Team:Freiburg Bioware/NoteBook/Labjournal/October
From 2010.igem.org
(→Sequencing analysis of HSPG-ko in several RepCap constructs) |
(→Sequencing analysis of HSPG-ko in several RepCap constructs) |
||
| Line 246: | Line 246: | ||
| - | <p style="color:#66bbff;"><i>Comment</i>: Sequencing results show that all plasmids contain the HSGP-ko. Working will be continued using P644, P646 and P687./p><br /> | + | <p style="color:#66bbff;"><i>Comment</i>: Sequencing results show that all plasmids contain the HSGP-ko. Working will be continued using P644, P646 and P687.</p><br /> |
<html> | <html> | ||
</div> | </div> | ||
</html> | </html> | ||
Revision as of 09:31, 2 October 2010
136. labday 01.10.2010
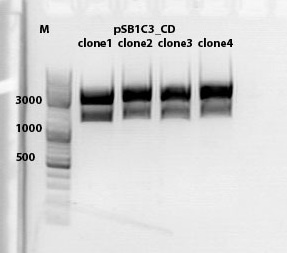
mini preps of CD clones
Investigator: Kira
c(p689)= 115 ng/ul
c(690)= 151, 20 ng/ul
c(691)= 122,27 ng/ul
c(p692) = 126,42 ng/ul
test digestion of CD clones
Investigator: Kira
| Components | sample Volume/µL |
| DNA | 4,0 µl |
| BSA (10x) | 2 µl |
| Buffer no. 4 | 2,0 µl |
| Enzyme 1 XbaI | 1,0 µl |
| Enzyme 2 AgeI | 1,5 µl |
| H2O | 9,5 µl |
| Total volume | 20 |
incubation @ 37 C for approx. 2 h
1% agarose gel
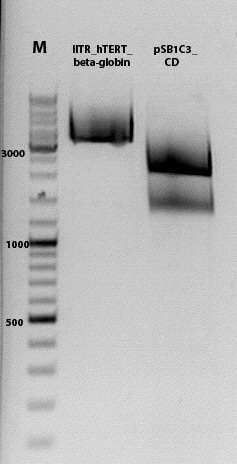
Biobrick assembly pSB1C3_lITR_hTERT_beta-globin_CD
Investigator: Kira
c(pSB1C3_lITR_hTERT_beta-globin)= 333 ng/ul
c(pSB1C3_CD)= 151 ng/ul
| Components | vector Volume/µL | insert Volume/µL |
| DNA | 4,5 µl | 6 |
| BSA (10x) | 3 µl | 3 |
| Buffer no. 4 | 3,0 µl | 3 |
| Enzyme 1 XbaI | 0 µl | 1,5 |
| Enzyme 2 SpeI | 1,5 µl | 0 |
| Enzyme 3 PstI-HF | 1,0 | 1 |
| H2O | 17 | 15,5 |
| Total volume | 25 |
incubation @ 37 C for approx. 2 h
1% agarose gel
Ligation
DNA-mix: 8 ul (vector 4,6ul)+(insert 3,4 ul)
T4 ligase: 1 ul
T4 buffer: 1 ul
Incubation @ RT for 30 min
Transformation was performed according to the standard protocol w BL21 cells.
Sequencing results of pCerulean_Zegfr:1907_MiddleLinker_VP2/3_HSPG-KO
Investigator: Hanna
Comment: All N-terminal fusion approaches with VP2/3_HSPG-KO revealed positive results except of pCerulean_Zegfr:1907_MiddleLinker_VP2/3_HSPG-KO. Another clone was picked, preped, test digested and sent for sequencing.
Conclusion: Sequencing results revealed positive results.
Picking clones of pGA14_MiddleLinker_VP2/3_insCap and pGA14_MiddleLinker_VP2/3_HSPG-KO
Investigator: Hanna
Unfortunately there grew a bacteria lawn over night - it was hardly not possible to pick clones. Nevertheless I tried and picked 2 clones of pGA14_MiddleLinker_VP2/3_insCap and pGA14_MiddleLinker_VP2/3_HSPG-KO.
To do: Mini-Prep and test digestion.
