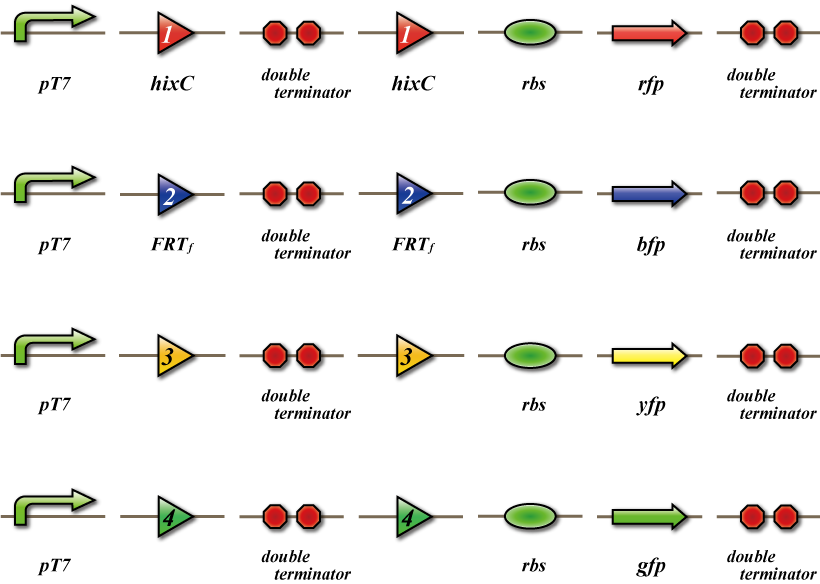
In order to be able to "see" that SUDOKU has been solved, we will prepare cluster of E. coli for detection. This detection E. coli will shut out all information from RNA virus except the virus produced by itself. By using antisense RNA, they will express SSRE which corresponds to the answer E. coli introduced.
In this detection E. coli, SSRE will cut off double terminator introduced ahead of fluorescent protein coding region. Thus, parallel fluorescent protein will be expresses according to the number information. For example, if the answer of grid 1-1 is 1, SSRE1 will be expressed inside the detection E. coli of the grid, and gfp, which is parallel fluorescent protein for 1, will be expressed.
Those detection E. coli will be placed into the detection plate, an imitating plate of the actual 4x4 plate for SUDOKU. The final step is to pour the medium in which E. coli finished solving SUDOKU to the plate. Each detection E. coli in the grid will begin to express fluorescent protein corresponding to the number and we can "see" the final answer!
 "
"