Team:Freiburg Bioware/NoteBook/Labjournal/August
From 2010.igem.org
(→Inoculation of CD colony) |
|||
| Line 4,654: | Line 4,654: | ||
One colony was picked from agar plate and dispersed into 10 ml DYT+ Amp and incubated over night @ 37 C. | One colony was picked from agar plate and dispersed into 10 ml DYT+ Amp and incubated over night @ 37 C. | ||
| + | |||
| + | ===91. labday 16.08.2010=== | ||
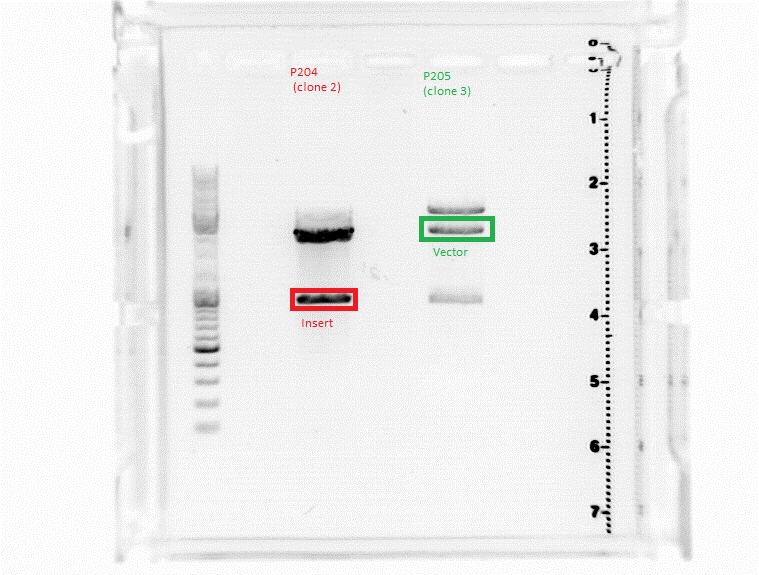
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Minipreps of P119, P122, P204, P205</b></p>==== | ||
| + | Investigator: Achim | ||
| + | |||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Cloning of pSB1C3_SspIdel_BLA clone 2 & 3 to get rid of mutations</b></p>==== | ||
| + | Investigator: Achim | ||
| + | |||
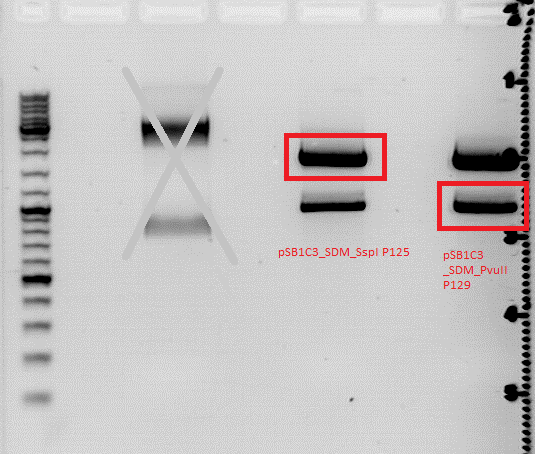
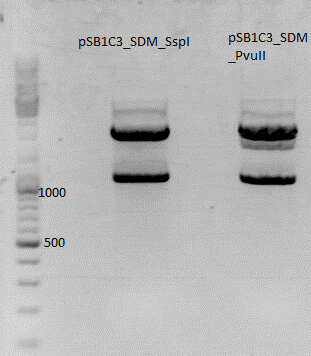
| + | *Sequencing showed that clone 2 had a mutation in the vector, clone 3 had one in the BLA sequence. Therefore, both were cut with BamHI and SalI, then the BLA fragment of clone 2 and the vector fragment of clone 3 were ligated. The final construct shouldn't contain any more mutations. | ||
| + | |||
| + | [[image:Freiburg10 16082010achim.jpg]] | ||
| + | <p style="font-size:15px; color:#ff00ff;">Nice picture, but where's the protocol? Results? (Hanna) ;) </p> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>New working solutions were prepared</b></p>==== | ||
| + | Investigator: Patrick | ||
| + | * DMEM | ||
| + | * Amp | ||
| + | * H2O millipore | ||
| + | * DYT | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Further pSB1C3_SDM_SspI_Bla14FM sequencing</b></p>==== | ||
| + | |||
| + | Two additional Clones were picked (clones 4 & 5) and sent for sequencing, labeled: iGEM4.1 and iGEM4.2. <br> | ||
| + | Used Primer: seq_pSB1C3_VR2_rev (O51) | ||
| + | |||
| + | Investigator: Patrick | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Mini prep of pKS-CD</b></p>==== | ||
| + | Investigator: Kira | ||
| + | |||
| + | c(pKS-CD) = 291, 39 ng/µl | ||
| + | |||
| + | Yielded DNA was used for site directed mutagenesis | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>SDM of PstI in pKS-CD</b></p>==== | ||
| + | Investigator: Jessica, Kira | ||
| + | |||
| + | Motivation: the desired gen contains 2 iGEM restrictions sites, thus in oder to use it for further cloning, these site have to be deleted. The first SDM was performed on PstI restriction site. | ||
| + | <ul> | ||
| + | <li>'''P223''' pKS_CD</li> | ||
| + | <li>'''O154''' CD_PstI FP</li> | ||
| + | <li>'''O155''' CD_PstI RP</li> | ||
| + | </ul> | ||
| + | {| border="1" | ||
| + | | '''component''' || '''Volume/µl''' | ||
| + | |- | ||
| + | | DNA: (1:20) || |0,5 | ||
| + | |- | ||
| + | | 10x reaction buffer: || |2,5 | ||
| + | |- | ||
| + | | primer forward PstI (1:10) || |0,56 | ||
| + | |- | ||
| + | | primer reverse PstI (1:10) || |0,56 | ||
| + | |- | ||
| + | | dNTP || |0,5 | ||
| + | |- | ||
| + | |DMSO || |0,5 | ||
| + | |- | ||
| + | |H2O || |14,38 | ||
| + | |- | ||
| + | |PfuTurbo DNA Polymerase || |0,5 | ||
| + | |} | ||
| + | <br /> | ||
| + | |||
| + | PCR | ||
| + | {| border="1" | ||
| + | | '''segment''' || '''cycles''' || '''temperature''' || '''time''' | ||
| + | |- | ||
| + | | 1 || |1 || |95 || |2min | ||
| + | |- | ||
| + | | 2 || |20 || |95 || |30sec | ||
| + | |- | ||
| + | | || | || |55 || | 1min | ||
| + | |- | ||
| + | | || | || |68 || | 4 min30 sec | ||
| + | |- | ||
| + | | 3 || | || | 4|| | hold | ||
| + | |} | ||
| + | <br /> | ||
| + | sample was digested with DpnI for 1h at 37°C | ||
| + | |||
| + | Transformation was performed according to the standard protocol. | ||
| + | <br/> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Biobrick production of pSB1C3_lITR_CMV_beta-globin_mVENUS_hgh_rITR and pSB1C3_lITR_CMV_mVENUS_hgh_rITR'''</p>==== | ||
| + | |||
| + | <b>Investigator: Anna, Anissa</b><br> | ||
| + | <p style="font-size:13px; color:#003399;"><b>Comments</b>: hgh_rITR was cloned one time into pSB1C3_lITR_CMV_beta-globin_mVENUS and one time into pSB1C3_lITR_CMV_mVENUS to receive a control for the improtance of beta-globin </p> | ||
| + | *<b>Digestion:</b> <br /> | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''volume of pBS1C3_hgh_rITR INSERT = p186 /µl''' || align="right" |'''volume of pBS1C3_lITR_CMV_beta-globin_mVENUS VECTOR = p208 /µl'''|| align="right" |'''volume of pBS1C3_lITR_CMV_mVENUS VECTOR = p207 /µl''' | ||
| + | |- | ||
| + | | DNA || align="right" |19|| align="right" |6 || align="right" |3,84 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" |3|| align="right" | 3|| align="right" |1,5 | ||
| + | |- | ||
| + | | Buffer 4 (10x)|| align="right" |3 || align="right"| 3 || align="right" |1,5 | ||
| + | |- | ||
| + | |SpeI° /XbaI*|| align="right" |1*|| align="right" |1° || align="right" |1° | ||
| + | |- | ||
| + | |PstI|| align="right" |1 || align="right" |1|| align="right" |1 | ||
| + | |- | ||
| + | |H2O|| align="right" |3 || align="right" |6 || align="right" |6,16 | ||
| + | |- | ||
| + | |'''Total volume'''|| align="right" |30|| align="right" |20 || align="right" |15 | ||
| + | |} | ||
| + | <br> | ||
| + | The samples were digested for 2 h at 37°C | ||
| + | <br/> | ||
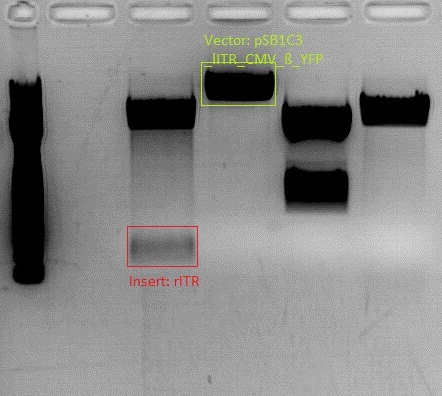
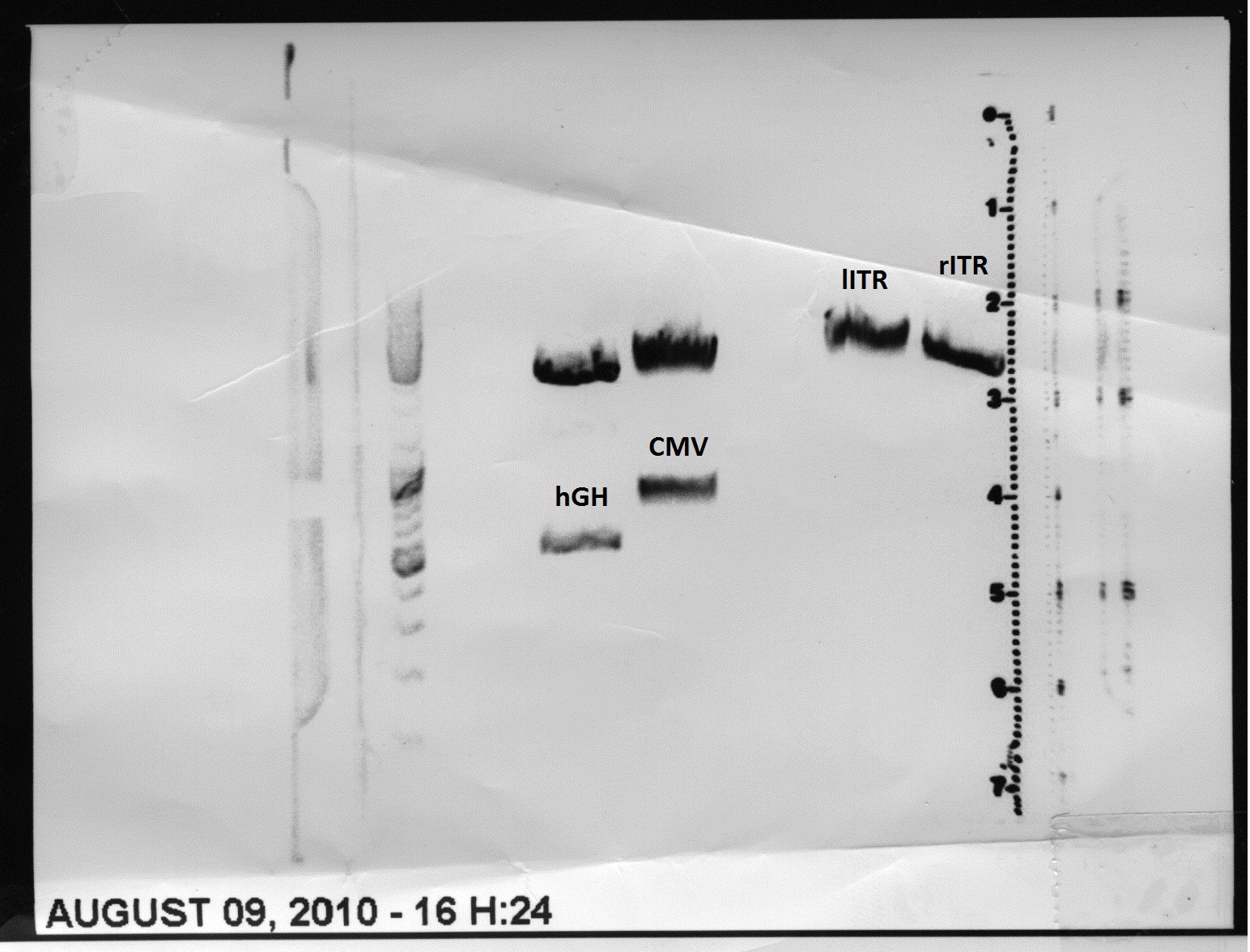
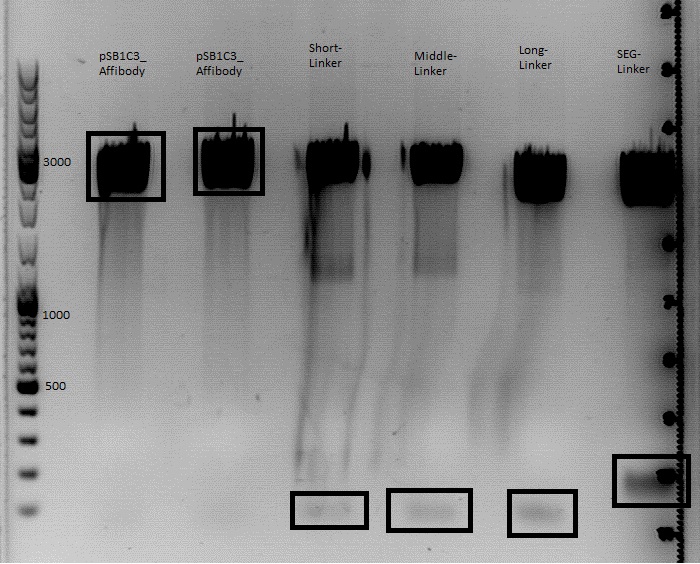
| + | * Gel: samples were loaded on a 1% agarose-gel. running time: 45 minutes | ||
| + | [[File:Freiburg10 psb1c3 lITR CMV ß-globin mVenus hgh rITR und psb1c3 lITR CMV mVenus hgh rITR.png|400px|]] | ||
| + | <br/> | ||
| + | *gelextraction was performed according with the standard-protocol | ||
| + | <br/> | ||
| + | *Ligation: | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''concentration/ ng/µl''' | ||
| + | |- | ||
| + | | p186 || align="right" |8,97 | ||
| + | |- | ||
| + | |p207 || align="right" |25,57 | ||
| + | |- | ||
| + | | p208|| align="right" |17,73 | ||
| + | |- | ||
| + | |} | ||
| + | <br/> | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''amount for ligation /µl''' | ||
| + | |- | ||
| + | | p186 (for p207)|| align="right" |4,2 | ||
| + | |- | ||
| + | |p207 || align="right" |3,8 | ||
| + | |- | ||
| + | |p186 (for p208)|| align="right" |4,7 | ||
| + | |- | ||
| + | | p208|| align="right" |3,3 | ||
| + | |- | ||
| + | | T4 ligation buffer|| align="right" |1 | ||
| + | |- | ||
| + | | T4 Ligase|| align="right" |1 | ||
| + | |- | ||
| + | |} | ||
| + | *Transformation: was performed according with the standard-protocol | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Preparation for Midi-Preps of B24 and B34'''</p>==== | ||
| + | |||
| + | Investigator: Anna | ||
| + | |||
| + | <ul><li>40 ml DYT was prepared with 40 µl Ampicillin and inoculated with B24 and B34, both were incubated over night in 37°C room.</li> | ||
| + | <br> | ||
| + | </ul> | ||
| + | <p style="font-size:13px; color:#ff00ff;">What is B24 and B34? Can you please be more precise - I also want to understand what's happening also when I am not able to check up in the excel sheets ;) (Hanna)</p> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Cloning of pAAV_RC (P50) with pMA_RC_insert (P190)</b></p>==== | ||
| + | Investigator: <b>Chris L.</B><br /> | ||
| + | |||
| + | *Vector: name: pAAV_RC <b>P50</b> | ||
| + | *Insert: name: pMA_RC_insert <b>P190</b> | ||
| + | |||
| + | *buffer used: 3 | ||
| + | *Restriction-enzymes used: | ||
| + | **BstEII (no. Lab: 17) | ||
| + | **SwaI (no.Lab: 135) | ||
| + | *DNA concentration (P50): 378,5 ng/µl | ||
| + | *DNA concentration (P190): 339,5 ng/µl | ||
| + | |||
| + | <br /> | ||
| + | |||
| + | <b>Digestion:</b> <br /> | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''volume of pMA_RC_insert /µl''' || align="right" |'''volume of pAAV_RC /µl''' | ||
| + | |- | ||
| + | | DNA || align="right" |4,4 || align="right" |2,6 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" |2 || align="right" | 2 | ||
| + | |- | ||
| + | | Buffer 3 (10x)|| align="right" |2 || align="right" |2 | ||
| + | |- | ||
| + | |SwaI (no.Lab:135)|| align="right" |0,5 || align="right" |0,5 | ||
| + | |- | ||
| + | |BstEII (no.Lab:17)|| align="right" |0,5 || align="right" |0,5 | ||
| + | |- | ||
| + | |H2O|| align="right" |10,6 || align="right" |12,4 | ||
| + | |- | ||
| + | |'''Total volume'''|| align="right" | 20|| align="right" |20 | ||
| + | |} | ||
| + | <br> | ||
| + | <p style="color:red;">Comments: Incubation at 25° after addition of SwaI for 1,5 hours. Then incubation at 60° after addition of BstEII for 1,5 hours.</p> <br /> | ||
| + | |||
| + | <b>Gel extraction:</b> <br /> | ||
| + | Preparation of gel:<br /> | ||
| + | 0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time: 45 minutes | ||
| + | <br /> | ||
| + | |||
| + | Expected sizes of constructs: | ||
| + | *pAAV_RC 6839 bp and 493 bp | ||