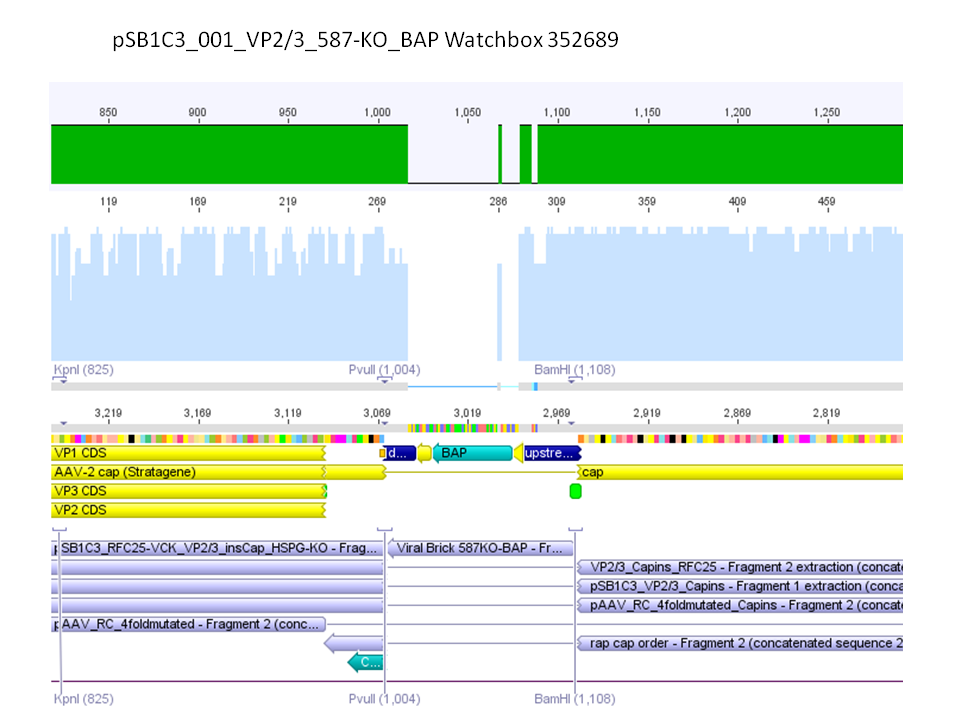
Sequencing results of pSB1C3_001_VP2/3_587-KO_BAP and pSB1C3_001_VP2/3_587-KO_6xHis
Investigator: Hanna
Comment: For the creation of our super constructs, the His-Tag and the BAP motif need to be cloned into VP2/3 for N-terminal fusion to VP2. Sequencing results showed that the 6xHis and BAP motif was not cloned into VP2/3_insCap.
To do: Clone 587-KO_6xHis and 587-KO_BAP into pSB1C3_001_VP2/3_insCap 1. via digestion of inserts and 2. via hybridization of referring oligos - digestion of vector with 800-900 ng.
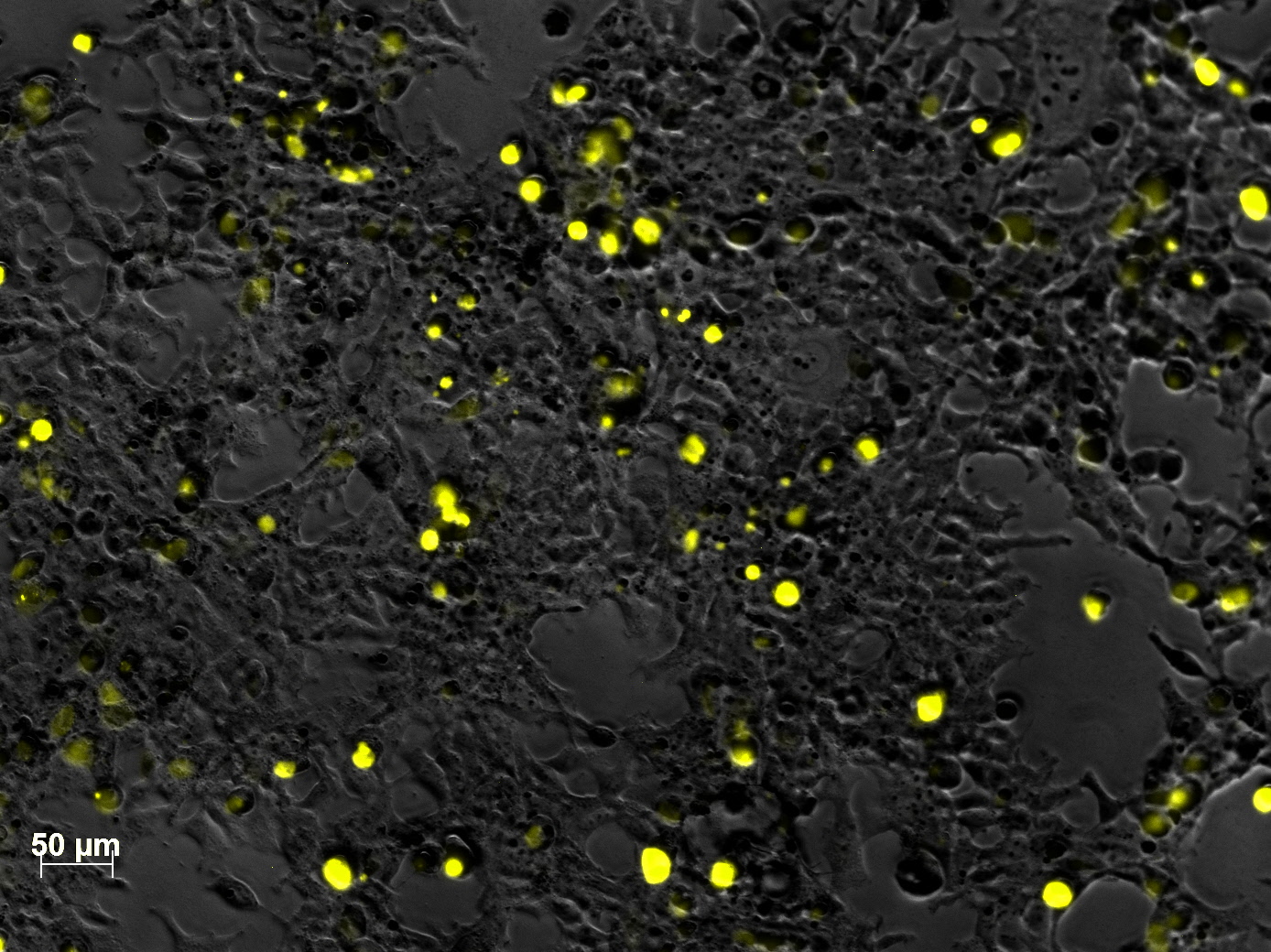
