Team:Harvard/allergy/notebook
From 2010.igem.org

notebook calendar
06-14-2010 [ top ]
- Task #1
- Task #2
- Task #3
Image of gel, photo of your work, diagram, etc.
Results/ Conclusions
- We successfully cloned x, y, z
- Next step is to ligate 1, 2, 3
06-15-2010 [ top ]
06-16-2010 [ top ]
06-17-2010 [ top ]
06-18-2010 [ top ]
06-21-2010 [ top ]
06-22-2010 [ top ]
06-23-2010 [ top ]
06-24-2010 [ top ]
06-25-2010 [ top ]
06-28-2010 [ top ]
06-29-2010 [ top ]
06-30-2010 [ top ]
07-01-2010 [ top ]
07-02-2010 [ top ]
07-05-2010 [ top ]
07-06-2010 [ top ]
07-07-2010 [ top ]
07-08-2010 [ top ]
07-09-2010 [ top ]
07-12-2010 [ top ]
07-13-2010 [ top ]
07-14-2010 [ top ]
07-15-2010 [ top ]
07-16-2010 [ top ]
07-19-2010 [ top ]
07-20-2010 [ top ]
07-21-2010 [ top ]
07-22-2010 [ top ]
07-23-2010 [ top ]
07-26-2010 [ top ]
07-27-2010 [ top ]
07-28-2010 [ top ]
07-29-2010 [ top ]
07-30-2010 [ top ]
08-02-2010 [ top ]
08-03-2010 [ top ]
- Grew up cultures of completed ihpRNA constructs (Bet, LTP, Ger) in pORE expression vector
- amiRNA PCR
- Will look at results of PCR tommorrow
08-04-2010 [ top ]
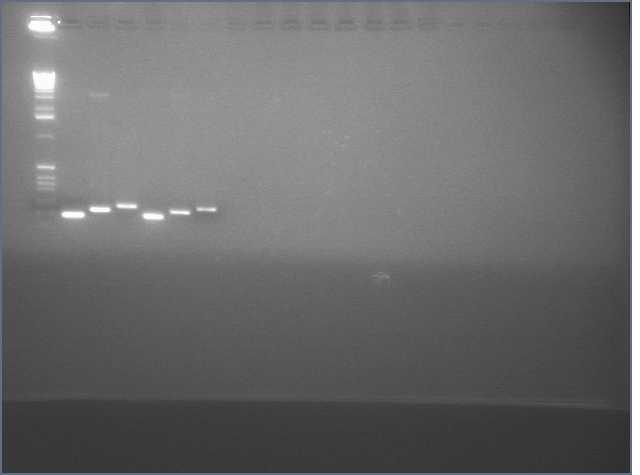
- amiRNA PCR appears to have worked at every Tm we tried:
- Digested V9/V10 to insert our constructs into
- Realized that we hadn't gel purified our ihpRNA inserts
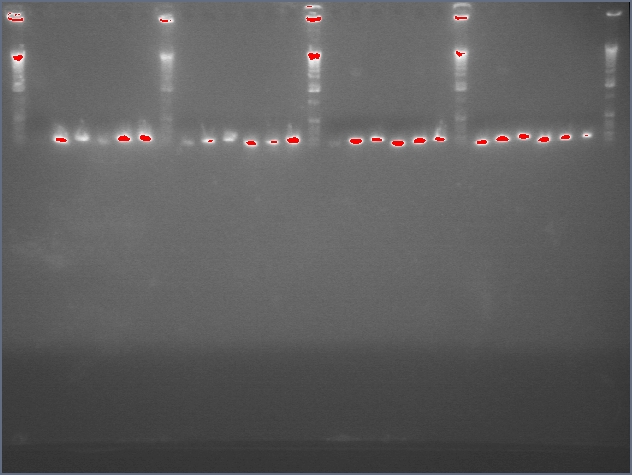
- Gel purification of inserts (entire ihpRNA parts)
Ladder, 9, 11c1, 11c2, ladder, 25c1, 28c1, 28c2, 36c1, 36c2, ladder
- Successfully gel purified our inserts and digested backbones that we will ligate into
08-05-2010 [ top ]
08-06-2010 [ top ]
08-09-2010 [ top ]
08-10-2010 [ top ]
08-11-2010 [ top ]
08-12-2010 [ top ]
08-13-2010 [ top ]
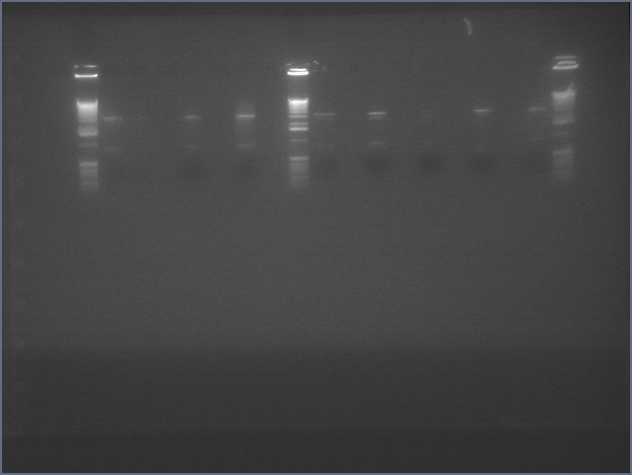
Results
If our Phusion Polymerase had worked, then we would have continued to extract the correctly formed DNA segments, performed a restriction digest and cut the PCR products into a B0120 biobrick with Xba1 and Pst1, ligate, and transform into E. coli. However, since we do not have the correct DNA, we cannot move forward.
Errors
There are several reasons for why our gel did not work. The most likely reasons is that the RNA extraction did not work properly. We used strawberry fruit as our sample for RNA extraction. Because RNA is constantly being degraded by RNAases, the integrity of our RNA may have been compromised before the addition of the denaturant.We will re-extract DNA from our plant samples, now including arabidopsis, probably tomorrow.
Another potential error could have occurred during the Phusion Polymerase. We set our annealing temperature at 72°C, which could have been too high for three of our four primers. We'll rerun our PCR with a gradient PCR of annealing temperatures to see if we can get a better result. The amount of DNA added by template was also much higher than the optimal 50~250 nanograms per 50 micrograms of PCR reactants. We will dilute the DNA templates to 100 micromolar and use one microliter per reaction so that there are approximately 100 nanograms of DNA per reaction.
 "
"