| + | *pMA_RC_Insert 3292 bp and 493 bp | ||
| + | <br> | ||
| + | {| border="1" | ||
| + | | | ||
| + | !Marker | ||
| + | !Sample P190, 20µl | ||
| + | !Sample P 50, 20µl | ||
| + | |- | ||
| + | !Lane | ||
| + | |3 | ||
| + | |5 | ||
| + | |7 | ||
| + | |- | ||
| + | |} | ||
| + | <br> | ||
| + | <b>Results:</b> <p style="color:#00bbff;"> Bands are really weak, tomorrow I`ll take more DNA in sample. | ||
| + | <br /> | ||
| + | <b>Next steps: Tomorrow repetition of experiment </b> </p> | ||
| + | <br /> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Repetition: Biobrick production of Rep40, Rep68, Rep78 and AAP'''</p>==== | ||
| + | |||
| + | '''Investigator: Stefan'''<br> | ||
| + | |||
| + | |||
| + | Aim of the experiment: | ||
| + | We want to produce biobricks from Rep40, Rep68, Rep78 and AAP. | ||
| + | |||
| + | *Plasmids used as template: | ||
| + | Rep_68_ex (p119): c = 470,6 ng/µl <br> | ||
| + | Rep_78_(p122): c = 566,8 ng/µl <br> | ||
| + | pAAV-RC containing AAP ORF (p50): c = 378,5 ng/µl <br> | ||
| + | |||
| + | |||
| + | |||
| + | *Primer used: | ||
| + | For Rep_68_ex: Praefix_68_78_ex & Suffix_40_68_ex<br> | ||
| + | For Rep_78_ex: Praefix_68_78_ex & Suffix_52_78_ex<br> | ||
| + | For AAP_ex: Praefix_AAP_ex & Suffix_AAP_ex | ||
| + | <br> | ||
| + | *'''PCR: | ||
| + | (was performed following the standard protocol) | ||
| + | <br> | ||
| + | {| border="1" | ||
| + | | '''Ingredients''' || align="right" |'''Volume / µl''' || align="right" |'''Rep68'''|| align="right" |'''Rep78''' || align="right" |'''AAP'''|| align="right" |'''Rep40''' | ||
| + | |- | ||
| + | | 5X Phusion HF buffer || align="right" |10 || align="right" | || align="right" | || align="right" | || align="right" | | ||
| + | |- | ||
| + | | 10 mM dNTP mix|| align="right" |1|| align="right" | || align="right" | || align="right" | || align="right" | | ||
| + | |- | ||
| + | | forward primer: || align="right" |2,5|| align="right" | || align="right" | || align="right" | || align="right" | | ||
| + | |- | ||
| + | | reverse primer: || align="right" |2,5|| align="right" | || align="right" | || align="right" | || align="right" | | ||
| + | |- | ||
| + | | DNA Template|| align="right" |***|| align="right" |2,5 µl || align="right" |2 µl|| align="right" |3 µl|| align="right" |1 µl | ||
| + | |- | ||
| + | | DMSO (2%)|| align="right" | || align="right" | - || align="right" | - || align="right" | 1 µl || align="right" |- | ||
| + | |- | ||
| + | | Phusion Polymerase|| align="right" |0,5|| align="right" | || align="right" | || align="right" | || align="right" | | ||
| + | |- | ||
| + | |H<sub>2</sub>O|| align="right" |*** || align="right" | 31 µl|| align="right" | 31,5 µl|| align="right" |29,5 µl || align="right" |31,5 µl | ||
| + | |- | ||
| + | |Total volume|| align="right" |50 | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | PCR program: | ||
| + | {| border="1" | ||
| + | |'''PCR Program'''|| align="right" |'''temperature/ °C'''|| align="right" |'''Time''' || align="right" |'''Rep68'''|| align="right" |'''Rep78'''|| align="right" |'''AAP''' || align="right" |'''Rep40''' | ||
| + | |- | ||
| + | |1|| align="right" |98 || align="right" |1min|| align="right" | || align="right" | || align="right" | || align="right" | | ||
| + | |- | ||
| + | |2|| align="right" |98 || align="right" |15s|| align="right" | || align="right" | || align="right" | || align="right" | | ||
| + | |- | ||
| + | |3|| align="right" |*** || align="right" |25s|| align="right" |63°C|| align="right" |62°C|| align="right" |64°C|| align="right" |63°C | ||
| + | |- | ||
| + | |4 (step 2-4 8x) || align="right" |72|| align="right" |***|| align="right" |24s || align="right" |27s || align="right" | 10s || align="right" |15s | ||
| + | |- | ||
| + | |5 || align="right" |98|| align="right" | 15s|| align="right" | || align="right" | || align="right" | || align="right" | | ||
| + | |- | ||
| + | |6|| align="right" |***|| align="right" |25s|| align="right" |68°C|| align="right" |64°C|| align="right" |68°C || align="right" |68°C | ||
| + | |- | ||
| + | |7 (step 5-7 17x)|| align="right" |72|| align="right" |***|| align="right" |24s|| align="right" |27s|| align="right" |10s || align="right" |15s | ||
| + | |- | ||
| + | |8|| align="right" |72|| align="right" |5min|| align="right" | || align="right" | || align="right" | || align="right" | | ||
| + | |- | ||
| + | |Hold|| align="right" |4°C|| align="right" | || align="right" | || align="right" | || align="right" | || align="right" | | ||
| + | |} | ||
| + | <br> | ||
| + | |||
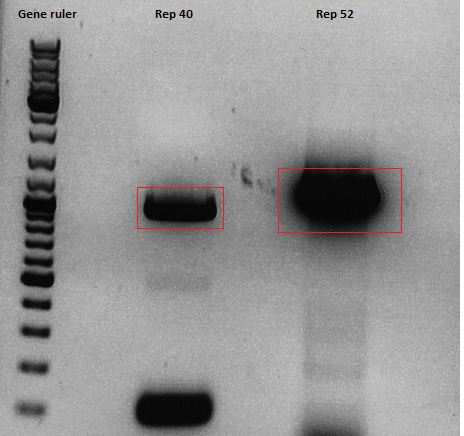
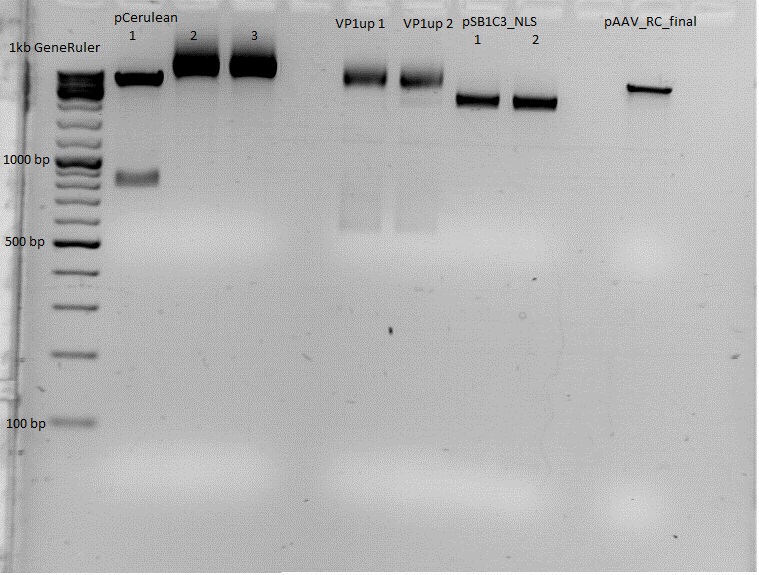
| + | Used agarose gel: | ||
| + | 0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time:45 minutes | ||
| + | |||
| + | [[File:Freiburg10 Rep40 68 78 AAP.png|400px|]]<br /> | ||
| + | <p style="font-size:13px; color:#68bbff;">'''''Comments''''': I forgot to dilute Rep68. Therefore, a new PCR will have to be performed tonight. </p> <br> | ||
| + | |||
| + | |||
| + | <p style="font-size:15px; font-weight: bold; color: blue;">Gel extraction</p> | ||
| + | <br /> | ||
| + | Gel extraction was perfomed according to protocol. | ||
| + | |||
| + | |||
| + | *Digestion of PCR products and vector: | ||
| + | |||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''PCR products /µl''' || align="right" |'''vector /µl''' | ||
| + | |- | ||
| + | | DNA || align="right" |29 || align="right" |11 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" |4 || align="right" | 2 | ||
| + | |- | ||
| + | | Buffer 4 (10x)|| align="right" |4 || align="right" |2 | ||
| + | |- | ||
| + | |Enzyme '''XbaI || align="right" |1 || align="right" |1 | ||
| + | |- | ||
| + | |Enzyme '''SpeI || align="right" |1 || align="right" |1 | ||
| + | |- | ||
| + | |H2O|| align="right" |1 || align="right" |3 | ||
| + | |- | ||
| + | |'''Total volume (e.g. 15,20,25,30 µl)'''|| align="right" | 40|| align="right" |20 | ||
| + | |} | ||
| + | <br /> | ||
| + | <p style="font-size:13px; color:#68bbff;">'''''Comments''''': Digestion was done with XbaI and SpeI, it has to be checked if the inserts are cloned into the vector in the right orientation. </p> <br> | ||
| + | |||
| + | *Purification of Rep40, 78 and AAP: | ||
| + | For the purification 200 µl of buffer PBI was used. | ||
| + | |||
| + | c(Rep40)= 6,5 ng/µl <br /> | ||
| + | c(Rep78)= 14,3 ng/µl <br /> | ||
| + | c(AAP)= 39,6 ng/µl | ||
| + | |||
| + | <br /> | ||
| + | |||
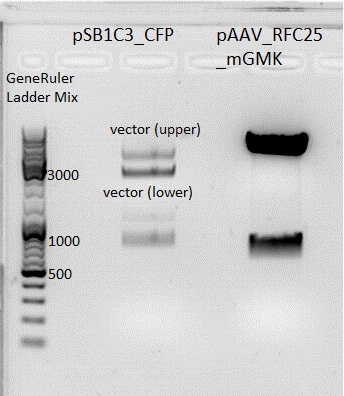
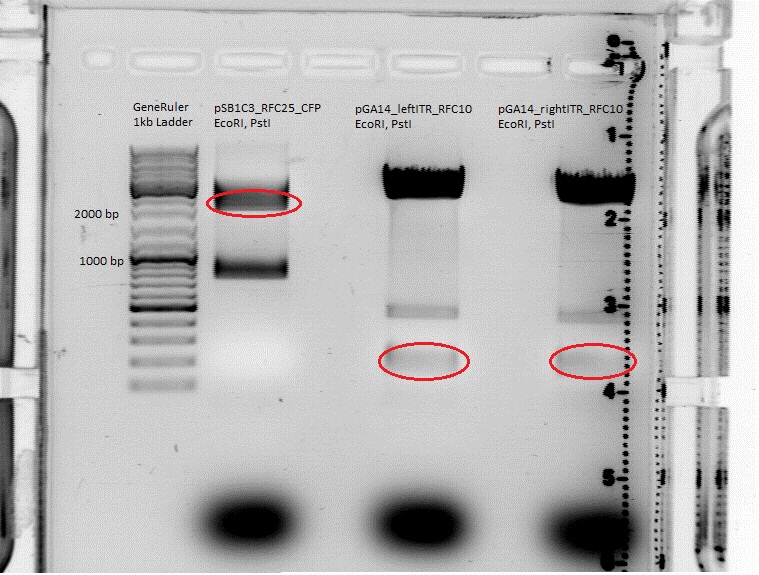
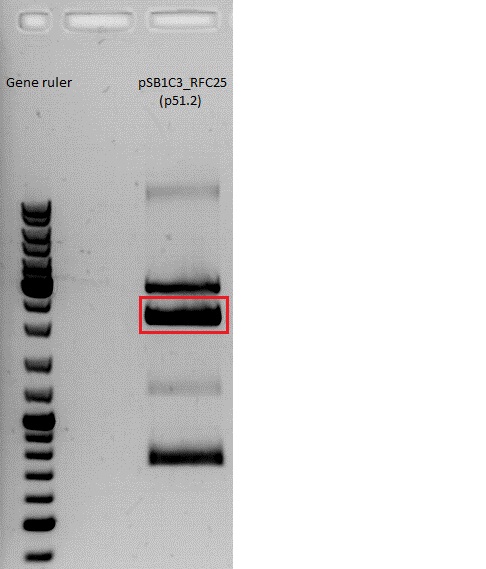
| + | *Gelextraction of pSB1C3_RFC25_CFP: | ||
| + | 0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time:50 minutes <br /> | ||
| + | 4µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas) | ||
| + | <br /> | ||
| + | <br /> | ||
| + | <br /> | ||
| + | <br /> | ||
| + | [[File:Freiburg10 pSB1C3_2x.jpg|500px|]] | ||
| + | |||
| + | |||
| + | <br /> | ||
| + | |||
| + | |||
| + | c(pSB1C3)= 19,5 ng/µl <br /> | ||
| + | |||
| + | |||
| + | <br /> | ||
| + | |||
| + | *T4 ligation of PCR products and vector: | ||
| + | |||
| + | For the Ligation 1µl buffer (10x) and 1µl T4 ligase were used. | ||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | ''' ''' || align="right" |'''vector /µl''' || align="right" |'''insert /µl''' | ||
| + | |- | ||
| + | | pSB1C3 + Rep40 || align="right" |1,57 || align="right" |6,43 | ||
| + | |- | ||
| + | | pSB1C32 + Rep78 || align="right" |1,71 || align="right" |6,29 | ||
| + | |- | ||
| + | | pSB1C3 + AAP || align="right" |5,56 || align="right" |2,44 | ||
| + | |- | ||
| + | |||
| + | |} | ||
| + | <br /> | ||
| + | |||
| + | *Transformation: | ||
| + | |||
| + | The transformation was done following the standard protocol using XL1b cells. | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#ff00ff;">'''Design and Ordering of Oligos'''</p>==== | ||
| + | |||
| + | '''Investigator: Hanna'''<br> | ||
| + | <br/> | ||
| + | <p style="font-size:15px;color:#ff00ff;">'''SV40 terminator'''</p> | ||
| + | <b>Comment:</b> The CMV promoter is often used in combination with a SV40 terminator. Because we want to use this promoter in the context of the VP2 fusion and VP1 insertion approaches, oligos were designed in order to produce a SV40 terminator BioBrick (from the pEGFP-C1 plasmid). | ||
| + | <br/> | ||
| + | [[File:Freiburg10 SV40Terminator.jpg|450px]] | ||
| + | <br/> | ||
| + | |||
| + | <p style="font-size:15px;color:#ff00ff;">'''pEGFP_backbone'''</p> | ||
| + | <b>Comment:</b> In order to perform the VP2 N-terminal fusion and VP1 insertion approaches, we decided to use pEGFP-C1 as expression plasmid. This plasmid contains an EGFP under the control of the CMV promoter and the SV40 terminator. In order to exchange the EGFP with our targeting constructs a PCR of the CMV_backbone_SV40Terminator should be performed. The referring primers were designed with EcoRI and PstI restriction sites. One should keep in mind that this expression plasmid mustn't digested with NgoMIV and it contains Kanamycin-resistance! | ||
