Team:Freiburg Bioware/NoteBook/Labjournal/October
From 2010.igem.org
(→Impressions of transfection of AAV293 with pCerulean_VP1up_NLS_mVenus_VP2/3_insCap) |
(→136. labday 01.10.2010) |
||
| Line 120: | Line 120: | ||
<br/><br/><br/><br/><br/><br/><br/> | <br/><br/><br/><br/><br/><br/><br/> | ||
<b>Conclusion:</b> The VP2/3 fusion particle was transcribed and translated AND was transported back into the nucleus in order to be packaged by the ITR-flanked gene of interest. <br/> | <b>Conclusion:</b> The VP2/3 fusion particle was transcribed and translated AND was transported back into the nucleus in order to be packaged by the ITR-flanked gene of interest. <br/> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Cloning of VP2/3_BAP_HSPG-KO and VP2/3_His_HSPG-KO into pCerulean_Zegfr:1907_Middlelinker and VP2/3_His_HSPG-KO into pCerulean_VP1up_NLS_mVenus</b></p>==== | ||
| + | <b>Investigator: Stefan </b><br> | ||
| + | |||
| + | <p style="font-size:13px; color:red;"></p> | ||
| + | |||
| + | Vector name: | ||
| + | <ul> | ||
| + | <li>pCerulean_Zegfr:1907_Middlelinker (P408)</li> | ||
| + | <li>pCerulean_VP1up_NLS_mVenus (P426)</li> | ||
| + | </ul> | ||
| + | Insert name: | ||
| + | <ul> | ||
| + | <li>pBS1C3_001_VP2/3_587_BAP_HSPG-KO clone1 (P660)</li> | ||
| + | <li>pBS1C3_001_VP2/3_587_BAP_HSPG-KO clone2 (P661)</li> | ||
| + | <li>pBS1C3_001_VP2/3_587_His_HSPG-KO clone1 (P662)</li> | ||
| + | <li>pBS1C3_001_VP2/3_587_His_HSPG-KO clone2 (P663)</li> | ||
| + | </ul> | ||
| + | |||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''volume for inserts (P660 - P663) /µl'''|| align="right" |'''volume of P408 /µl''' || align="right" |'''volume of P426 /µl''' | ||
| + | |- | ||
| + | | DNA || align="right" |10 || align="right" |3|| align="right" |3 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" |3 || align="right" |2|| align="right" | 2 | ||
| + | |- | ||
| + | | Buffer 4 (10x)|| align="right" |3 || align="right" |2 || align="right" | 2 | ||
| + | |- | ||
| + | |Enzyme NgoMIV|| align="right" |1|| align="right" |- || align="right" |- | ||
| + | |- | ||
| + | |Enzyme PstI|| align="right" |1|| align="right" |1 || align="right" |1 | ||
| + | |- | ||
| + | |Enzyme MscI|| align="right" |1|| align="right" |- || align="right" |- | ||
| + | |- | ||
| + | |Enzyme AgeI|| align="right" |-|| align="right" |1 || align="right" |1 | ||
| + | |- | ||
| + | |H2O|| align="right" |11|| align="right" |11|| align="right" |11 | ||
| + | |- | ||
| + | |'''Total volume (e.g. 15,20,25,30 µl)'''|| align="right" | 30|| align="right" | 20|| align="right" |20 | ||
| + | |} | ||
| + | |||
| + | <br /> | ||
| + | <b>Gel:</b><br /> | ||
| + | 0,5 g Agarose, 50 ml TAE (1%), 3 µl GELRED , at 115 Volt<br /> | ||