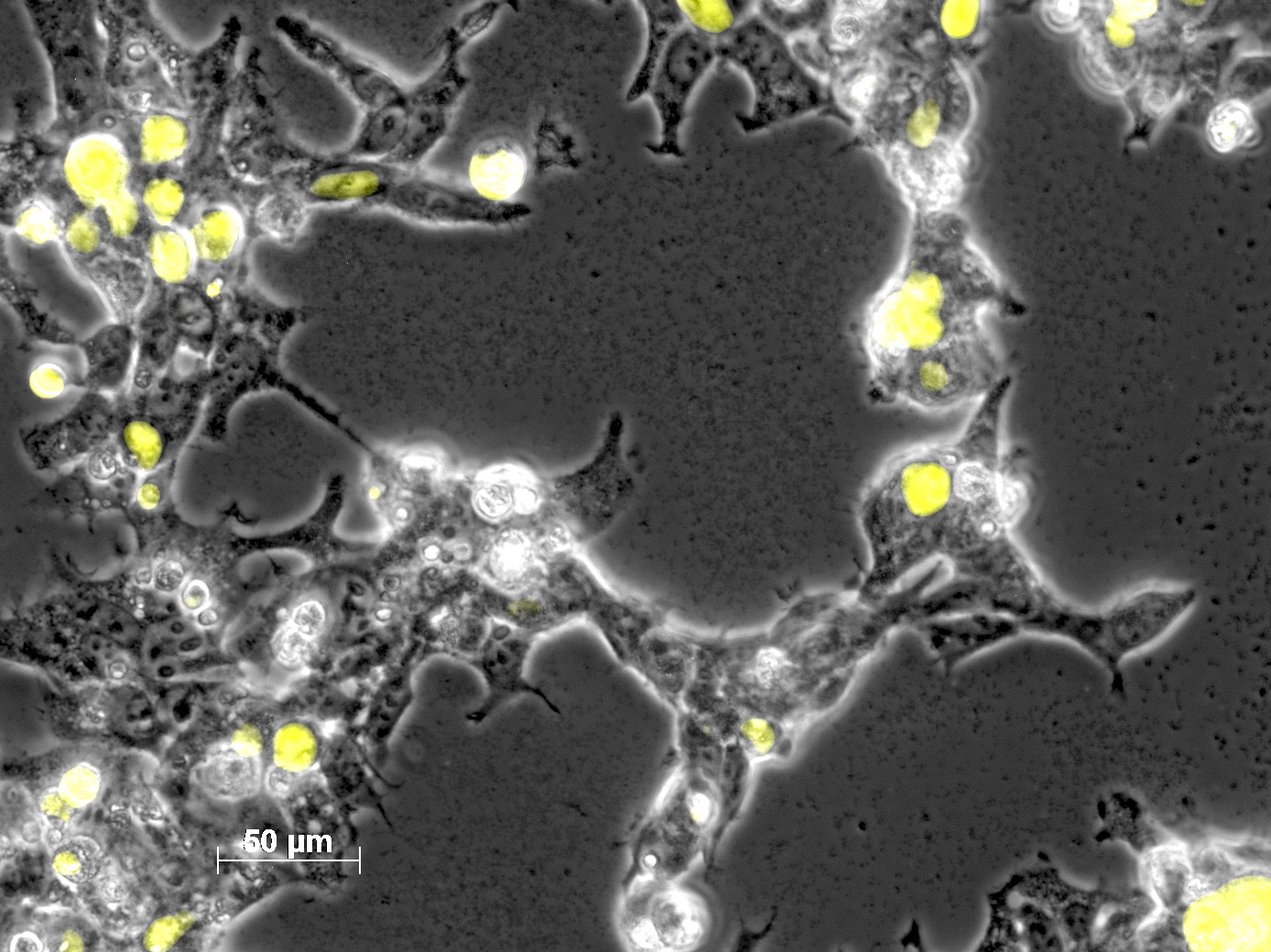
Impressions of transfection of AAV293 with pCerulean_VP1up_NLS_mVenus_VP2/3
Investigator: Adrian
Yesterday AAV293 cells were transfected with pCerulean_VP1up_NLS_mVenus_VP2/3 and GMK-TK30 as gene of interest. Today, nuclear localization of the produced mVenus_VP2/3 fusion protein was visible and can be nicely seen in the following pictures:
Conclusion: The VP2/3 fusion particle was transcribed and translated AND was transported back into the nucleus in order to be packaged by the ITR-flanked gene of interest.
Cloning of hGH_rITR into pSB1C3_lITR_phTERT_beta-globin_CFP and lITR_phTERT_beta-globin_mGMK_TK30 into pSB1C3_hGH_rITR