| + | <br/> | ||
| + | [[File:Freiburg10 pEGFP-C1 backbone.jpg|400px]] | ||
| + | |||
| + | ===92. labday 17.08.2010=== | ||
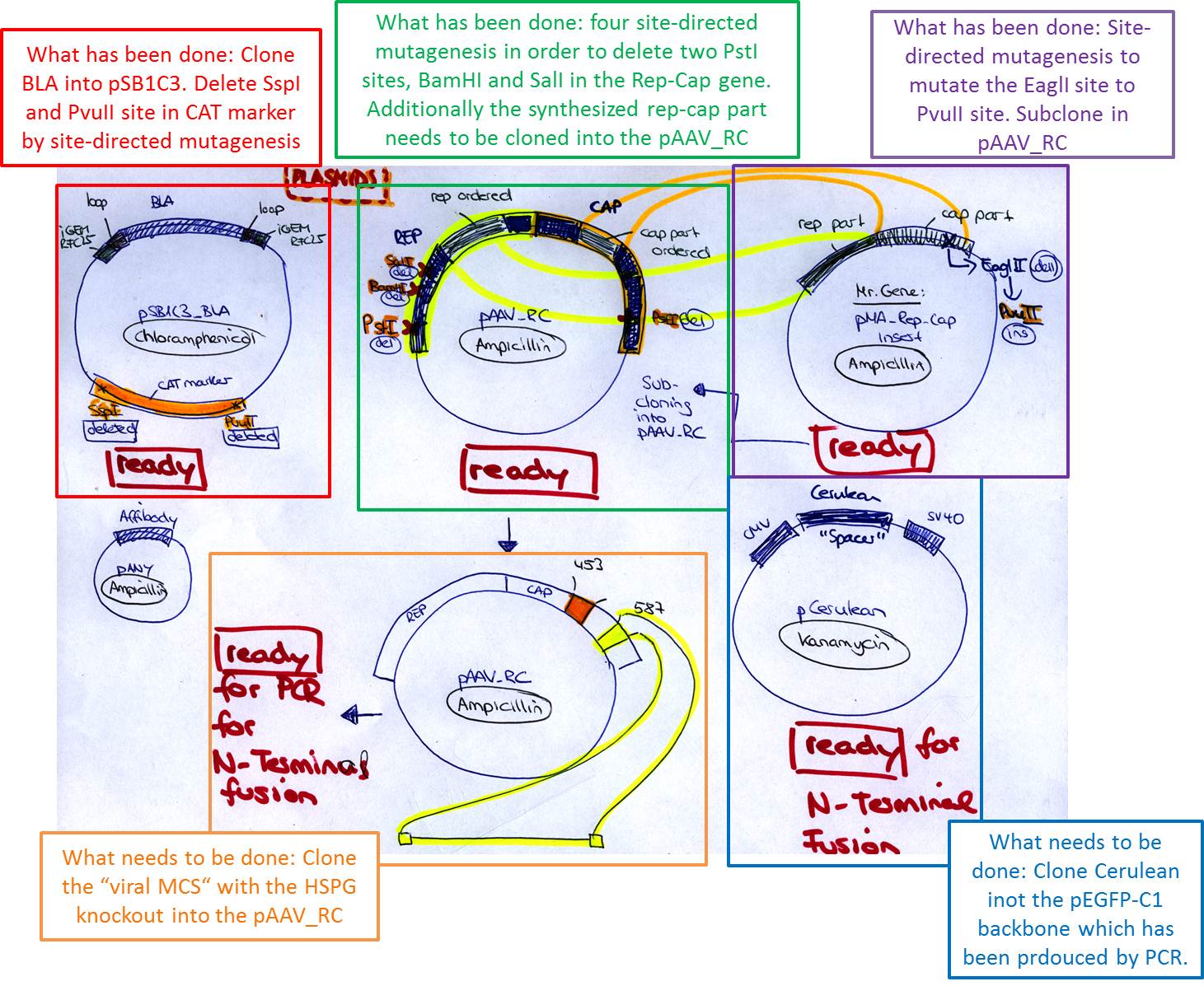
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Overview of the state-of the art plasmid preparation</b></p>==== | ||
| + | <b>Investigator: Bea </b> | ||
| + | <br /> | ||
| + | <p style="font-size:13px; color:#003399;"><b>Comments</b>: In order to obtain a overview (more or less plausible ;-)) I sorted out how the final plasmid need to be constructed. In the picture you can see the most important plasmids which we will need for the retargeting approaches starting tomorrow.</p> | ||
| + | |||
| + | <gallery widths=600px heights=500px caption="Overview scheme"> | ||
| + | Image:Freiburg10 Overview of plasmid preparation for retargeting 16 08 2010.jpg | ||
| + | </gallery> | ||
| + | <br /> | ||
| + | <br /> | ||
| + | [http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/index.php/Laborjournal top of page]<br /> | ||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Further pSB1C3_SDM_SspI_Bla14FM sequencing </p>==== | ||
| + | |||
| + | Investigators: Bea, Patrick | ||
| + | |||
| + | Sequencing results: there is no BLA in clone 4. Clone 5 showed a flawless BLA so it will be used for the next site directed mutagenesis. | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''pSB1C3_SDM_SspI_Bla14FM clone 5 SDM (P222)</p>==== | ||
| + | |||
| + | Investigator: Patrick <br> | ||
| + | Intention: remove PvuII restriction site <br> | ||
| + | |||
| + | PCR program: | ||
| + | *95°C 2 min (1x) | ||
| + | *95°C 20s, 60°C 10 s, 68°C 30 s (30s/kb) (18x) | ||
| + | *68°C 5 minutes | ||
| + | |||
| + | {| border="1" | ||
| + | | '''Ingredients''' || |'''Volume''' | ||
| + | |- | ||
| + | | 10x reaction buffer || |2,5 | ||
| + | |- | ||
| + | | DNA template ( about 10 ng)|| 1 µl diluted P222 | ||
| + | |- | ||
| + | | forward primer: || | 0,58 µl O108 (pSB1C3 PvuII rev) | ||
| + | |- | ||
| + | | reverse primer: || | 0,59 µl O109 (pSB1C3 PvuII for) | ||
| + | |- | ||
| + | | DMSO (2%)|| | 0,5 | ||
| + | |- | ||
| + | | dNTP Mix from the kit|| | 0,5 µl | ||
| + | |- | ||
| + | | QuickSolution Reagent|| |0,75 µl | ||
| + | |- | ||
| + | | Quickchange Lightning Enzyme (1.25U)|| |0,5 µl | ||
| + | |- | ||
| + | |H<sub>2</sub>O|| | 18,08 µl | ||
| + | |- | ||
| + | |Total volume|| |25 µl | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | Further procedure including the transformation was performed according to the standard protocol. | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">Sent for sequencing</p>==== | ||
| + | * pMA_RC (P222) sent for sequencing with O41 (Cap 4000 for) to check if the EaglII restiction site was successfully changed to PvuII. | ||
| + | * pAAV_RC (P158) sent for sequencing with GATC_std_SK, Rep_1250 for (O35), GATC_std_UP-2 to check if all deleted restriction sites are present (PstI, BamHI, SalI). | ||
| + | |||
| + | Investigators: Bea, Patrick | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">Cloning of pMA_RC-Insertparts in pAAV_RC 1.2 SDM SalI</p>==== | ||
| + | '''Investigator: Jessica'''<br> | ||
| + | '''P211''' will be cloned in the '''P158''' vector that contains 4 mutation (BamHI, SalI, PstI 320 + 4073)<br> | ||
| + | <b>Digestion:</b> | ||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''pMA_RC_Insertparts''' || align="right" |'''pAAV_RC 1.2 SDM SalI''' | ||
| + | |- | ||
| + | | DNA || align="right" |5 || align="right" |4 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" |1 || align="right" | 1 | ||
| + | |- | ||
| + | | Buffer 3 (10x)|| align="right" |1 || align="right" |1 | ||
| + | |- | ||
| + | |Enzyme1 BstEII || align="right" |0,5 || align="right" |0,5 | ||
| + | |- | ||
| + | |Enzyme2 SwaI|| align="right" |0,5 || align="right" |0,5 | ||
| + | |- | ||
| + | |H2O|| align="right" |2 || align="right" |3 | ||
| + | |- | ||
| + | |'''Total volume'''|| align="right" |10|| align="right" |10 | ||
| + | |} | ||
| + | <br /> | ||
| + | digestion was splitted in two parts: <ul> | ||
| + | <li>1. digestion with SwaI at 25°C for 1,5h</li> | ||
| + | <li>2. digestion with BstEII at 60°C for 1,5h</li> | ||
| + | </ul> | ||
| + | Preparation of gel:<br /> | ||
| + | 0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at Volt, running time: minutes | ||
| + | <br /> | ||
| + | |||
| + | Expected sizes of constructs: | ||
| + | *pMA_RC-Insertparts: '''493bp''', 3287bp | ||
| + | *pAAV_RC 1.2 SDM SalI: '''6839bp''', 478bp | ||
| + | |||
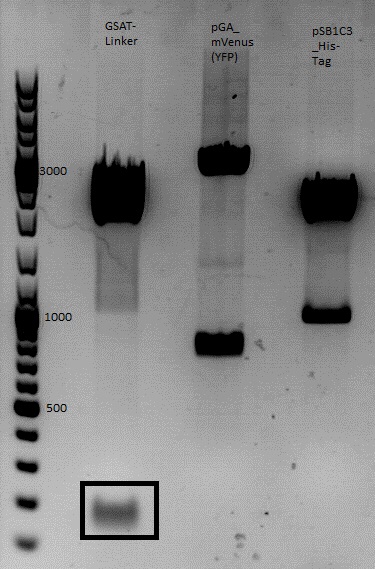
| + | The corresponding bands were cut out and Gel-Extraction was performed according to protocol. | ||
| + | <br /> | ||
| + | [[File:Freiburg1 cloning of pMA_RC-Insertparts in pAAV_RC 1.2 SDM SalI.jpg|400px|]] | ||
| + | <br> | ||
| + | |||
| + | '''concentrations''' measured via NanoDrop: | ||
| + | *pMA_RC-Insertparts: 5,9 ng/µl | ||
| + | *pAAV_RC 1.2 SDM SalI:8,6 ng/µl | ||
| + | <br> | ||
| + | '''Ligation'''<br> | ||
| + | Volume insert: 1,95 µl<br> | ||
| + | Volume vector: 6,08 µl<br> | ||
| + | <br> | ||
| + | '''Trafo''' was prepared with BL21 | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Repetition: Biobrick production of Rep 68'''</p>==== | ||
| + | |||
| + | '''Investigator: Christian L. , Stefan'''<br> | ||
| + | |||
| + | |||
| + | Aim of the experiment: | ||
| + | We want to produce biobricks from Rep 68. | ||
| + | |||
| + | *Plasmid used as template: | ||
| + | Rep_68_ex (p224): c = 350,29 ng/µl <br> | ||
| + | |||
| + | |||
| + | |||
| + | *Primer used: | ||
| + | For Rep_68_ex: Praefix_68_78_ex & Suffix_40_68_ex<br> | ||
| + | <br> | ||
| + | *'''PCR: | ||
| + | (was performed following the standard protocol) | ||
| + | <br> | ||
| + | {| border="1" | ||
| + | | '''Ingredients''' || |'''Volume Rep68 / µl''' | ||
| + | |- | ||
| + | | 5x Phusion HF buffer || |10 | ||
| + | |- | ||
| + | | 10 mM dNTP mix|| |1 | ||
| + | |- | ||
| + | | forward primer: || |2,5 | ||
| + | |- | ||
| + | | reverse primer: || |2,5 | ||
| + | |- | ||
| + | | DNA Template|| |3 | ||
| + | |- | ||
| + | | DMSO (2%)|| | - | ||
| + | |- | ||
| + | | Phusion Polymerase|| |0,5 | ||
| + | |- | ||
| + | |H<sub>2</sub>O|| | 30,5 | ||
| + | |- | ||
| + | |Total volume|| |50 | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | PCR program: | ||
| + | {| border="1" | ||
| + | |'''PCR Program'''|| align="right" |'''temperature/ °C'''|| align="right" |'''Time''' | ||
| + | |- | ||
| + | |1|| align="right" |98 || align="right" |1min | ||
| + | |- | ||
| + | |2|| align="right" |98 || align="right" |15s | ||
| + | |- | ||
| + | |3|| align="right" |63 || align="right" |25s | ||
| + | |- | ||
| + | |4 (step 2-4 8x) || align="right" |72|| align="right" |24s | ||
| + | |- | ||
| + | |5 || align="right" |98|| align="right" | 15s | ||
| + | |- | ||
| + | |6|| align="right" |68|| align="right" |25s | ||
| + | |- | ||
| + | |7 (step 5-7 17x)|| align="right" |72|| align="right" |24s | ||
| + | |- | ||
| + | |8|| align="right" |72|| align="right" |5min | ||
| + | |- | ||
| + | |Hold|| align="right" |4°C | ||
| + | |} | ||
| + | <br> | ||
| + | |||
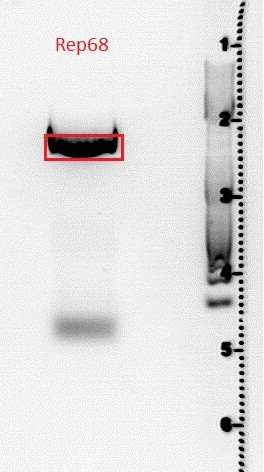
| + | Used agarose gel: | ||
| + | 0,5 g Agarose,50 ml TBE (1%), 3 µl GELRED , at 115 Volt, running time:45 minutes | ||
| + | |||
| + | [[File:Freiburg10 Rep68.jpg|400px|]] | ||
| + | |||
| + | <br /> | ||
| + | Gel extraction was perfomed according to protocol.<br /> | ||
| + | <br /> | ||
| + | comment: Elution was put in 4°C room. Work will be continued tomorrow. | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Transduction of HT1080 cells'''</p>==== | ||
| + | |||
| + | <b>Transduction plan</b> | ||
| + | |||
| + | <b> Investigator: Adrian, Kerstin</b> | ||
| + | |||
| + | <li>1. plate 50.000 cells, YFP | ||
| + | {| align=right | ||
| + | |} | ||
| + | {| border="1" | ||
| + | !A | ||
| + | |control no cells | ||
| + | |300 µl virus (40 µg YFP) | ||
| + | |300 µl virus (20 µg YFP) | ||
| + | |- | ||
| + | !B | ||
| + | |control no virus | ||
| + | |500µl P38 (pAAV2_mVenus) | ||
| + | |1000µl P38 (pAAV2_mVenus) | ||
| + | |- | ||
| + | |} | ||
| + | <li>2. plate 50.000 cells, YFP | ||
| + | {| align=right | ||
| + | |} | ||
| + | {| border="1" | ||
| + | !A | ||
| + | |control no cells | ||
| + | |300 µl virus (40 µg YFP) | ||
| + | |300 µl virus (20 µg YFP) | ||
| + | |- | ||
| + | !B | ||
| + | |control no virus | ||
| + | |500µl P38 (pAAV2_mVenus) | ||
| + | |1000µl P38 (pAAV2_mVenus) | ||
| + | |- | ||
| + | |} | ||
| + | <li>3. plate 100.000 cells, YFP | ||
| + | {| align=right | ||
