Team:Cambridge/Bioluminescence/Bacterial Codon optimisation
From 2010.igem.org
(→What is codon usage?) |
|||
| Line 2: | Line 2: | ||
{{:Team:Cambridge/Templates/headerbar|colour=#386abc|title=Project Vibrio: Bacterial Codon Optimisation}} | {{:Team:Cambridge/Templates/headerbar|colour=#386abc|title=Project Vibrio: Bacterial Codon Optimisation}} | ||
=What is codon usage?= | =What is codon usage?= | ||
| - | + | {{:Team:Cambridge/RightImage|image=codons.jpg}} | |
One of the fascinating features of life is the '''universal genetic code'''. In all known organisms, from bacteria to man, the same triplets of DNA bases code for the same amino acids. However this does not mean that all species encode their genomes in exactly the same way. | One of the fascinating features of life is the '''universal genetic code'''. In all known organisms, from bacteria to man, the same triplets of DNA bases code for the same amino acids. However this does not mean that all species encode their genomes in exactly the same way. | ||
The code is ''redundant'': a number of triplets code for the same amino acid. While all species are able to translate any sequence of DNA interchangeably, E. coli prefers to use certain triplets to code for certain amino acids which may be different to the ones we use. This 'preference' is reflected in the levels of tRNA which match such a triplet. In this project we resynthesised a number of genes ''de novo'' and thus were able to codon optimise them for expression in E. coli. | The code is ''redundant'': a number of triplets code for the same amino acid. While all species are able to translate any sequence of DNA interchangeably, E. coli prefers to use certain triplets to code for certain amino acids which may be different to the ones we use. This 'preference' is reflected in the levels of tRNA which match such a triplet. In this project we resynthesised a number of genes ''de novo'' and thus were able to codon optimise them for expression in E. coli. | ||
Revision as of 03:26, 27 October 2010

What is codon usage?
Team:Cambridge/RightImage One of the fascinating features of life is the universal genetic code. In all known organisms, from bacteria to man, the same triplets of DNA bases code for the same amino acids. However this does not mean that all species encode their genomes in exactly the same way. The code is redundant: a number of triplets code for the same amino acid. While all species are able to translate any sequence of DNA interchangeably, E. coli prefers to use certain triplets to code for certain amino acids which may be different to the ones we use. This 'preference' is reflected in the levels of tRNA which match such a triplet. In this project we resynthesised a number of genes de novo and thus were able to codon optimise them for expression in E. coli.
Improved translational speed
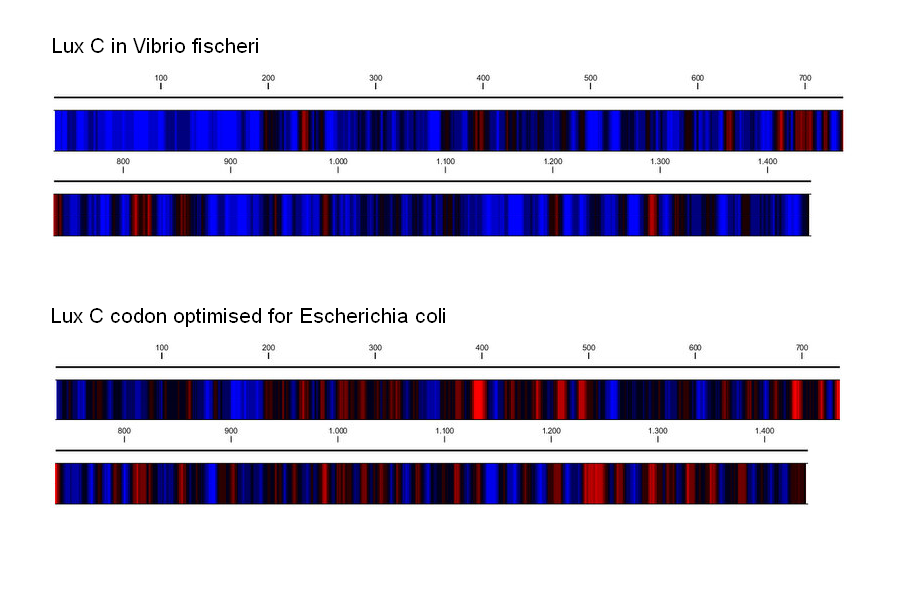
Starting with the DNA sequence of the Vibrio fischeri lux operon found on the NCBI database, we used a number of tools to replace the codons used with the most common codons found in the E.coli genome. To achieve optimal expression of the Lux operon in E.coli, we had the operon re-synthesized after optimising the usage of codons. This conserves the sequence of amino acids in the gene products, but improves the rate of translation, as more common tRNAs are recruited. Codon usage optimization can yield dramatic increases in the expression of foreign genes, especially if they are introduced from less closely related species [http://www.sciencedirect.com/science?_ob=ArticleURL&_udi=B6TCW-4C8NKCY-3&_user=6094838&_coverDate=07%2F31%2F2004&_rdoc=1&_fmt=high&_orig=search&_origin=search&_sort=d&_docanchor=&view=c&_searchStrId=1511316646&_rerunOrigin=scholar.google&_acct=C000053194&_version=1&_urlVersion=0&_userid=6094838&md5=c970323b521fa1ec9d3050d4c4970eb1&searchtype=a Gustafsson et al. 2004]
Altered G-C content