| + | |||
| + | |||
| + | <br/> | ||
| + | [[Image:Freiburg10 digestion cloning BAP His to Affi Middle.jpg|450px|]]<br/> | ||
| + | |||
| + | <br/> | ||
| + | |||
| + | |||
| + | <b>Gel extraction</b>: <br> | ||
| + | Was performed according to protocol. | ||
| + | |||
| + | <br> | ||
| + | <b>T4 Ligation</b>: <br> | ||
| + | {| border="1" | ||
| + | |ligation name || align="right" |408 + 660|| align="right" |408 + 661|| align="right" |408 + 662|| align="right" |408 + 663|| align="right" |426 + 662|| align="right" |426 + 663 | ||
| + | |- | ||
| + | |volume of vector || align="right" |3,4 || align="right" | 3,05|| align="right" | 3,35|| align="right" | 2,56|| align="right" | 2,74|| align="right" | 3,13 | ||
| + | |- | ||
| + | |volume of insert|| align="right" |4,6 || align="right" |4,95|| align="right" |4,65|| align="right" |5,44|| align="right" |5,26|| align="right" |4,87 | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | <br> | ||
| + | <b>Transformation</b>: <br> | ||
| + | Was performed according to standard protocol using BL21 cells. | ||
| + | <br/> | ||
| + | |||
<html> | <html> | ||
</div> | </div> | ||
</html> | </html> | ||
Revision as of 21:45, 1 October 2010
136. labday 01.10.2010
mini preps of CD clones
Investigator: Kira
c(p689)= 115 ng/ul
c(690)= 151, 20 ng/ul
c(691)= 122,27 ng/ul
c(p692) = 126,42 ng/ul
test digestion of CD clones
Investigator: Kira
| Components | sample Volume/µL |
| DNA | 4,0 µl |
| BSA (10x) | 2 µl |
| Buffer no. 4 | 2,0 µl |
| Enzyme 1 XbaI | 1,0 µl |
| Enzyme 2 AgeI | 1,5 µl |
| H2O | 9,5 µl |
| Total volume | 20 |
incubation @ 37 C for approx. 2 h
1% agarose gel
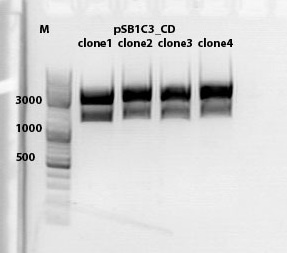
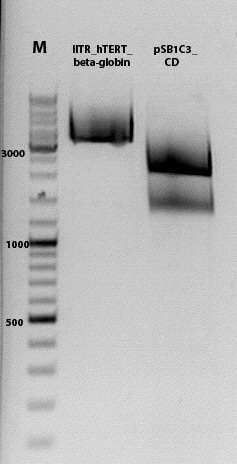
Biobrick assembly pSB1C3_lITR_hTERT_beta-globin_CD
Investigator: Kira
c(pSB1C3_lITR_hTERT_beta-globin)= 333 ng/ul
c(pSB1C3_CD)= 151 ng/ul
| Components | vector Volume/µL | insert Volume/µL |
| DNA | 4,5 µl | 6 |
| BSA (10x) | 3 µl | 3 |
| Buffer no. 4 | 3,0 µl | 3 |
| Enzyme 1 XbaI | 0 µl | 1,5 |
| Enzyme 2 SpeI | 1,5 µl | 0 |
| Enzyme 3 PstI-HF | 1,0 | 1 |
| H2O | 17 | 15,5 |
| Total volume | 25 |
incubation @ 37 C for approx. 2 h
1% agarose gel
Ligation
DNA-mix: 8 ul (vector 4,6ul)+(insert 3,4 ul)
T4 ligase: 1 ul
T4 buffer: 1 ul
Incubation @ RT for 30 min
Transformation was performed according to the standard protocol w BL21 cells.
Sequencing results of pCerulean_Zegfr:1907_MiddleLinker_VP2/3_HSPG-KO
Investigator: Hanna
Comment: All N-terminal fusion approaches with VP2/3_HSPG-KO revealed positive results except of pCerulean_Zegfr:1907_MiddleLinker_VP2/3_HSPG-KO. Another clone was picked, preped, test digested and sent for sequencing.
Conclusion: Sequencing results revealed positive results.