| + | |} | ||
| + | {| border="1" | ||
| + | !A | ||
| + | |control no cells | ||
| + | |300 µl virus (40 µg YFP) | ||
| + | |300 µl virus (20 µg YFP) | ||
| + | |- | ||
| + | !B | ||
| + | |control no virus | ||
| + | |500µl P38 (pAAV2_mVenus) | ||
| + | |1000µl P38 (pAAV2_mVenus) | ||
| + | |- | ||
| + | |} | ||
| + | <li>4. plate 100.000 cells, YFP | ||
| + | {| align=right | ||
| + | |} | ||
| + | {| border="1" | ||
| + | !A | ||
| + | |control no cells | ||
| + | |300 µl virus (40 µg YFP) | ||
| + | |300 µl virus (20 µg YFP) | ||
| + | |- | ||
| + | !B | ||
| + | |control no virus | ||
| + | |500µl P38 (pAAV2_mVenus) | ||
| + | |1000µl P38 (pAAV2_mVenus) | ||
| + | |- | ||
| + | |} | ||
| + | <li>5. plate 100.000 cells TKGMK | ||
| + | {| align=right | ||
| + | |} | ||
| + | {| border="1" | ||
| + | !A | ||
| + | |control no cells | ||
| + | |150 µl virus (TKGMK clone 1) | ||
| + | |180 µl virus (TKGMK clone 1) | ||
| + | |- | ||
| + | !B | ||
| + | |control no virus | ||
| + | |200 µl (TKGMK clone2) | ||
| + | |200µl (no GOI) | ||
| + | |- | ||
| + | |} | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Midi-Prep of pAAV_igEM_mVenus_YFP and dsAAV_CMV_EGFP</b></p>==== | ||
| + | |||
| + | '''Investigators: Chris, Anna <br> | ||
| + | <p style="font-size:13px; color:#003399;"><b>Comment</b>: Midi-Preps of dsAAV_CMV_EGFP (p31, B24) and pAAV_iGEM_mVenus_YFP (p39, B34)</p> | ||
| + | The Midi-Preps were performed according to the standard protocol yielding the following concentrations: | ||
| + | |||
| + | {| border="1" | ||
| + | | plasmid-no. || align="right" |p31|| align="right" |p39 | ||
| + | |- | ||
| + | | concentration (ng/µl)|| align="right" |539,05 || align="right" |162,37 | ||
| + | |- | ||
| + | |} | ||
| + | <br> | ||
| + | |||
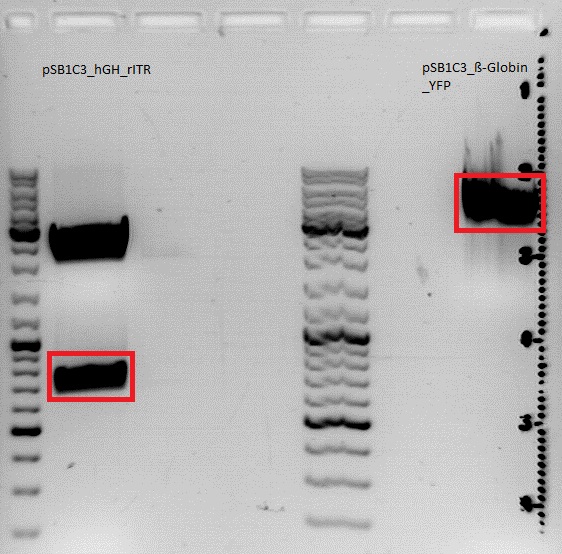
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''BioBrick assembly for pSB1C3_lITR_CMV_beta-globin_YFP_rITR'''</p>==== | ||
| + | investigator:Achim | ||
| + | *This construct without the hGH polyadenylation sequence will be used to determine wether/how much hGH affects translation of the virus gene. | ||
| + | |||
| + | Vector: <br> | ||
| + | *pSB1C3_lITR_CMV_ß_YFP (P209): c=274,86 ng/µl<br> | ||
| + | Insert: <br> | ||
| + | *pSB1C3_rITR (P172/P191): c=73,3 ng/µl<br> | ||
| + | |||
| + | |||
| + | |||
| + | {| border="1" | ||
| + | | components || align="right" |Vector|| align="right" |Insert | ||
| + | |- | ||
| + | | DNA || align="right" |3,6 || align="right" | 20,5 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" |3|| align="right" | 3 | ||
| + | |- | ||
| + | | Buffer 4 (10x)|| align="right" |3 || align="right" | 3 | ||
| + | |- | ||
| + | |Enzyme 1 || align="right" |1(SpeI) || align="right" | 1(XbaI) | ||
| + | |- | ||
| + | |Enzyme 2 || align="right" |1(PstI)|| align="right" | 1 (PstI) | ||
| + | |- | ||
| + | |H<sub>2</sub>O|| align="right" |18,4 || align="right" | 1,5 | ||
| + | |- | ||
| + | |'''Total volume '''|| align="right" |30 || align="right" | 30 | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | * 1% Agarose gel | ||
| + | * 3 µl Gelred | ||
| + | *7 µl DNA-Ladder-Mix | ||
| + | * 115 Volt, running time: 30 minutes | ||
| + | The Insert containing the right ITR was cut out after 30 minutes, the Vector was cut out after 45 minutes<br> | ||
| + | |||
| + | [[File:Freiburg10 Achim 17 8 10.jpg|400px|thumb|left|]]<br> | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <br /> | ||
| + | |||
| + | {| border="1" | ||
| + | | '''Sample/µl ''' ||''' Expected size/bp | ||
| + | |- | ||
| + | | Vector || align="right" |4100 | ||
| + | |- | ||
| + | | Insert || align="right" |150 | ||
| + | |- | ||
| + | |} | ||
| + | <br /> | ||
| + | * weight of insert gel extract: 170 mg | ||
| + | * weight of vector gel extract: 200 mg | ||
| + | |||
| + | |||
| + | '''Nanodrop''' | ||
| + | * insert : 11,4 ng/µl | ||
| + | * vector : 26,7 ng/µl | ||
| + | |||
| + | <br> | ||
| + | |||
| + | '''Ligation''' | ||
| + | *vector: 6,36 µl | ||
| + | *insert: 1,64 µl<br> | ||
| + | |||
| + | '''Transformation''' | ||
| + | <br> | ||
| + | <br> | ||
| + | |||
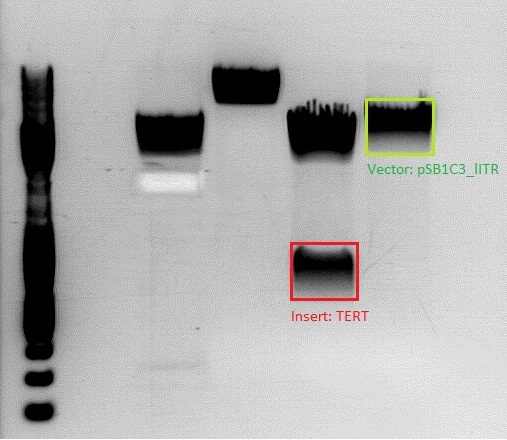
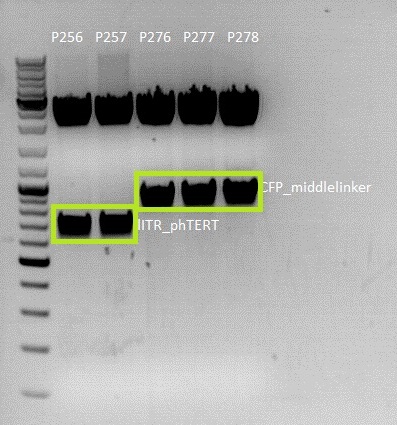
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''BioBrick assembly for pSB1C3_lITR_pTERT'''</p>==== | ||
| + | Investigator:Anissa | ||
| + | *Construct with the pTERT tumor specific promoter | ||
| + | |||
| + | Vector: <br> | ||
| + | *pSB1C3_lITR (P196): 174 ng/µl<br> | ||
| + | Insert: <br> | ||
| + | *pSB1C3_phTERT (P177): c=312 ng/µl<br> | ||
| + | |||
| + | |||
| + | |||
| + | {| border="1" | ||
| + | | components || align="right" |Vector|| align="right" |Insert | ||
| + | |- | ||
| + | | DNA || align="right" |5,75 || align="right" | 6,41 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" |1,5|| align="right" | 1,5 | ||
| + | |- | ||
| + | | Buffer 4 (10x)|| align="right" |1,5 || align="right" | 1,5 | ||
| + | |- | ||
| + | |Enzyme 1 || align="right" |1(SpeI) || align="right" | 1(XbaI) | ||
| + | |- | ||
| + | |Enzyme 2 || align="right" |1(PstI)|| align="right" | 1 (PstI) | ||
| + | |- | ||
| + | |H<sub>2</sub>O|| align="right" |4,25 || align="right" | 3,59 | ||
| + | |- | ||
| + | |'''Total volume '''|| align="right" |15 || align="right" | 15 | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | * 1% Agarose gel | ||
| + | * 3 µl Gelred | ||
| + | *7 µl DNA-Ladder-Mix | ||
| + | * 115 Volt, running time: 45 minutes | ||
| + | |||
| + | |||
| + | [[File:Freiburg10 Achim anissa 17 8.jpg|400px|thumb|left|]]<br> | ||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | |||
| + | <br /> | ||
| + | |||
| + | {| border="1" | ||
| + | | '''Sample/µl ''' ||''' Expected size/bp | ||
| + | |- | ||
| + | | Vector || align="right" |2200 | ||
| + | |- | ||
| + | | Insert || align="right" |500 | ||
| + | |- | ||
| + | |} | ||
| + | <br /> | ||
| + | * weight of insert gel extract: 350 mg | ||
| + | * weight of vector gel extract: 250 mg | ||
| + | |||
| + | |||
| + | '''Nanodrop''' | ||
| + | * insert : 17,7 ng/µl | ||
| + | * vector : 27,6 ng/µl | ||
| + | |||
| + | <br> | ||
| + | |||
| + | '''Ligation''' | ||
| + | *vector: 5,57 µl | ||
| + | *insert: 2,43 µl<br> | ||
| + | |||
| + | '''Transformation''' | ||
| + | <br> | ||
| + | <br/> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbFF;"><b>Picking clones of pSB1C3_lITR_CMV_(betaglobin)_mVenus_hGH_rITR, EGFP_C1 and pMA_T_affibody</b></p>==== | ||
| + | |||
| + | <p><b>Investigator: Anna</b></p> | ||
| + | |||
| + | {| border="1" | ||
| + | | Plasmid || align="right" |Resistance || align="right" |Bacterial strain | ||
| + | |- | ||
| + | | pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR° || align="right" |Chloramphenicol|| align="right" |XL1 blue | ||
| + | |- | ||
| + | | pSB1C3_lITR_CMV_mVEnus_hGH_rITR° || align="right" |Chloramphenicol|| align="right" |XL1 blue | ||
| + | |- | ||
| + | | EGFP_C1|| align="right" |Kanamycin || align="right" | | ||
| + | |- | ||
| + | |pMA_T_affibody || align="right" |Ampicillin|| align="right" | | ||
| + | |- | ||
| + | |} | ||
| + | °: 3 clones each<br> | ||
| + | |||
| + | Clones were inoculated in 10 mL DYT medium containing 10 µL of antibiotics and were put in 37°C room on shaker.<br> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Trafo evaluation of SDM PstI in CD and inoculation'''</p>==== | ||
| + | Investigator: Kira | ||
| + | |||
| + | The plate contained lots of colonies and 2 of them were picked and inoculated into DYT+Amp | ||
| + | |||
| + | ===93. labday 18.08.2010=== | ||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Continuation: pSB1C3_SDM_SspI_Bla14FM clone 5 SDM (P222)</p>==== | ||
| + | There is a lawn on the plate although chloramphenicol was added. Some clones were picked and a new agar plate inoculated to hopefully receive some 'pickable' clones. | ||
| + | <br> | ||
| + | |||
| + | Additionally a new SDM was performed: | ||
| + | PCR program: | ||
| + | *95°C 2 min (1x) | ||
| + | *95°C 20s, 60°C 10 s, 68°C 30 s (30s/kb) (18x) | ||
| + | *68°C 5 minutes | ||
| + | |||
| + | {| border="1" | ||
| + | | '''Ingredients''' || |'''Volume''' | ||
| + | |- | ||
| + | | 10x reaction buffer || |2,5 | ||
| + | |- | ||
| + | | DNA template ( about 10 ng)|| 1 µl diluted P222 | ||
| + | |- | ||
| + | | forward primer: || | 0,58 µl O108 (pSB1C3 PvuII rev) | ||
| + | |- | ||
| + | | reverse primer: || | 0,59 µl O109 (pSB1C3 PvuII for) | ||
| + | |- | ||
| + | | DMSO (2%)|| | 0,5 | ||
| + | |- | ||
| + | | dNTP Mix from the kit|| | 0,5 µl | ||
| + | |- | ||
| + | | QuickSolution Reagent|| |0,75 µl | ||
| + | |- | ||
| + | | Quickchange Lightning Enzyme (1.25U)|| |0,5 µl | ||
| + | |- | ||
| + | |H<sub>2</sub>O|| | 18,08 µl | ||
| + | |- | ||