Investigator: Stefan
Vector name:
- pSB1C3_lITR_phTERT_beta-globin_CFP (P682)
- pSB1C3_hGH_rITR (P186)
Insert name:
- pSB1C3_hGH_rITR (P186)
- pSB1C3_lITR_phTERT_beta-globin_mGMK_TK30 (P672)
- pSB1C3_lITR_phTERT_beta-globin_mGMK_TK30 (P673)
| components | volume for P186 /µl | volume of P186 X+E /µl | volume of P672 + P673 /µl | volume of P682 /µl |
| DNA | 12 | 12 | 5 | 14 |
| BSA (10x) | 2,5 | 2 | 2 | 2 |
| Buffer 4 (10x) | 3 | 2 | 2 | 2 |
| Enzyme XbaI | 1 | 1 | - | - |
| Enzyme PstI | 1 | - | - | 1 |
| Enzyme EcoI | - | 1 | 1 | - |
| Enzyme SpeI | - | - | 1 | 1 |
| H2O | 6 | 6 | 9 | - |
| Total volume (e.g. 15,20,25,30 µl) | 25 | 25 | 20 | 20 |
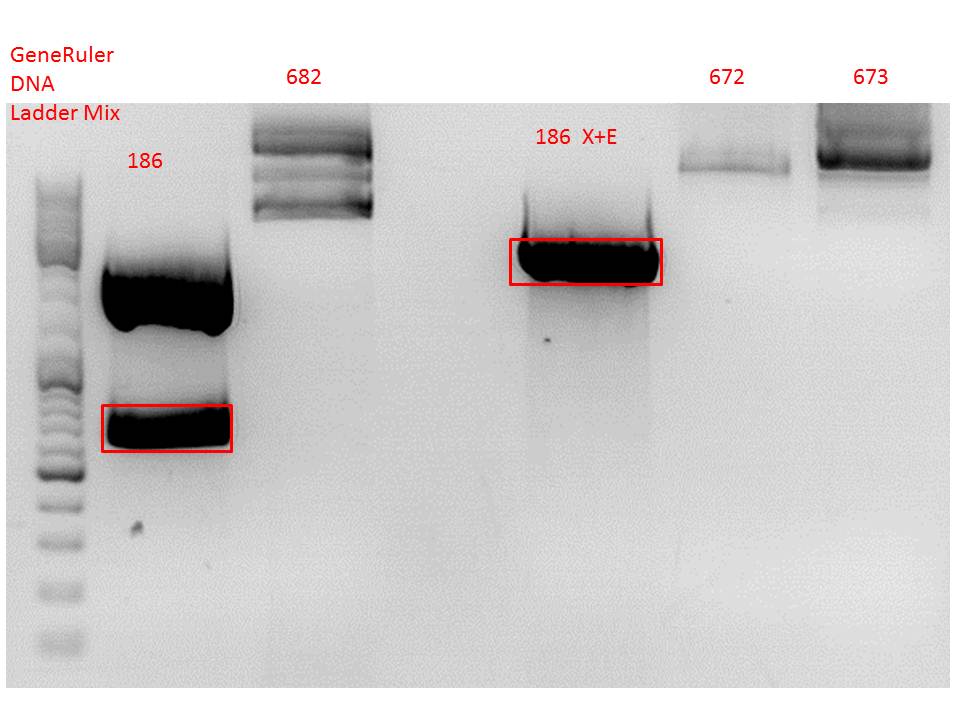
Gel:
0,5 g Agarose, 50 ml TAE (1%), 3 µl GELRED , at 115 Volt
Comment: Since the first gel revealed problems with samples 672, 673 and 682 another digestion was prepared (same approach as shown above), this time using a 0,8% gel and another loading dye (without SDS).
0,4 g Agarose, 50 ml TAE (0,8%), 3 µl GELRED , at 115 Volt
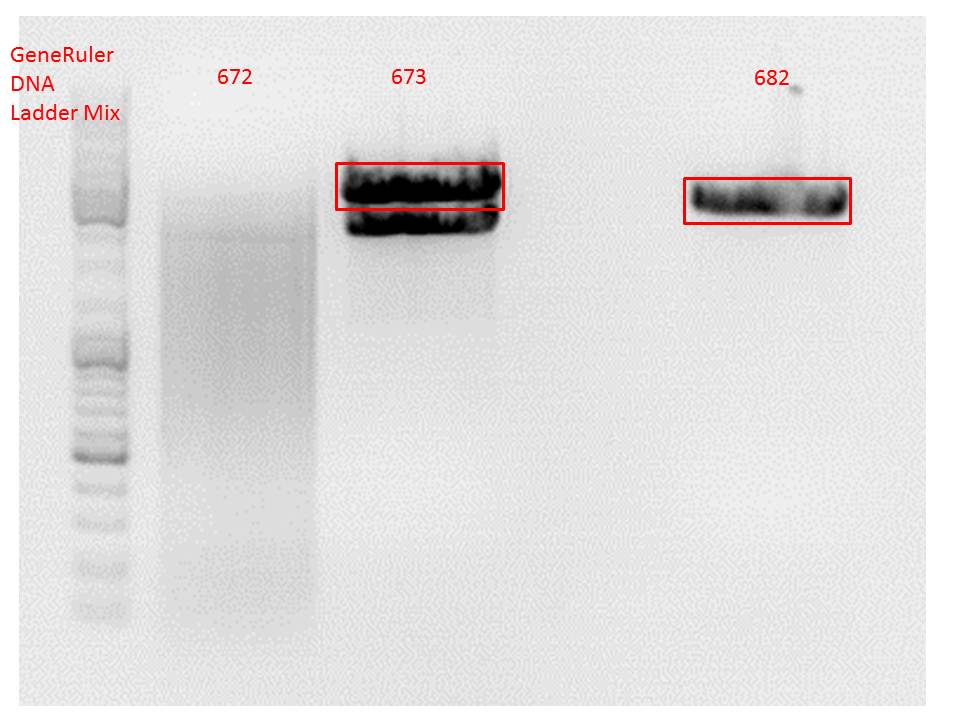
Comment: Since sample 672 showed no bands this sample was discarded and cloning continued only using 673.