Picking clones of pGA14_MiddleLinker_VP2/3_insCap and pGA14_MiddleLinker_VP2/3_HSPG-KO
Investigator: Hanna
Unfortunately there grew a bacteria lawn over night - it was hardly not possible to pick clones. Nevertheless I tried and picked 2 clones of pGA14_MiddleLinker_VP2/3_insCap and pGA14_MiddleLinker_VP2/3_HSPG-KO.
To do: Mini-Prep and test digestion.
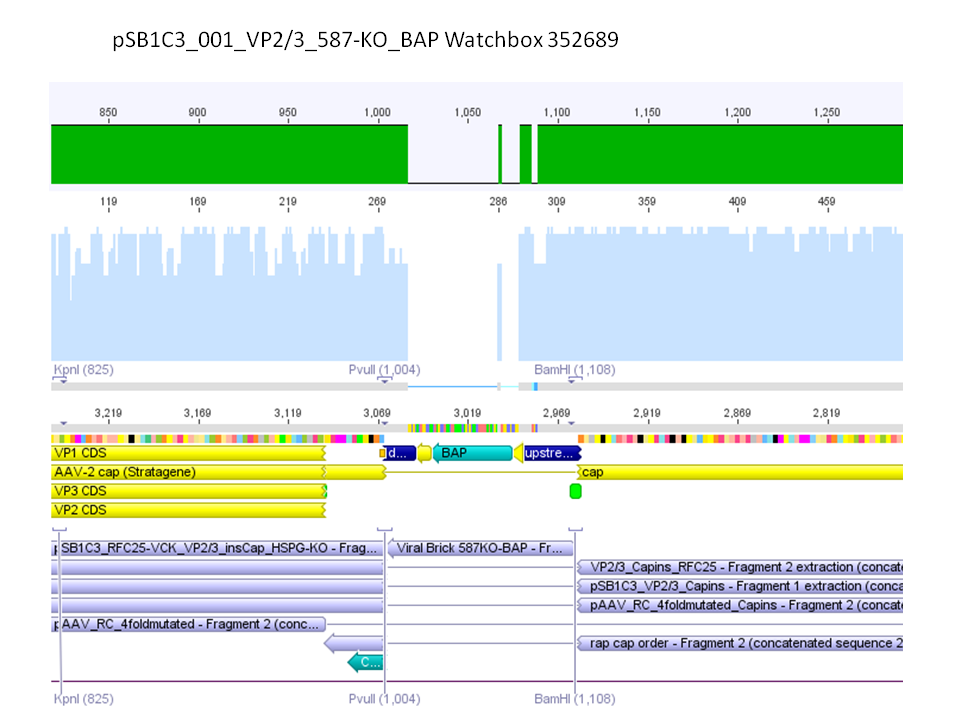
Sequencing results of pSB1C3_001_VP2/3_587-KO_BAP and pSB1C3_001_VP2/3_587-KO_6xHis
Investigator: Hanna
Comment: For the creation of our super constructs, the His-Tag and the BAP motif need to be cloned into VP2/3 for N-terminal fusion to VP2. Sequencing results showed that the 6xHis and BAP motif was not cloned into VP2/3_insCap.
To do: Clone 587-KO_6xHis and 587-KO_BAP into pSB1C3_001_VP2/3_insCap 1. via digestion of inserts and 2. via hybridization of referring oligos - digestion of vector with 800-900 ng.