| + | |Total volume|| |25 µl | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | All further steps including the transformation were performed according to the standard protocol. | ||
| + | This time 2 agar plates were inoculated. One of them with the resuspended pellet and the other one with a 1:100 dilution of the resuspended pellet. | ||
| + | |||
| + | Investigator: Patrick | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Mini Preps pEGFP_C1,pSB1C3_Rep40,pSB1C3_Rep78,pMA_T_Affibody,pSB1C3_SspIdel_BLA clone 2+3,pSB1C3_AAP, pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR,pSB1C3_lITR_CMV_mVenus_hGH_rITR,pSK_CD_SDMPstI'''</p>==== | ||
| + | |||
| + | '''Investigator: Jessica'''<br> | ||
| + | Preps and test digestions were made:<br> | ||
| + | '''pSK_CD_SDMPstI''' | ||
| + | <ul> | ||
| + | <li>Given Plasmid-Number: P248; DNA concentration: 302,7 ng/µL ;</li> | ||
| + | <li>Given Plasmid-Number: P249; DNA concentration: 267,0 ng/µL ;</li> | ||
| + | </ul> | ||
| + | |||
| + | <ul><li>'''P248''', '''P249''' and '''P223''' as control digested with PstI --> no satisfying result,control was not cut but samples two times... sent for sequencing with primer: GATC_std_T7minus1 </li></ul> | ||
| + | <br> | ||
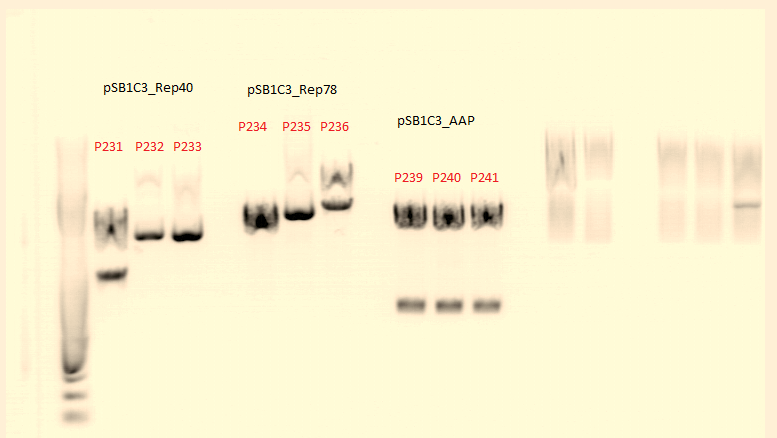
| + | '''pSB1C3_Rep40''' | ||
| + | <ul> | ||
| + | <li>Given Plasmid-Number: P231 (clone3); DNA concentration: 234,00 ng/µL</li> | ||
| + | <li>Given Plasmid-Number: P232 (clone2); DNA concentration: 234,96 ng/µL</li> | ||
| + | <li>Given Plasmid-Number: P233 (clone1); DNA concentration: 237,12 ng/µL</li> | ||
| + | </ul> | ||
| + | '''pSB1C3_Rep78''' | ||
| + | <ul> | ||
| + | <li>Given Plasmid-Number: P234 (clone2); DNA concentration: 277,56 ng/µL</li> | ||
| + | <li>Given Plasmid-Number: P235 (clone1); DNA concentration: 294,19 ng/µL</li> | ||
| + | <li>Given Plasmid-Number: P236 (clone3); DNA concentration: 378,87 ng/µL</li> | ||
| + | </ul> | ||
| + | '''pSB1C3_AAP''' | ||
| + | <ul> | ||
| + | <li>Given Plasmid-Number: P239 (clone1); DNA concentration: 219,82 ng/µL</li> | ||
| + | <li>Given Plasmid-Number: P240 (clone2); DNA concentration: 220,45 ng/µL</li> | ||
| + | <li>Given Plasmid-Number: P241 (clone3); DNA concentration: 209,57 ng/µL</li> | ||
| + | </ul> | ||
| + | '''has to be edited''' | ||
| + | [[File:Freiburg10_Kopfkratzen.gif]] | ||
| + | |||
| + | @Jessica: What are these plasmid numbers?? (Bea) | ||
| + | |||
| + | <ul> | ||
| + | <li>Given Plasmid-Number: P230; DNA concentration: 367,36 ng/µL ;</li> | ||
| + | <li>Given Plasmid-Number: P242; DNA concentration: 202,09 ng/µL ;</li> | ||
| + | <li>Given Plasmid-Number: P243; DNA concentration: 369,34 ng/µL ;</li> | ||
| + | <li>Given Plasmid-Number: P245; DNA concentration: 207,12 ng/µL ;</li> | ||
| + | <li>Given Plasmid-Number: P246; DNA concentration: 198,35 ng/µL ;</li> | ||
| + | <li>Given Plasmid-Number: P247; DNA concentration: 212,22 ng/µL ;</li> | ||
| + | </ul> | ||
| + | |||
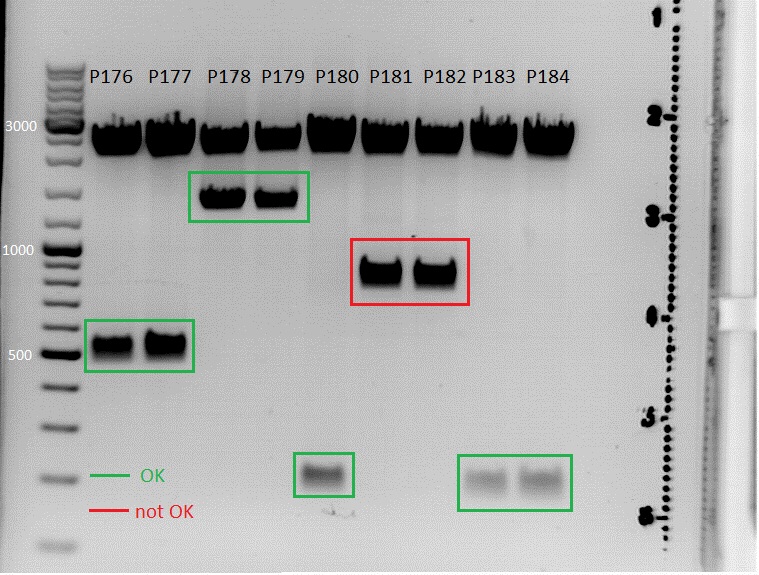
| + | <p>Test digestion</p> | ||
| + | <ul> | ||
| + | <li>buffer used: 4 ; Restriction-enzymes used: Enzyme XbaI ; Enzyme SpeI </li> | ||
| + | <li>Plasmid | ||
| + | <br /> | ||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | align="left" | '''Components''' ||align="left"| '''P231-233 Volume/µL''' ||align="left"| '''P234 Volume/µL''' ||align="left"| '''P235 Volume/µL''' ||align="left"| '''236 Volume/µL''' ||align="left"| '''P239-242 Volume/µL''' | ||
| + | |- | ||
| + | | align="left" | DNA ||align="left"| 4 ||align="left"| 3.5 ||align="left"| 3 ||align="left"| 2.5 ||align="left"| 4 | ||
| + | |- | ||
| + | | align="left" | BSA (10x) ||align="left"| 2 ||align="left"| 2 ||align="left"| 2 ||align="left"| 2 ||align="left"| 2 | ||
| + | |- | ||
| + | | align="left" | Buffer 4 (10x) ||align="left"| 2 ||align="left"| 2 ||align="left"| 2 ||align="left"| 2 ||align="left"| 2 | ||
| + | |- | ||
| + | | align="left" | SpeI ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 | ||
| + | |- | ||
| + | | align="left" | XbaI ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 | ||
| + | |- | ||
| + | | align="left" | H<sub>2</sub>O ||align="left"| 11 ||align="left"| 11,5 ||align="left"| 12 ||align="left"| 12.5 ||align="left"| 11 | ||
| + | |- | ||
| + | | align="left" | <b>Total volume</b> ||align="left"| <b>20</b> ||align="left"| <b>20</b> ||align="left"| <b>20</b> ||align="left"| <b>20</b> ||align="left"| <b>20</b> | ||
| + | |} | ||
| + | <br /> | ||
| + | |||
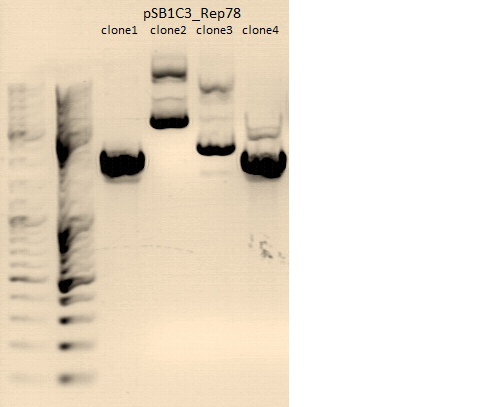
| + | <p style="font-size:13px; color:#003399;"><b>Comment</b>: Test digestion of P231-233 and P239-242 contained equal volumes of ingredients. Therefore the charts were simplyfied. </p> | ||
| + | |||
| + | [[File:Freiburg10 test digestion Rep40,78,AAP.png|500px]] | ||
| + | |||
| + | <p style="font-size:13px; color:#003399;"><b>Comment</b>: P231 and P239 will be sent for sequenzing tomorrow. We picked for additional clones of Rep78 for Mini-Prep and test digestion tomorrow.</p> | ||
| + | </ul> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Continuation of Repetition: Biobrick production of Rep 68 (17.08.2010)'''</p>==== | ||
| + | |||
| + | '''Investigator: Christian L. , Stefan'''<br> | ||
| + | |||
| + | |||
| + | Aim of the experiment: | ||
| + | We want to produce biobricks from Rep 68. | ||
| + | |||
| + | *Vector: name: pSB1C3_CFP <b>P51.1</b> | ||
| + | *Insert: name: Rep68_SDM_PstI clone 1 <b>P224</b> | ||
| + | |||
| + | *buffer used: 4 | ||
| + | *Restriction-enzymes used: | ||
| + | **XbaI | ||
| + | **SpeI | ||
| + | *DNA concentration (P51.1): 132,1 ng/µl | ||
| + | *DNA concentration (P224): 350.29 ng/µl | ||
| + | |||
| + | <br /> | ||
| + | |||
| + | <b>Digestion of plasmid backbone and PCR product:</b> <br /> | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''volume of pSB1C3_CFP /µl''' || align="right" |'''volume of Rep68_SDM_PstI clone 1 /µl''' | ||
| + | |- | ||
| + | | DNA || align="right" |10 || align="right" |29 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" |2 || align="right" | 4 | ||
| + | |- | ||
| + | | Buffer 4 (10x)|| align="right" |2 || align="right" | 4 | ||
| + | |- | ||
| + | |SpeI || align="right" |1 || align="right" |1 | ||
| + | |- | ||
| + | |XbaI || align="right" |1 || align="right" |1 | ||
| + | |- | ||
| + | |H2O|| align="right" |4 || align="right" |1 | ||
| + | |- | ||
| + | |'''Total volume'''|| align="right" | 20|| align="right" |40 | ||
| + | |} | ||
| + | <br> | ||
| + | Incubation at 37°C, 2 h | ||
| + | <br> | ||
| + | <b>Dephoshorylation of Vector pSB1C3_CFP after digestion:</b> | ||
| + | *add 2 µl of 10x Antarctic Phopshatase Reaction Buffer to DNA cut with XbaI and SpeI | ||
| + | *add 1 µl of Antarctic Phosphatase and mix | ||
| + | *incubate for 60 minutes at 37°C<br> | ||
| + | <br /> | ||
| + | <b>Gelextration of pSB1C3</b> | ||
| + | <br /> | ||
| + | Gelextration of pSB1C3 was performed according to standard protocol | ||
| + | <br /> | ||
| + | 0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time:45 <br /> | ||
| + | |||
| + | [[File:Freiburg10 pSB1C3 dephos.png|400px]] | ||
| + | |||
| + | <br /> | ||
| + | <b>PCR Purification</b> | ||
| + | Purification of Rep68 was performed according to the standard protocol | ||
| + | <br /> | ||
| + | Concentration measured: 16,4 ng/µl | ||
| + | <br /> | ||
| + | <b>Ligation</b> | ||
| + | For the ligation 1µl T4 buffer (10x) and 1µl T4 ligase were used. | ||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | ''' ''' || align="right" |'''vector /µl''' || align="right" |'''insert /µl''' | ||
| + | |- | ||
| + | | pSB1C3 + Rep68 || align="right" | 3,9 || align="right" | 4,1 | ||
| + | |- | ||
| + | |} | ||
| + | <br> | ||
| + | Incubation at room temperature for 45 minutes. | ||
| + | <br /> | ||
| + | <b>Transformation:</b> | ||
| + | <br /> | ||
| + | The transformation was done following the standard protocol using XL1b cells. | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Transfection of AAV293 cells'''</p>==== | ||
| + | |||
| + | <b> Investigator:</b> Adrian, Kerstin | ||
| + | |||
| + | * 2x10<sup>6</sup> cells were seeded (instead of 3x10<sup>6</sup>) --> hopefully better virus-production | ||
| + | * used plasmids: | ||
| + | **P50 c= 429 ng/µl (RC) | ||
| + | **P64 c= 500 ng/µl (pHelper) | ||
| + | **P229 c= 162 ng/µl (GOI=YFP) | ||
| + | **P228 c= 540 ng/µl (GOI=eGFP) | ||
| + | **P158a c= 1800ng/µl (RC mutant) | ||
| + | **P158a c= 1770ng/µl (RC mutant) | ||
| + | |||
| + | |||
| + | * <b> 1. plate:</b> Lipo-Transfection | ||
| + | * <b> 2. plate:</b> Lipo-Transfection | ||
| + | |||
| + | |||
| + | * <b> 3. plate:</b> 3,3 µg (RC), 3,3 µg (pHelper), 3,3 µg (YFP) | ||
| + | * <b> 4. plate:</b> 10 µg (RC), 10 µg (pHelper), 3,3 µg (YFP) | ||
| + | * <b> 5. plate:</b> 10 µg (RC mutant P158a), 10 µg (pHelper), 3,3 µg (YFP) | ||