DNA curvature is increased by sequences rich in A-T or G-C pairs. The natural V.fischeri Lux operon, and especially its intergenic regions, contains stretches rich in A-T, resulting in the curvature that H-NS proteins bind to preferentially. Changing the coding DNA sequence also meant changing the curvature of the DNA, which affects the binding affinity of H-NS proteins. To alleviate the repression that H-NS exerts, we took care to raise the G-C content of intergenic regions and coding sequences (at times resorting to suboptimal codons). According to a computational prediction, this resulted in greatly reduced DNA curvature, and thus hopefully to a reduced affinity for H-NS proteins.
Differential Expression
In Vibrio fischeri, LuxA and B are expressed at five times the levels of LuxC, D, E and G. Since all these genes are transcribed on the same mRNA, but have their own Ribosome Binding Sites, this is probably due to differences in codon usage ([http://www.annualreviews.org/doi/pdf/10.1146/annurev.mi.42.100188.001055 Meighen et al. 1988]). A number of rare codons are found in Lux C, D, E and G, but not in LuxA and B. Since we did not receive the newly synthesised LuxA and B in time, we constructed a new operon using our LuxC, D, E and G, and the LuxAB BioBrick that was put into the registry by the Edinburgh iGEM team 2009 (BBa_216008). These genes originate from Xenorhabdus luminescens. Compared to Vibrio fischeri, there is only limited amino acid identity in the Lux A and B genes (52% and 66% respectively). Yet the literature describes both as using the same substrates. [http://aem.asm.org/cgi/content/abstract/55/10/2607 Schmidt et al. 1989] describe positive complementation test using components from V. fisheri and Xenorhabdus luminescens. To use foreign genes from two very different donor species in one pathway in E.coli is an exciting test of our understanding of the processes involved in bacterial bioluminescence and of the power of synthetic biology in general. In this construct, LuxC, D, E and G are codon optimised, but LuxA and B are not. In order to adjust the ratio of gene expression between these genes to the state found in nature, we chose to put LuxA and B under a very strong, phage derived promoter (plambda) to be constitutively expressed. The other genes can now be put under any promoter to create a PoPS-to-light device. In conjunction with an inducible or repressible promoter, this could be used as a reporter device. To test the system, we placed the LuxCDEG under an arabinose induced pbad promoter (BBa_i0500).
Parts submitted to the registry
Ideally we would have submitted a codon-optimised version of the entire Lux operon. Unfortunately Mr Gene, the company we employed for synthesis, still had not completed the order two months after placing, and at the point of the wiki-freeze we still have not received the optimised versions of LuxA and B. In order to complete our aim of creating a PoPs->light device that can be placed under any promoter, we combined the optimised Vibrio LuxCDEG genes with the Edinburgh 2009 Xenorhabdus LuxAB. The assembly was achieved using the Gibson method. Since we only received LuxCD and LuxEG from Mr Gene two weeks before the documentation deadline, we could not properly characterise these parts. For a complete list of the BioBricks we submitted see the BioBricks section.
Codon optimised bacterial luminescence parts:
[http://partsregistry.org/Part:BBa_K325902 luxCD (BBa_K325902)] derived from V. fischeri, but optimised for E.coli
[http://partsregistry.org/Part:BBa_K325903 luxEG (BBa_K325903)] derived from V. fischeri, but optimised for E.coli
[http://partsregistry.org/Part:BBa_K325905 CDEG pLambda AB (BBa_K325905)] a PoPs->light device that can be placed under any promoter, hybrid device containing promoterless V. fisheri derived CDEG (optimised) and Xenorhabdus AB under a strong constitutive promoter
[http://partsregistry.org/Part:BBa_K325906 pbad CDEG pLambda AB (BBa_K325906)] the above part under the arabinose induced pbad promoter
 "
"