Gel extraction:
Was performed according to protocol.
T4 Ligation:
Ligation was performed over night.
| ligation name | 186 (X+E) + 673 | 186 + 682 |
| volume of vector | 3,51 | 4,49 |
| volume of insert | 5,38 | 2,64 |
Transformation:
Will be done tomorrow using BL21 cells.
137. labday 02.10.2010
Mini Prep and test digestion of pGa14_MiddleLinker_VP2/3_HSPG-KO
Investigator: Hanna
Comment: In order to fuse the DARPin to the N-terminus of VP2/3_insCap and VP2/3_HSPG-KO, these motifs need to be fused to the Middle Linker (performed on 30.9.2010). In order to be able to immediately continue with cloning, BL21 cells were transformed and clones were picked in the noon. Now, the DNA needs to be preped and test digested in order to perform a 3-fragment-ligation with the DARPin and the pSB1C3_001_CMV plasmid. Unfortunately the cells of the VP2/3_insCap construct grew not densely enough. Therefore two new clones were picked from the plate and were inoculated in 10 mL LB medium. The other construct (pGa14_MiddleLinker_VP2/3_HSPG-KO) was preped and test digested.
TEST DIGESTION:
For both clones:
- DNA: 4 µL
- Buffer 4: 1 µL
- BSA: 1 µL
- EcoRI: 0.5 µL
- PstI: 0.5 µL
- H2O: 3 µL
Incubation: 1 hour.
Agarose gel: 0.8 %, 115 Volt, 1 hour.
Comment: The choice of these restriction enzymes was not clever, because if cloning didn't work, I expect one band at 2379 bp (the insert should hardly be seen) - if it worked one band at 2379 bp and a second at 2005 bp. Fortunately the bands could be separated ans revealed positive results for both clones :
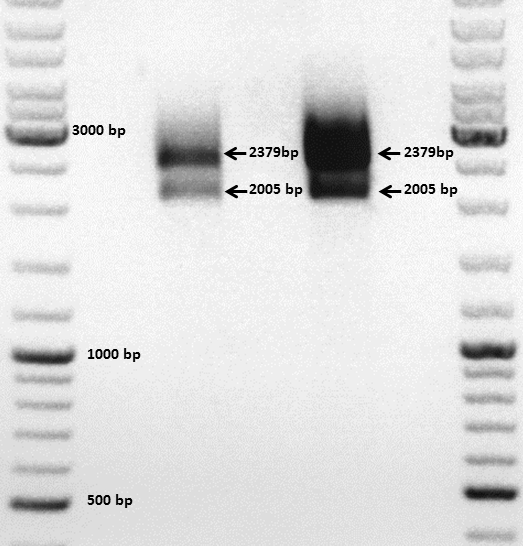
Next steps:
- Mini-Prep and test digestion of pGa14_MiddleLinker_VP2/3_ins Cap.
- 3 fragment ligation of MiddleLinker_VP2/3_ins Cap or MiddleLinker_VP2/3_HSPG-KO + DARPin + pSB1C3_001_CMV backbone.