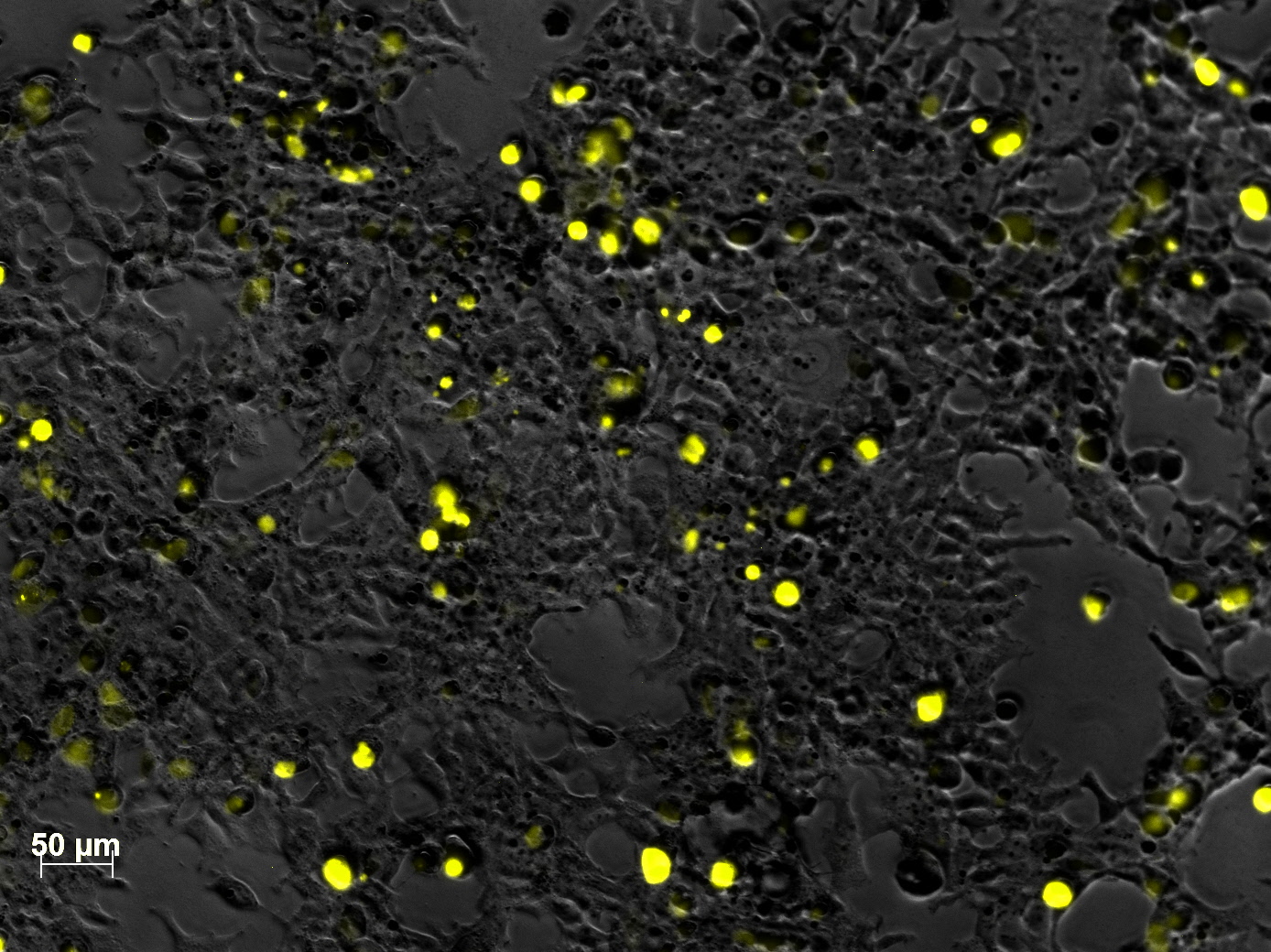
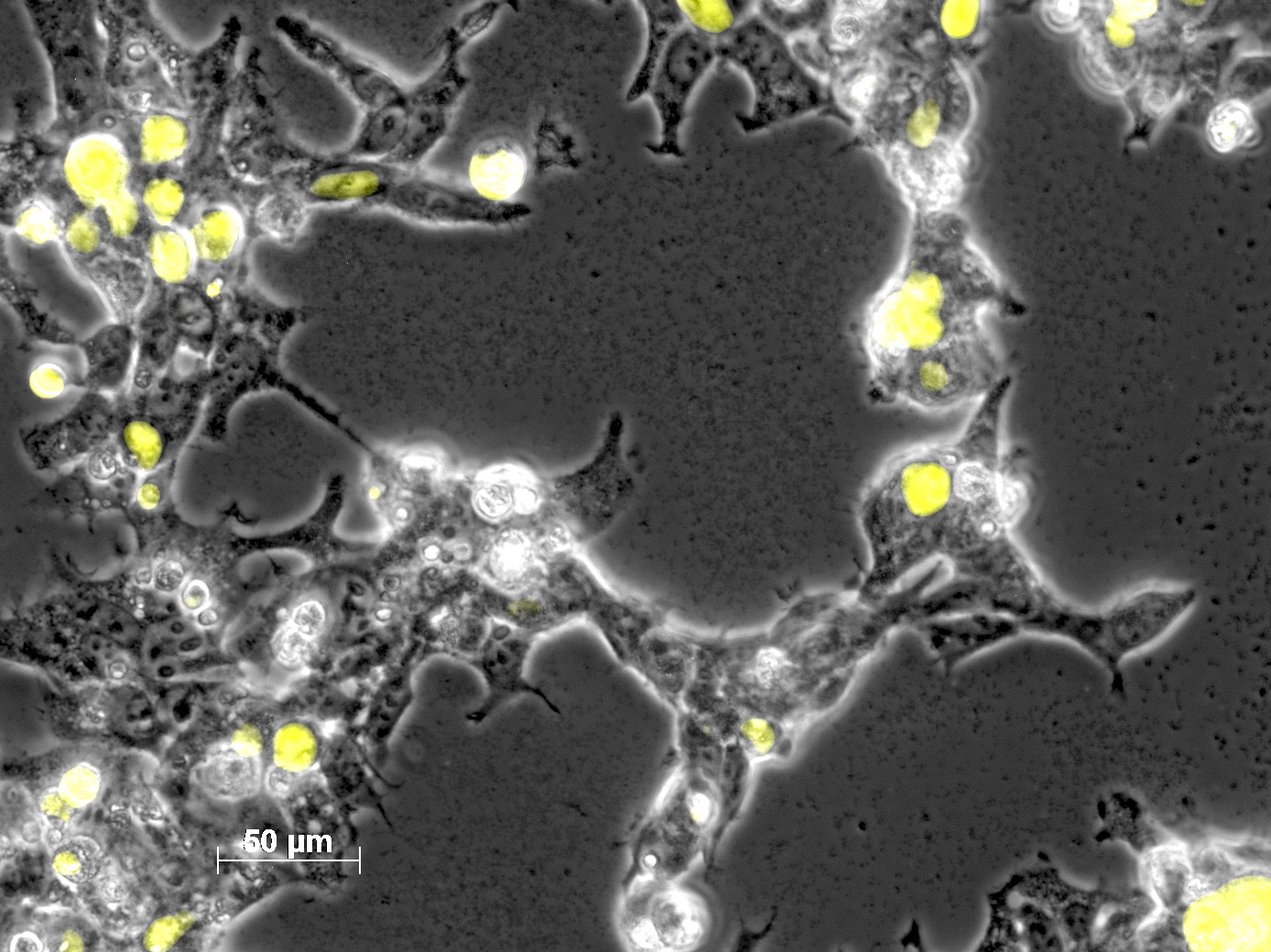
Impressions of transfection of AAV293 with pCerulean_VP1up_NLS_mVenus_VP2/3
Investigator: Adrian
Yesterday AAV293 cells were transfected with pCerulean_VP1up_NLS_mVenus_VP2/3 and GMK-TK30 as gene of interest. Today, nuclear localization of the produced mVenus_VP2/3 fusion protein was visible and can be nicely seen in the following pictures:
Conclusion: The VP2/3 fusion particle was transcribed and translated AND was transported back into the nucleus in order to be packaged by the ITR-flanked gene of interest.