| + | * <b> 6. plate:</b> 10 µg (RC mutant P158b), 10 µg (pHelper), 3,3 µg (YFP) | ||
| + | * <b> 7. plate:</b> 3,3 µg (RC mutant P158a), 3,3 µg (pHelper), 3,3 µg (YFP) | ||
| + | * <b> 8. plate:</b> 3,3 µg (RC mutant P158b), 3,3 µg (pHelper), 3,3 µg (YFP) | ||
| + | * <b> 9. plate:</b> 10 µg (RC), 10 µg (pHelper), 3,3 µg (eGFP) | ||
| + | * <b> 10. plate:</b> 10 µg (RC), 10 µg (pHelper), 10 µg (eGFP) | ||
| + | * <b> 11. plate:</b> 10 µg (RC), 10 µg (pHelper), 20 µg (eGFP) | ||
| + | * <b> 12. plate:</b> 10 µg (RC), 10 µg (pHelper), 40 µg (eGFP) | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Seeding Ht1080'''</p>==== | ||
| + | |||
| + | Investigators: Adrian, Kerstin | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Cloning of pMA_T_affibody and pSB1C3_CFP'''</p>==== | ||
| + | |||
| + | <p style="font-size:13px; color:#003399;"><b>Comment</b>: pMA_T_affibody (p237) from Retrafo and pSB1C3_CFP (p51.5) were digested with EcoRI and PstI.</p> | ||
| + | |||
| + | <b> Investigator: Anna</b> | ||
| + | |||
| + | Vector: <br> | ||
| + | *pSB1C3_CFP (p51.1): c= 132,1 ng/µl<br> | ||
| + | Insert: <br> | ||
| + | *pMA_T_affibody (p237): c= 233,96 ng/µl<br> | ||
| + | |||
| + | |||
| + | |||
| + | {| border="1" | ||
| + | | components || align="right" |Vector p51.1|| align="right" |Insert p237 | ||
| + | |- | ||
| + | | DNA || align="right" |9 || align="right" | 8 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" |2|| align="right" | 2 | ||
| + | |- | ||
| + | | Buffer 4 (10x)|| align="right" |2 || align="right" | 2 | ||
| + | |- | ||
| + | |Enzyme EcoRI || align="right" |1 || align="right" | 1 | ||
| + | |- | ||
| + | |Enzyme PstI || align="right" |1|| align="right" | 1 | ||
| + | |- | ||
| + | |H<sub>2</sub>O|| align="right" |5 || align="right" | 6 | ||
| + | |- | ||
| + | |'''Total volume '''|| align="right" |20 || align="right" | 20 | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | *Gelextraction of pSB1C3_RFC25_CFP: | ||
| + | 0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 120 Volt, running time:55 <br /> | ||
| + | 4µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas) | ||
| + | <br /> | ||
| + | <br /> | ||
| + | <br /> | ||
| + | <br /> | ||
| + | |||
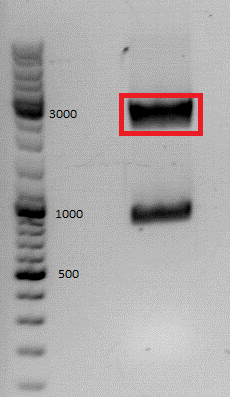
| + | [[file:Freiburg10_pSB1C3_affibody.jpg]] | ||
| + | <br /> | ||
| + | |||
| + | |||
| + | *c(pSB1C3)= 5,4 ng/µl <br /> | ||
| + | *c(affibody)= 5,6 ng/µl | ||
| + | |||
| + | <br /> | ||
| + | |||
| + | *Ligation of PCR products and vector: | ||
| + | |||
| + | For the Ligation 1µl T4 buffer (2x) and 1µl T4 ligase were used. Incubation time: 30 min | ||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | ''' ''' || align="right" |'''vector /µl''' || align="right" |'''insert /µl''' | ||
| + | |- | ||
| + | | pSB1C3 + affibody || align="right" |5,91 || align="right" |2,09 | ||
| + | |- | ||
| + | |} | ||
| + | <br /> | ||
| + | |||
| + | *Transformation: | ||
| + | |||
| + | The transformation was done following the standard protocol using XL1 blue cells. | ||
| + | |||
| + | <p style="font-size:13px; color:#003399;"><b>Result</b>: Clones of pSB1C3_affibody will be picked tomorrow.</p> | ||
| + | |||
| + | ===94. labday 19.08.2010=== | ||
| + | ====<p style="font-size:15px; background-color:#66bbFF;"><b>Continuation: pSB1C3_SDM_SspI_Bla14FM clone 5 SDM (P222)</b></p>==== | ||
| + | Again there was a bacterial lawn on the agar plate. Therefore some clones from the overgrown agar plated were picked and diluted. The 1:10^4, 1:10^5, 1:10^6 dilution each were put onto an agar plate (37°C, over night). Additionally a transformation with XL1B cells from the remaining SDM DNA sample was performed and diluted, too. The 1:10^4 and the 1:10^6 dilution were plated. Hopefully there will be some 'pickable' clones tomorrow. | ||
| + | |||
| + | Investigator: Patrick | ||
| + | |||
| + | [[image:Freiburg10_Kopfkratzen.gif]] | ||
| + | |||
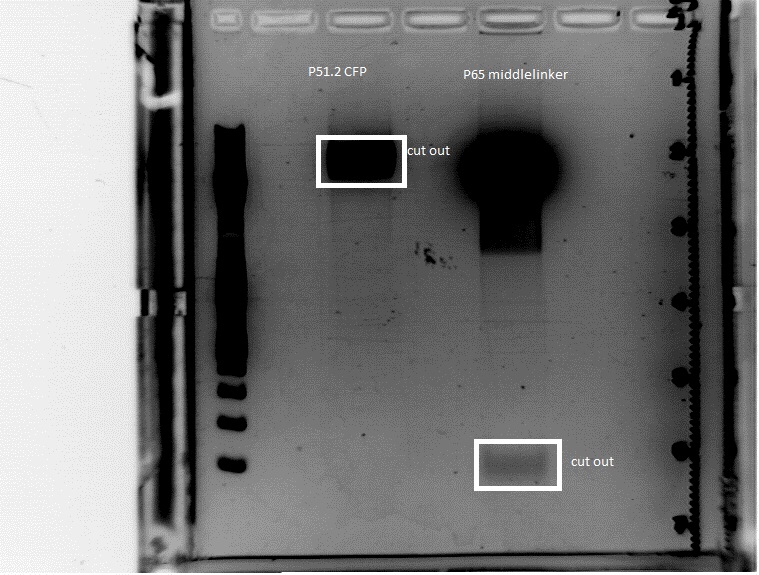
| + | ====<p style="font-size:15px; background-color:#66bbFF;"><b>Cloning of pSB1C3_CFP and middlelinker </b></p>==== | ||
| + | '''Investigator: Jessica'''<br> | ||
| + | *buffer used: 4; Restriction-enzymes used: Enzyme 1: AgeI ; Enzyme 2: PstI ; Enzyme3: NgoMIV | ||
| + | <ul><li>Plasmids:</li> | ||
| + | <ul><li>pSB1C3_CFP '''P51.2'''</li> | ||
| + | <li>pGA14_middle linker '''P65'''</li> | ||
| + | </ul> | ||
| + | </ul> | ||
| + | |||
| + | |||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | align="left" | '''Components''' ||align="left"| '''Mastermix''' ||align="left"| '''P51.2/µL''' ||align="left"| '''P65/µL''' | ||
| + | |- | ||
| + | | align="left" | DNA ||align="left"| -||align="left"| 9,9||align="left"| 10,8 | ||
| + | |- | ||
| + | | align="left" | BSA (10x) ||align="left"|-||align="left"|2||align="left"| 2 | ||
| + | |- | ||
| + | | align="left" | Buffer 4 (10x) ||align="left"| -||align="left"| 2||align="left"| 2 | ||
| + | |- | ||
| + | | align="left" | Enzyme 1 AgeI ||align="left"|-||align="left"| 1||align="left"| - | ||
| + | |- | ||
| + | | align="left" | Enzyme 2 PstI ||align="left"| -||align="left"| 1||align="left"| 1 | ||
| + | |- | ||
| + | | align="left" | Enzyme 3 NgoMIV ||align="left"| -||align="left"| -||align="left"| 1 | ||
| + | |- | ||
| + | | align="left" | H<sub>2</sub>O ||align="left"| -||align="left"| 4,1||align="left"| 3,2 | ||
| + | |- | ||
| + | | align="left" | '''Total volume''' ||align="left"| -||align="left"| 20||align="left"| 20 | ||
| + | |} | ||
| + | <br /> | ||
| + | |||
| + | *Incubation: 90 minutes<br> | ||
| + | '''Agarosegel''' | ||
| + | <br /> | ||
| + | 0.5 g Agarose, 50 ml TAE (1 %), 3 µL GELRED, at 115 Volt, running time: 45 minutes | ||
| + | <br /> | ||
| + | <br /> | ||
| + | |||
| + | |||
| + | <br /> | ||
| + | [[File:Freiburg10 gel cut n terminus.jpg|500px|left|thumb|]] | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | '''concentrations''' measured via NanoDrop: | ||
| + | *SB1C3_CFP: 5,0 ng/µl | ||
| + | *middle linker:1,7 ng/µl | ||
| + | <br> | ||
| + | '''T4 Ligation'''<br> | ||
| + | Volume insert: 1,24 µl<br> | ||
| + | Volume vector: 6,76 µl<br> | ||
| + | <br> | ||
| + | '''Trafo''' was prepared with XL1blue | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Test digestion of pSB1C3_Rep78'''</p>==== | ||
| + | |||
| + | '''Investigator: Chris L., Stefan'''<br> | ||
| + | Preps and test digestions were made:<br> | ||
| + | '''pSB1C3_Rep78''' | ||
| + | <ul> | ||
| + | <li>Given Plasmid-Number: P252 (clone4); DNA concentration: 352,47 ng/µL</li> | ||
| + | <li>Given Plasmid-Number: P253 (clone5); DNA concentration: 408,23 ng/µL</li> | ||
| + | <li>Given Plasmid-Number: P254 (clone6); DNA concentration: 349,18 ng/µL</li> | ||
| + | <li>Given Plasmid-Number: P255 (clone7); DNA concentration: 320,25 ng/µL</li> | ||
| + | </ul> | ||
| + | |||
| + | <p>Test digestion</p> | ||
| + | <ul> | ||
| + | <li>buffer used: 4 ; Restriction-enzymes used: Enzyme XbaI ; Enzyme SpeI </li> | ||
| + | <li>Plasmid | ||
| + | <br /> | ||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | align="left" | '''Components''' ||align="left"| '''P252 Volume/µL''' ||align="left"| '''P253 Volume/µL''' ||align="left"| '''P254 Volume/µL''' ||align="left"| '''255 Volume/µL''' | ||
| + | |- | ||
| + | | align="left" | DNA ||align="left"| 2.5 ||align="left"| 2 ||align="left"| 2.5 ||align="left"| 3 | ||
| + | |- | ||
| + | | align="left" | BSA (10x) ||align="left"| 2 ||align="left"| 2 ||align="left"| 2 ||align="left"| 2 | ||
| + | |- | ||
| + | | align="left" | Buffer 4 (10x) ||align="left"| 2 ||align="left"| 2 ||align="left"| 2 ||align="left"| 2 | ||
| + | |- | ||
| + | | align="left" | SpeI ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 | ||
| + | |- | ||
| + | | align="left" | XbaI ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 | ||
| + | |- | ||
| + | | align="left" | H<sub>2</sub>O ||align="left"| 12.5 ||align="left"| 13 ||align="left"| 12.5 ||align="left"| 12 | ||
| + | |- | ||
| + | | align="left" | <b>Total volume</b> ||align="left"| <b>20</b> ||align="left"| <b>20</b> ||align="left"| <b>20</b> ||align="left"| <b>20</b> | ||
| + | |} | ||
| + | <br /> | ||
| + | Incubation time: 1 h; Incubation temperature: 37° | ||
| + | <br /> | ||
| + | 0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , running time:5 minutes at 90 Volt and 40 minutes at 115 Volt <br /> | ||
| + | 4µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas) | ||
| + | |||
| + | [[File:PSB1C3_Rep78_18.08.png|PSB1C3_Rep78_18.08.png]] | ||
| + | </ul> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">Sent for sequencing</p>==== | ||
| + | |||
| + | <p><b>Investigator: Chris L., Stefan</b></p> | ||
| + | |||
| + | pSB1C3_Rep52 clone3 (P202) sent for sequencing with O162 (VF2). | ||
| + | pSB1C3_Rep40 clone3 (P231), pSB1C3_AAP clone1 (P239), pSB1C3_Rep78 clone4 (P252) and pSB1C3_Rep78 clone7 (255) sent for sequencing with O162 (VF2) and O51 (VR2) | ||
