Team:TU Delft/22 July 2010 content
From 2010.igem.org
Ravandervalk (Talk | contribs) (→Salt tolerance) |
(→Lab work) |
||
| Line 116: | Line 116: | ||
A number of colonies look promising! To check if they really are the BioBricks we want, tomorrow we will do a plasmid isolation with the cultures of lane 2, 4, 6, 7, 9 and 14. We will cut the isolated plasmids with various restriction enzymes and analyze the digestion products on gel. | A number of colonies look promising! To check if they really are the BioBricks we want, tomorrow we will do a plasmid isolation with the cultures of lane 2, 4, 6, 7, 9 and 14. We will cut the isolated plasmids with various restriction enzymes and analyze the digestion products on gel. | ||
| + | |||
| + | ==Characterisation of Anderson RBS sequences== | ||
| + | |||
| + | ====Assembly of reference construct==== | ||
| + | |||
| + | =====Method 3===== | ||
| + | |||
| + | The Amp plates contained very few transformants. These were [https://2010.igem.org/Team:TU_Delft#page=protocols/colony_PCR single colony PCR'd] and run on [https://2010.igem.org/Team:TU_Delft#page=protocols/agarose_gel 1% agarose gel]: | ||
| + | |||
| + | |||
| + | |||
| + | =====Method 4===== | ||
| + | |||
| + | After having stored yesterday's Amp plates in the fridge it was decided to screen once more for GFP fluorescence. This time multiple fluorescing colonies were found under UV excitation. Three of the colonies were brought over to 5 mL of LB+AMP for over night culturing and characterization. | ||
Revision as of 21:22, 16 August 2010
Contents |
Lab work
Alkane degradation
There were some colonies on Tuesday's plates! We had left the plates @ 37°C yesterday after having seen that there were no colonies. When checking this morning on all plates (except the negative control) there were a few colonies! (2-50 colonies). Chances are it's not what we're looking for, but maybe they are good transformants... to check we will do a colony PCR.
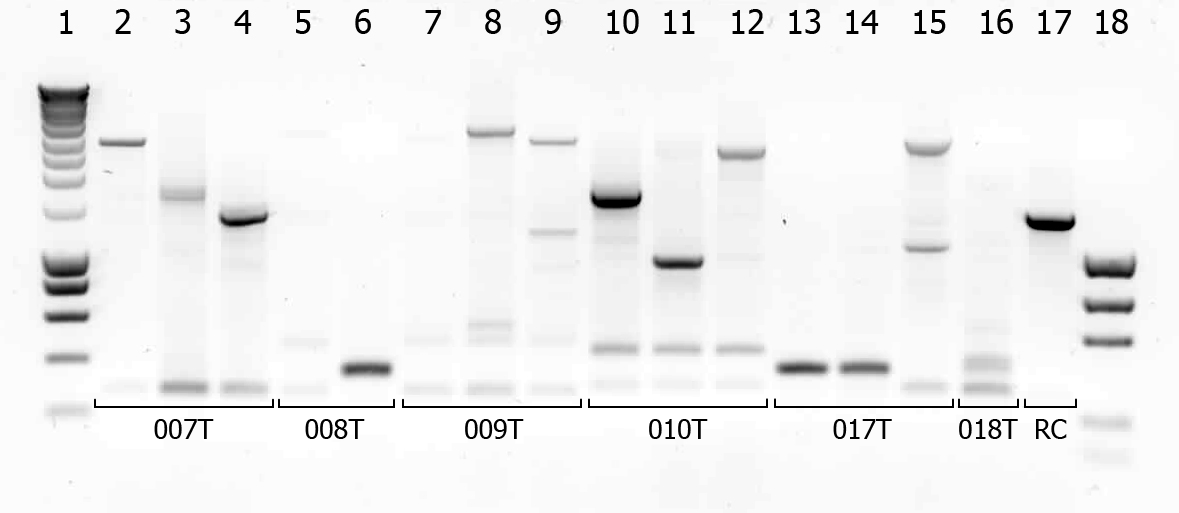
| # | Description | Expected length (bp) | Primers | Status |
| 1 | SmartLadder | n/a | n/a | n/a |
| 2 | Transformant #1 of ligation mix 007T | 1616 | G00100 + G00101 | |
| 3 | Transformant #2 of ligation mix 007T | 1616 | G00100 + G00101 | |
| 4 | Transformant #3 of ligation mix 007T | 1616 | G00100 + G00101 | |
| 5 | Transformant #1 of ligation mix 008T | 551 | G00100 + G00101 | |
| 6 | Transformant #2 of ligation mix 008T | 551 | G00100 + G00101 | |
| 7 | Transformant #1 of ligation mix 009T | 551 | G00100 + G00101 | |
| 8 | Transformant #1 of ligation mix 009T | 560 | G00100 + G00101 | |
| 9 | Transformant #1 of ligation mix 009T | 560 | G00100 + G00101 | |
| 10 | Transformant #1 of ligation mix 010T | 1657 | G00100 + G00101 | |
| 11 | Transformant #1 of ligation mix 010T | 1657 | G00100 + G00101 | |
| 12 | Transformant #1 of ligation mix 010T | 1657 | G00100 + G00101 | |
| 13 | Transformant #1 of ligation mix 017T | 1130 | G00100 + G00101 | |
| 14 | Transformant #1 of ligation mix 017T | 1130 | G00100 + G00101 | |
| 15 | Transformant #1 of ligation mix 017T | 1130 | G00100 + G00101 | |
| 16 | Transformant #1 of ligation mix 018T | 1874 | G00100 + G00101 | |
| 17 | Transformant #1 of Red colony | 1360 | G00100 + G00101 |
A number of colonies look promising! To check if they really are the BioBricks we want, tomorrow we will do a plasmid isolation with the cultures of lane 2, 4, 6, 7, 9 and 14. We will cut the isolated plasmids with various restriction enzymes and analyze the digestion products on gel.
Characterisation of Anderson RBS sequences
Assembly of reference construct
Method 3
The Amp plates contained very few transformants. These were single colony PCR'd and run on 1% agarose gel:
Method 4
After having stored yesterday's Amp plates in the fridge it was decided to screen once more for GFP fluorescence. This time multiple fluorescing colonies were found under UV excitation. Three of the colonies were brought over to 5 mL of LB+AMP for over night culturing and characterization.
 "
"