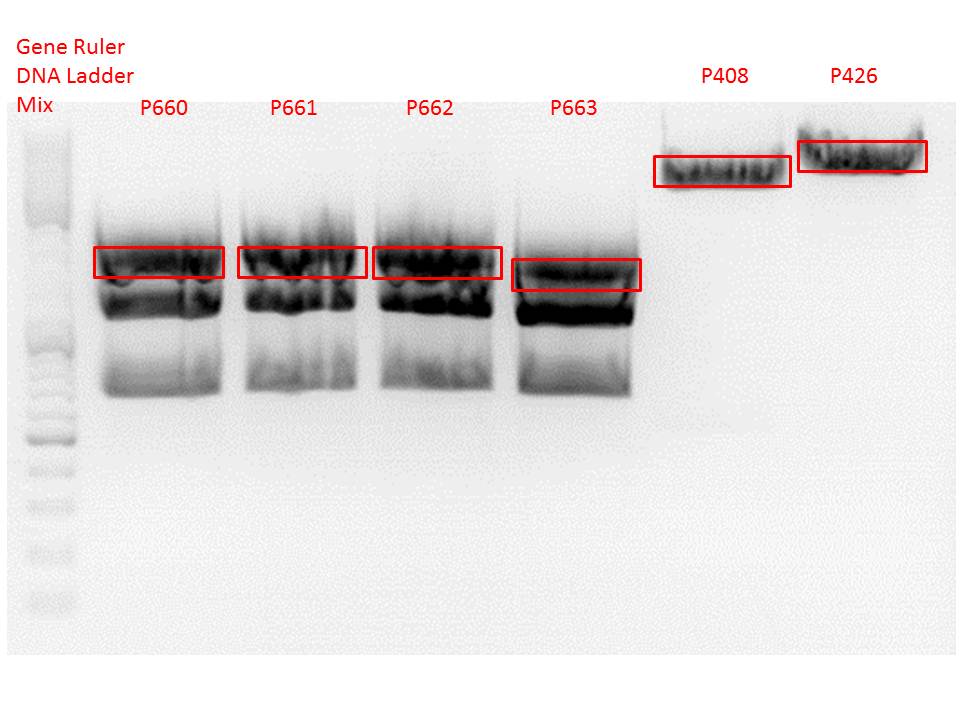
Cloning of VP2/3_BAP_HSPG-KO and VP2/3_His_HSPG-KO into pCerulean_Zegfr:1907_Middlelinker and VP2/3_His_HSPG-KO into pCerulean_VP1up_NLS_mVenus
Investigator: Stefan
Vector name:
- pCerulean_Zegfr:1907_Middlelinker (P408)
- pCerulean_VP1up_NLS_mVenus (P426)
Insert name:
- pBS1C3_001_VP2/3_587_BAP_HSPG-KO clone1 (P660)
- pBS1C3_001_VP2/3_587_BAP_HSPG-KO clone2 (P661)
- pBS1C3_001_VP2/3_587_His_HSPG-KO clone1 (P662)
- pBS1C3_001_VP2/3_587_His_HSPG-KO clone2 (P663)
| components | volume for inserts (P660 - P663) /µl | volume of P408 /µl | volume of P426 /µl |
| DNA | 10 | 3 | 3 |
| BSA (10x) | 3 | 2 | 2 |
| Buffer 4 (10x) | 3 | 2 | 2 |
| Enzyme NgoMIV | 1 | - | - |
| Enzyme PstI | 1 | 1 | 1 |
| Enzyme MscI | 1 | - | - |
| Enzyme AgeI | - | 1 | 1 |
| H2O | 11 | 11 | 11 |
| Total volume (e.g. 15,20,25,30 µl) | 30 | 20 | 20 |
Gel:
0,5 g Agarose, 50 ml TAE (1%), 3 µl GELRED , at 115 Volt
Gel extraction:
Was performed according to protocol.
T4 Ligation:
| ligation name | 408 + 660 | 408 + 661 | 408 + 662 | 408 + 663 | 426 + 662 | 426 + 663 |
| volume of vector | 3,4 | 3,05 | 3,35 | 2,56 | 2,74 | 3,13 |
| volume of insert | 4,6 | 4,95 | 4,65 | 5,44 | 5,26 | 4,87 |
Transformation:
Was performed according to standard protocol using BL21 cells.
 "
"