| + | pAAV_RC_ins-rep clone1 (P250) and pAAV_RC_ins-rep clone2 (P251) sent for sequencing with O19 (VP1 primer for pKEX reverse) | ||
| + | |||
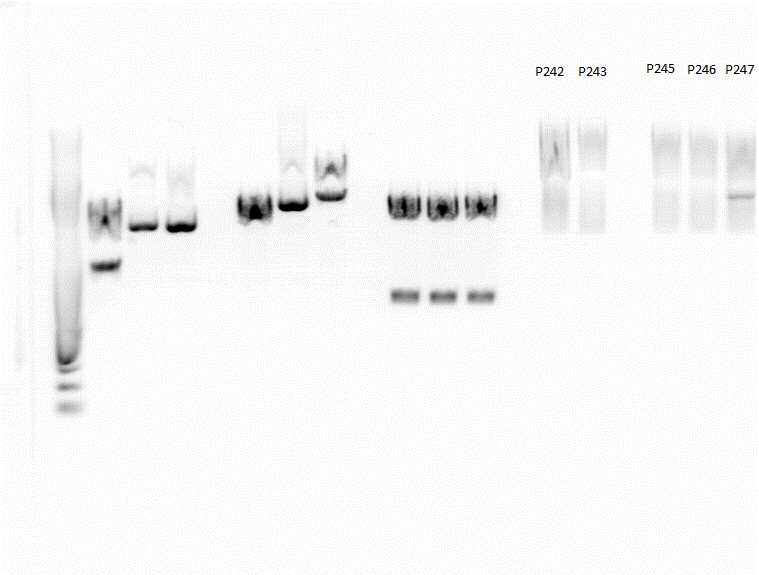
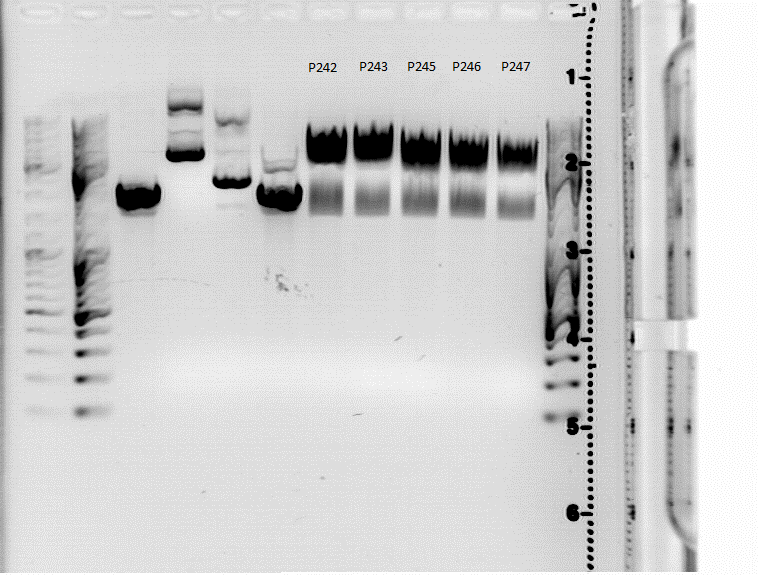
| + | ====<p style="font-size:15px; background-color:#66bbFF;"><b>Test Digestion of P242, P243, P245, P246, P247</b></p>==== | ||
| + | |||
| + | Investigator: Achim | ||
| + | |||
| + | P242: pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR clone1 | ||
| + | |||
| + | P243: pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR clone2 | ||
| + | |||
| + | P245: pSB1C3_lITR_CMV_mVenus_hGH_rITR clone1 | ||
| + | |||
| + | P246: pSB1C3_lITR_CMV_mVenus_hGH_rITR clone2 | ||
| + | |||
| + | P247: pSB1C3_lITR_CMV_mVenus_hGH_rITR clone3 | ||
| + | |||
| + | <gallery widths=450px heights=300px perrow=2 caption=""> | ||
| + | Image:Freiburg10 test digestion 18082010.png | ||
| + | Image:Freiburg10 test digestion 19082010.png | ||
| + | </gallery> | ||
| + | |||
| + | <br><br><br> | ||
| + | |||
| + | (Left picture: Test digestion from 18.8., right picture: Repetition of test digestion from 19.8.) | ||
| + | |||
| + | Fragments sizes: | ||
| + | *P242/243: 1400/3400 | ||
| + | *P245/P246/P247: 1400/2900 | ||
| + | |||
| + | Bands are diffuse and size estimation is difficult because of strange marker behaviour, but at least the second digestion shows two distinct bands which look like they have the correct sizes. P242 and P245 were sent for sequencing. | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Test digestion of pAAV_RC-ins-rep'''</p>==== | ||
| + | |||
| + | '''Investigator: Chris L., Stefan'''<br> | ||
| + | Preps and test digestions were made:<br> | ||
| + | '''pAAV_RC-ins-rep''' | ||
| + | <ul> | ||
| + | <li>Given Plasmid-Number: P250 (clone1); DNA concentration: 467,5 /µL</li> | ||
| + | <li>Given Plasmid-Number: P251 (clone2); DNA concentration: 108,4 ng/µL</li> | ||
| + | </ul> | ||
| + | |||
| + | <p>Test digestion</p> | ||
| + | <ul> | ||
| + | <li>buffer used: 3 ; Restriction-enzymes used: Enzyme SwaI ; Enzyme BstEII </li> | ||
| + | <li>Plasmid | ||
| + | <br /> | ||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | align="left" | '''Components''' ||align="left"| '''P250 Volume/µL''' ||align="left"| '''P251 Volume/µL''' | ||
| + | |- | ||
| + | | align="left" | DNA ||align="left"| 1,7 ||align="left"| 7.3 | ||
| + | |- | ||
| + | | align="left" | BSA (10x) ||align="left"| 1 ||align="left"| 1 | ||
| + | |- | ||
| + | | align="left" | Buffer 3 (10x) ||align="left"| 1 ||align="left"| 1 | ||
| + | |- | ||
| + | | align="left" | SpeI ||align="left"| 0.5 ||align="left"| 0.5 | ||
| + | |- | ||
| + | | align="left" | XbaI ||align="left"| 0.5 ||align="left"| 0.5 | ||
| + | |- | ||
| + | | align="left" | H<sub>2</sub>O ||align="left"| 5.3 ||align="left"| - | ||
| + | |- | ||
| + | | align="left" | <b>Total volume</b> ||align="left"| <b>10</b> ||align="left"| <b>10.3</b> | ||
| + | |} | ||
| + | <br /> | ||
| + | Incubation time with SwaI: 60 minutes; Incubation temperature: 25° | ||
| + | Incubation time with BstEII: 75 minutes; Incubation temperature: 60° | ||
| + | <br /> | ||
| + | 0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , running time: 45 minutes at 115 Volt <br /> | ||
| + | 4µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas) | ||
| + | </ul> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Midiprep of dsAAV_CMV_EGFP and pAAV_iGEM_mVenus_YF '''</p>==== | ||
| + | '''Investigators: Chris W. <br> | ||
| + | <p style="font-size:13px; color:#003399;"><b>Comment</b>: Midi-Preps of dsAAV_CMV_EGFP (p31, B24) and pAAV_iGEM_mVenus_YFP (p39, B34)</p> | ||
| + | The Midi-Preps were performed according to the standard protocol yielding the following concentrations: | ||
| + | |||
| + | {| border="1" | ||
| + | | plasmid-no. || align="right" |p31|| align="right" |p39 | ||
| + | |- | ||
| + | | concentration (ng/µl)|| align="right" |2420,46 & 2586,64 || align="right" |1148,07 & 978,99 | ||
| + | |- | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | |||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Harvest viral particles and Transduction of HT1080 cells '''</p>==== | ||
| + | '''Investigators: Adrian, Kerstin <br> | ||
| + | |||
| + | |||
| + | |||
| + | <li> <b> Transfectionplan </b> (16.08.2010) | ||
| + | |||
| + | |||
| + | {| border="1" | ||
| + | | Plate || align="right" |amount of cells | ||
| + | |- | ||
| + | | 1|| align="right" |100 000 | ||
| + | |- | ||
| + | | 2|| align="right" |200 000 | ||
| + | |- | ||
| + | | 3|| align="right" |400 000 | ||
| + | |- | ||
| + | | 4|| align="right" |500 000 | ||
| + | |- | ||
| + | | 5|| align="right" |800 000 | ||
| + | |- | ||
| + | | 6|| align="right" |1 000 000 | ||
| + | |- | ||
| + | | 7|| align="right" |1 200 000 | ||
| + | |- | ||
| + | | 8|| align="right" |1 500 000 | ||
| + | |- | ||
| + | | 9|| align="right" |1 750 000 | ||
| + | |- | ||
| + | | 10|| align="right" |1 750 000 not sure | ||
| + | |- | ||
| + | |} | ||
| + | The cells were transfected with 40µg YFP, 3,3µg Rep/Cap and 3,3µg pHelper. | ||
| + | |||
| + | <li> <b> Transduction of HT1080 </b> | ||
| + | |||
| + | <p style="font-size:13px; color:#66bbff;"><b>Comments: We want to find out, if the efficiency of virus production can be influenced by the amount of transfected cells. </b></p><br> | ||
| + | |||
| + | <br /> | ||
| + | Plate 1 YFP: 200.000 cells per well<br /> | ||
| + | <table border="3" rules="all" cellpadding="5" cellspacing="1" style="font-size:13pt;text-align:center" > | ||
| + | <tr> | ||
| + | <th width="80"> </th> | ||
| + | <th width="80">1</th> | ||
| + | <th width="80">2</th> | ||
| + | <th width="80">3</th> | ||
| + | |||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>A</td> | ||
| + | <td>control, no cells</td> | ||
| + | <td>165µl virus (1) </td> | ||
| + | <td>165µl virus (2) </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>B</td> | ||
| + | <td>control, no virus</td> | ||
| + | <td> 165µl virus (1) </td> | ||
| + | <td> 165µl virus (2) </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | Plate 2 YFP: 200.000 cells per well<br /> | ||
| + | <table border="3" rules="all" cellpadding="5" cellspacing="1" style="font-size:13pt;text-align:center" > | ||
| + | <tr> | ||
| + | <th width="80"> </th> | ||
| + | <th width="80">1</th> | ||
| + | <th width="80">2</th> | ||
| + | <th width="80">3</th> | ||
| + | |||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>A</td> | ||
| + | <td>control, no cells</td> | ||
| + | <td>165µl virus (3) </td> | ||
| + | <td>165µl virus (4) </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>B</td> | ||
| + | <td>control, no virus</td> | ||
| + | <td> 165µl virus (3) </td> | ||
| + | <td> 165µl virus (4) </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | Plate 3 YFP: 200.000 cells per well<br /> | ||
| + | <table border="3" rules="all" cellpadding="5" cellspacing="1" style="font-size:13pt;text-align:center" > | ||
| + | <tr> | ||
| + | <th width="80"> </th> | ||
| + | <th width="80">1</th> | ||
| + | <th width="80">2</th> | ||
| + | <th width="80">3</th> | ||
| + | |||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>A</td> | ||
| + | <td>control, no cells</td> | ||
| + | <td>165µl virus (5) </td> | ||
| + | <td>165µl virus (6) </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>B</td> | ||
| + | <td>control, no virus</td> | ||
| + | <td> 165µl virus (5) </td> | ||
| + | <td> 165µl virus (6) </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | |||
| + | Plate 4 YFP: 200.000 cells per well<br /> | ||
| + | <table border="3" rules="all" cellpadding="5" cellspacing="1" style="font-size:13pt;text-align:center" > | ||
| + | <tr> | ||
| + | <th width="80"> </th> | ||
| + | <th width="80">1</th> | ||
| + | <th width="80">2</th> | ||
| + | <th width="80">3</th> | ||
| + | |||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>A</td> | ||
| + | <td>control, no cells</td> | ||
| + | <td>165µl virus (7) </td> | ||
| + | <td>165µl virus (8) </td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>B</td> | ||
| + | <td>control, no virus</td> | ||
| + | <td> 165µl virus (7) </td> | ||
| + | <td> 165µl virus (8) </td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | |||
| + | Plate 5 YFP: 200.000 cells per well<br /> | ||
| + | <table border="3" rules="all" cellpadding="5" cellspacing="1" style="font-size:13pt;text-align:center" > | ||
| + | <tr> | ||
| + | <th width="80"> </th> | ||
| + | <th width="80">1</th> | ||
| + | <th width="80">2</th> | ||
| + | <th width="80">3</th> | ||
| + | |||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>A</td> | ||
| + | <td>control, no cells</td> | ||
| + | <td>165µl virus (9) </td> | ||
| + | <td>165µl virus (10)</td> | ||