Sequencing analysis of HSPG-ko in several RepCap constructs
Investigator: Stefan
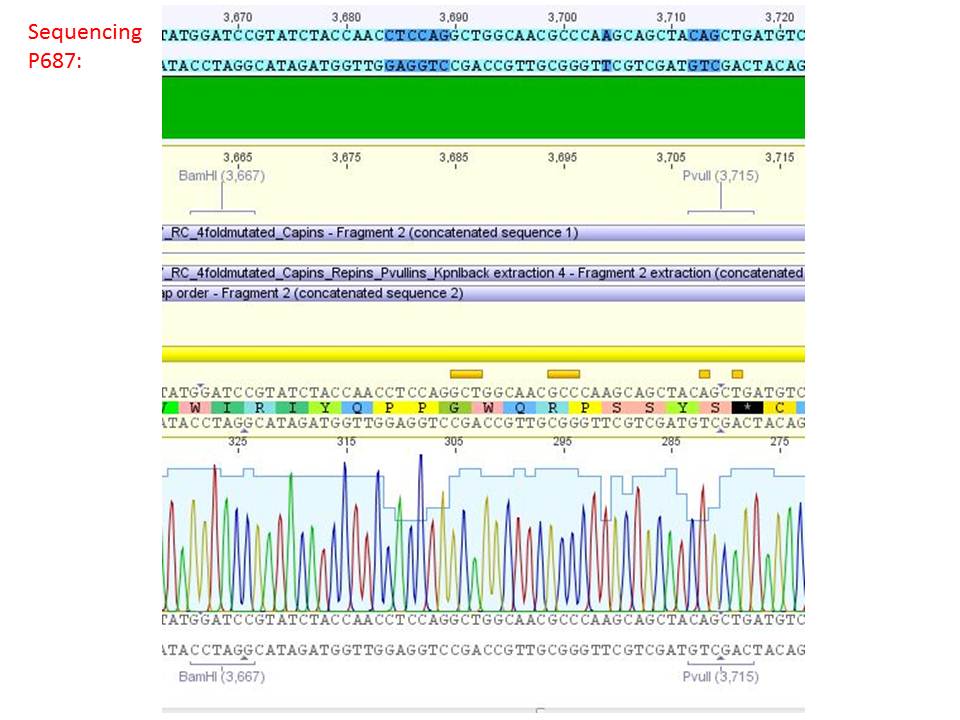
pSB1C3_001_RC_IRCK_HSPG-ko_P5tataless_RFC10 clone 1 (P687):
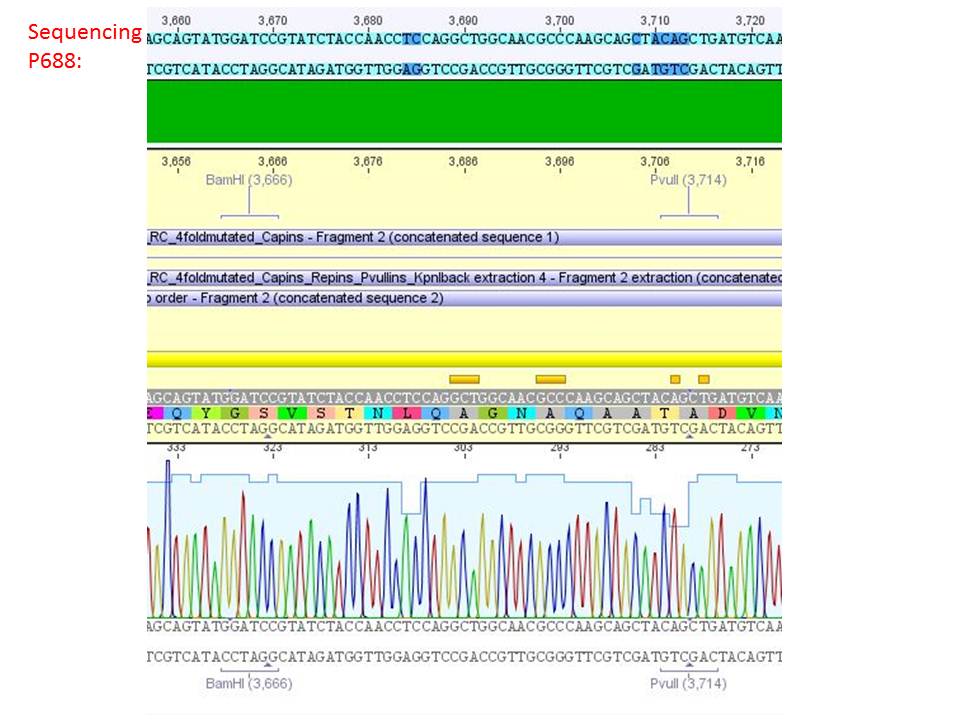
pSB1C3_001_RC_IRCK_HSPG-ko_P5tataless_RFC10 clone 2(P688):
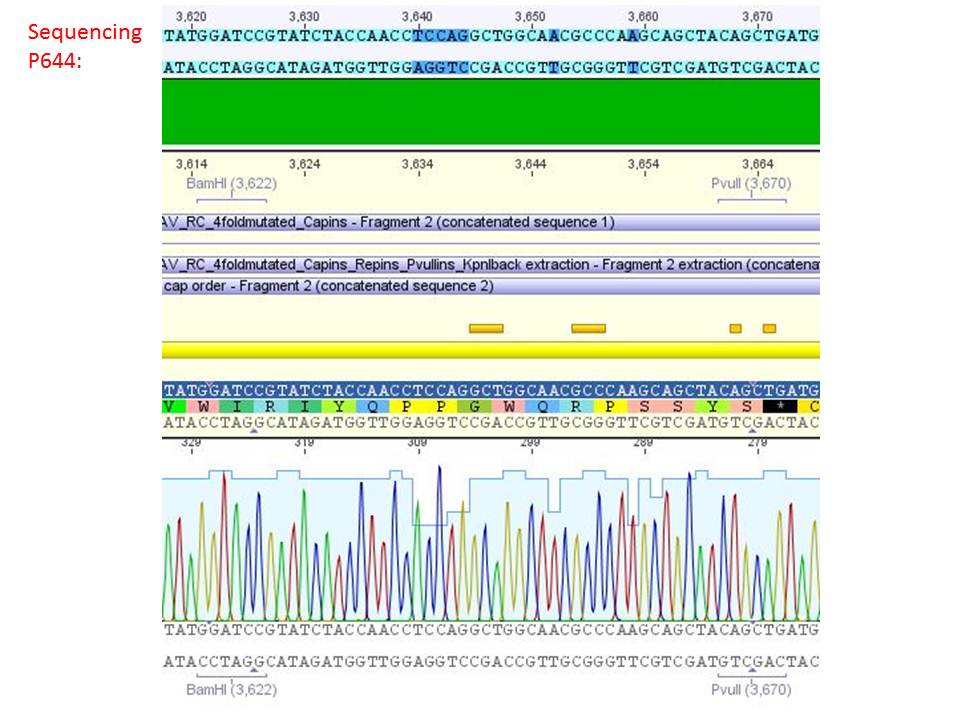
pSB1C3_001_RC_IRCK_VP1-ko_HSPG-ko_P5tataless cl1 (P644):
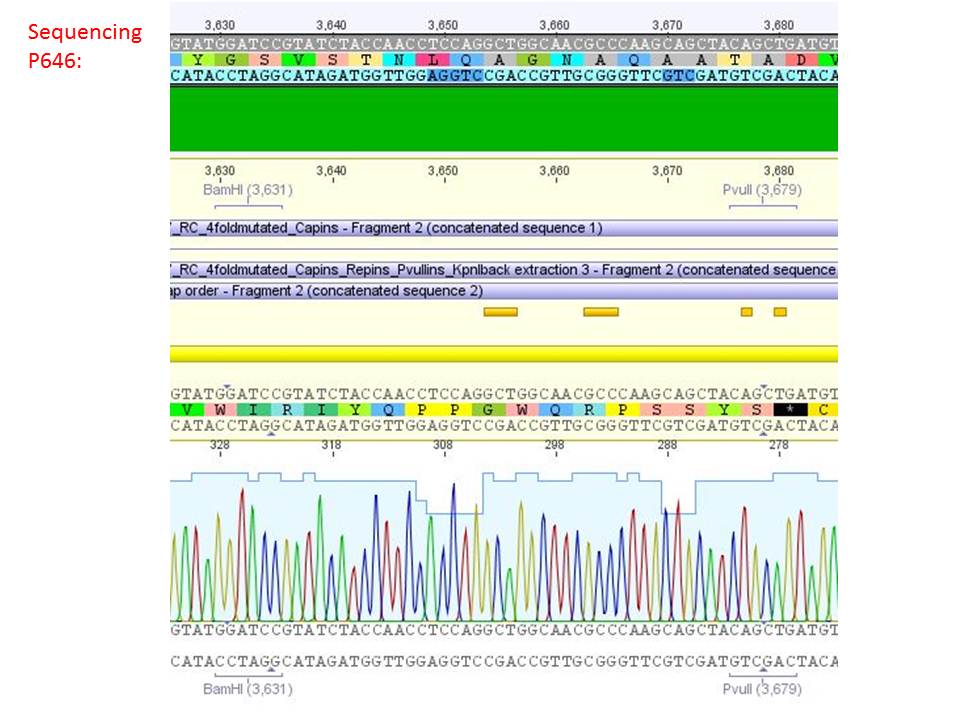
pSB1C3_001_RC_IRCK_VP2-ko_HSPG-ko_P5tataless cl1 (P646):
Comment: Sequencing results show that all plasmids contain the HSGP-ko. Working will be continued using P644, P646 and P687.
 "
"