| + | </tr> | ||
| + | <tr> | ||
| + | <td>B</td> | ||
| + | <td>control, no virus</td> | ||
| + | <td> 165µl virus (9) </td> | ||
| + | <td> 165µl virus (10)</td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | |||
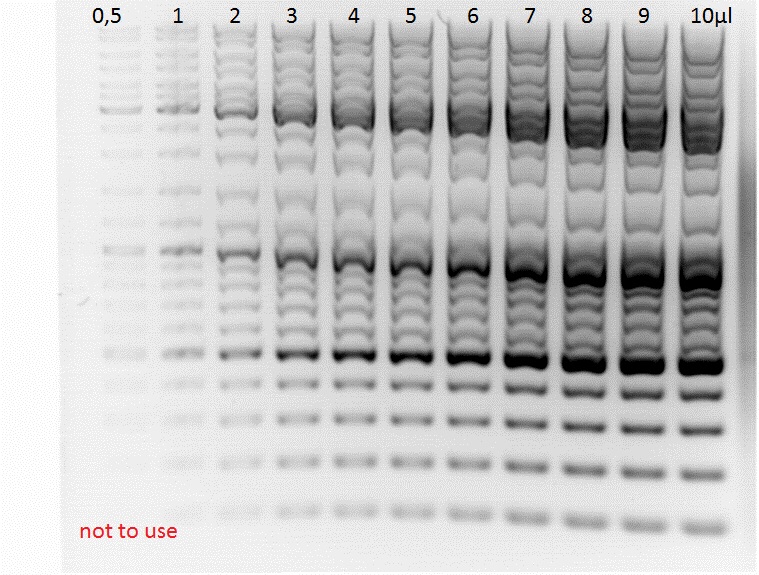
| + | ====<p style="font-size:15px; background-color:#66bbFF;"><b>New ladder mix and testing </b></p>==== | ||
| + | '''Investigator: Jessica'''<br> | ||
| + | new Generuler ladder mix from Fermentas arrived<br> | ||
| + | one tube (containing 100µl solution) was prepared with 100µl loading dye and 400 µl miliporewater<br> | ||
| + | <br> | ||
| + | two test-gels were prepared: | ||
| + | <ul> | ||
| + | <li>1% TAE agarose gel with samples of</li> | ||
| + | <ul> | ||
| + | <li> 0,5 µl </li> | ||
| + | <li> 1,0 µl </li> | ||
| + | <li> 2,0 µl</li> | ||
| + | <li>3,0 µl</li> | ||
| + | <li> 4,0 µl </li> | ||
| + | <li> 5,0 µl </li> | ||
| + | <li> 6,0 µl</li> | ||
| + | <li> 7,0 µl</li> | ||
| + | <li> 8,0 µl</li> | ||
| + | <li> 9,0 µl</li> | ||
| + | <li> 10,0 µl</li> | ||
| + | <li> sample P250 </li> | ||
| + | |||
| + | </ul> | ||
| + | </ul> | ||
| + | [[File:Freiburg10 gel mix.jpg|500px|left|thumb|]] | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <ul> | ||
| + | <li>1% TAE agarose gel containing 20% H2O with samples of ( 20% H2O was an idea I have read in a journal but it doesn't seem like a good idea</li> | ||
| + | <ul> | ||
| + | <li> 0,5 µl </li> | ||
| + | <li> 1,0 µl </li> | ||
| + | <li> 2,0 µl</li> | ||
| + | <li> 3,0 µl</li> | ||
| + | <li> 4,0 µl </li> | ||
| + | <li> 5,0 µl </li> | ||
| + | <li> 6,0 µl</li> | ||
| + | <li> 7,0 µl</li> | ||
| + | <li> 8,0 µl</li> | ||
| + | <li> 9,0 µl</li> | ||
| + | <li> 10,0 µl</li> | ||
| + | <li> sample P251 </li> | ||
| + | </ul> | ||
| + | </ul> | ||
| + | |||
| + | [[File:Freiburg10 gel mix 20% wasser.jpg|500px|left|thumb|]] | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <br> | ||
| + | <p style="font-size:13px; color:#66bbff;"><b>Comments:</b> an example of these laddervolumes is patches at the bench (right up)</p><br> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbFF;"><b>SDM of CD NgoMIV </b></p>==== | ||
| + | '''Investigator:Kira'''<br> | ||
| + | |||
| + | Comment: Lightning kit was used for quickchange reaction | ||
| + | {| border="1" | ||
| + | | '''component''' || '''Volume/µl''' | ||
| + | |- | ||
| + | | DNA: (1:100) || |3,3 | ||
| + | |- | ||
| + | | 10x reaction buffer: || |2,5 | ||
| + | |- | ||
| + | | primer forward NgoMIV (1:10) || |0,55 | ||
| + | |- | ||
| + | | primer reverse NgoMIV (1:10) || |0,55 | ||
| + | |- | ||
| + | | dNTP || |0,5 | ||
| + | |- | ||
| + | |DMSO || |0 | ||
| + | |- | ||
| + | |QuickSolution Reagent ||0,75 | ||
| + | |- | ||
| + | |H2O || |16,35 | ||
| + | |- | ||
| + | |Lightning Polymerase || |0,5 | ||
| + | |} | ||
| + | <br /> | ||
| + | |||
| + | PCR | ||
| + | {| border="1" | ||
| + | | '''segment''' || '''cycles''' || '''temperature''' || '''time''' | ||
| + | |- | ||
| + | | 1 || |1 || |95 || |2min | ||
| + | |- | ||
| + | | 2 || |18 || |95 || |20sec | ||
| + | |- | ||
| + | | || | || |60 || | 10 sec | ||
| + | |- | ||
| + | | || | || |68 || | 2 min15 sec | ||
| + | |- | ||
| + | | 3 || | || | 68|| | 5 min | ||
| + | |} | ||
| + | <br /> | ||
| + | sample was digested with 1 ul DpnI for 10 min at 37°C | ||
| + | |||
| + | Transformation was performed according to the standard protocol. | ||
| + | <br/> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbFF;"><b>Picking clones of pSB1C3_affibody and pSB1C3_Rep68</b></p>==== | ||
| + | |||
| + | <p><b>Investigator: Anna</b></p> | ||
| + | |||
| + | {| border="1" | ||
| + | | Plasmid || align="right" |Resistance || align="right" |Bacterial strain | ||
| + | |- | ||
| + | | pSB1C3_affibody° || align="right" |Chloramphenicol|| align="right" |XL1 blue | ||
| + | |- | ||
| + | | pSB1C3_Rep68°° || align="right" |Chloramphenicol|| align="right" |XL1 blue | ||
| + | |- | ||
| + | |} | ||
| + | °: 2 clones | ||
| + | °°: 3 clones each<br> | ||
| + | |||
| + | Clones were inoculated in 10 mL DYT medium containing 10 µL of antibiotics and were put in 37°C room on shaker.<br> | ||
| + | |||
| + | ===95. labday 20.08.2010=== | ||
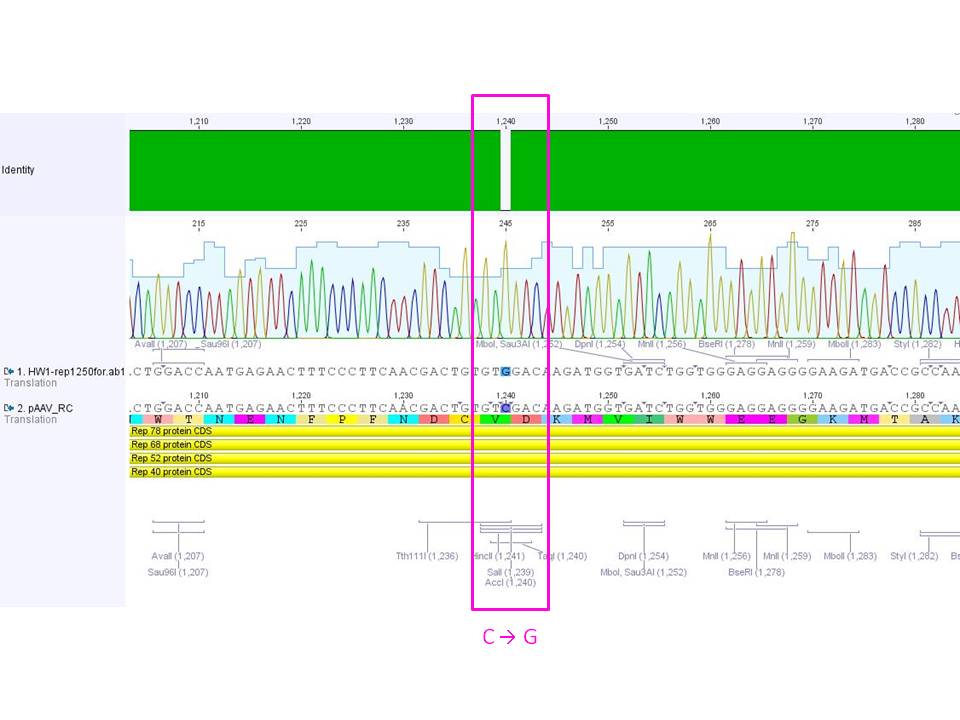
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Sequence analysis of sequencing reads prepared yesterday (19.08.2010)</b></p>==== | ||
| + | <b>Investigator: Bea </b> | ||
| + | [[Image:Mistake.png|thumb|200px]] | ||
| + | <br /> | ||
| + | <p style="font-size:13px; color:#cc0033;"><I><b>GENERAL COMMENT:</b> pSB1C3 {Rep 52 and Rep 78{} were sent for sequencing BUT they cannot be used for subcloning the ordered rep protein because in the produced BioBricks the restriction enzyme needed for subcloning (SwaI) is NOT present in the BioBricks pSB1C3_Rep52 and rep78.</i> </p> | ||
| + | <br /> | ||
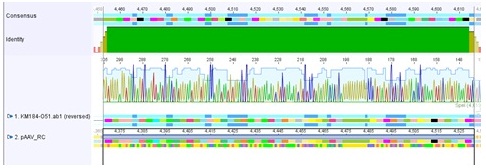
| + | <p style="font-size:13px; color:#003399;"><b>Comments</b>: Sequence analysis of pAAV_RC containing the subcloned "rep" (ordered) </p> | ||
| + | <ul> | ||
| + | <li>Primer used: VP1 primer fro pKex</li> | ||
| + | <li>Plasmid sequenced: P250 </li> | ||
| + | <li>Sequence sample: CL-9 </li> | ||
| + | <li>Stored in folder: </li> | ||
| + | </ul> | ||
| + | |||
| + | <gallery widths=700px heights=300px caption="xy"> | ||
| + | Image: | ||
| + | </gallery> | ||
| + | <br /> | ||
| + | <b>Results:</b> <p style="color:#00bbff;"> </p> | ||
| + | |||
| + | <br /> | ||
| + | <b>Next steps: xy </b> | ||
| + | <br /> | ||
| + | <br /> | ||
| + | |||
| + | <p style="font-size:13px; color:#003399;"><b>Comments</b>: Sequence analysis of pAAV_RC containing the subcloned "rep" (ordered) </p> | ||
| + | <ul> | ||
| + | <li>Primer used: VP1 primer fro pKex</li> | ||
| + | <li>Plasmid sequenced: P251 </li> | ||
| + | <li>Sequence sample: CL-10 </li> | ||
| + | <li>Stored in folder: </li> | ||
| + | </ul> | ||
| + | |||
| + | <gallery widths=700px heights=300px caption="xy"> | ||
| + | Image: | ||
| + | </gallery> | ||
| + | <br /> | ||
| + | <b>Results:</b> <p style="color:#00bbff;"> </p> | ||
| + | |||
| + | <br /> | ||
| + | <b>Next steps: xy </b> | ||
| + | <br /> | ||
| + | <br /> | ||
| + | <p style="font-size:13px; color:#003399;"><b>Comments</b>: Sequence analysis of pSB1C3_Rep40 </p> | ||
| + | <ul> | ||
| + | <li>Primer used: VR-2</li> | ||
| + | <li>Plasmid sequenced: P231 </li> | ||
| + | <li>Sequence sample: CL-2 </li> | ||
| + | <li>Stored in folder: DKFZ Rep40ex</li> | ||
| + | </ul> | ||
| + | <gallery widths=700px heights=300px caption="xy"> | ||
| + | Image: | ||
| + | </gallery> | ||
| + | <br /> | ||
| + | <b>Results:</b> <p style="color:#00bbff;"> Sequence read is good (prefix is ok) but there was found a mutation in Rep52 protein: G2886A (nucleotides) --> V337I. </p> | ||
| + | |||
| + | <br /> | ||
| + | <b>Next steps: xy </b> | ||
| + | <br /> | ||
| + | <br /> | ||
| + | <p style="font-size:13px; color:#003399;"><b>Comments</b>: Sequence analysis of pSB1C3_AAP </p> | ||
| + | <ul> | ||
| + | <li>Primer used: VR-2</li> | ||
| + | <li>Plasmid sequenced: P239 </li> | ||
| + | <li>Sequence sample: CL-3 </li> | ||
| + | <li>Stored in folder: DKFZ AAP</li> | ||
| + | </ul> | ||
| + | <gallery widths=700px heights=300px caption="xy"> | ||
| + | Image: | ||
| + | </gallery> | ||
| + | <br /> | ||
| + | <b>Results:</b> <p style="color:#00bbff;"> xy</p> | ||
| + | |||
| + | <br /> | ||
| + | <b>Next steps: xy </b> | ||
| + | <br /> | ||
| + | <br /> | ||
| + | |||
| + | [http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/index.php/Laborjournal top of page]<br /> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbFF;"><b>Continuation: pSB1C3_SDM_SspI_Bla14FM clone 5 SDM (P222)</b></p>==== | ||
| + | |||
| + | Investigator: Patrick <br> | ||
| + | |||
| + | The XL1B transformation didnt work or was too much diluted because there were no clones on the plates. The dilution steps of the transformed XL10-Gold cells worked quite well. Now there are 'pickable' clones. | ||
| + | <br /> | ||
| + | [[File:Freiburg10_Kopfkratzen.gif]] | ||
| + | <br /> | ||
| + | [http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/index.php/Laborjournal top of page]<br /> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbFF;"><b>Subcloning cap from pMA_RepCap into pAAV_RC which contains four mutations and the synthesized Rep part</b></p>==== | ||
| + | Investigator: Patrick <br> | ||
| + | |||
| + | pAAV_RC_ins-rep clone 1 (P250) and pMA_RepCap Vector_SDM_InsPvuII clone 1 (P211) were digested with PspLI and BspMI according to the standard protocol (Subcloning_cap_into_pAAV_RC). | ||
| + | |||
| + | Expected size of the bands: | ||
| + | *Insert: 920 bp (P211) | ||
| + | *Vector: 6414 bp (P250) | ||
| + | |||
| + | The gelextraction yielded the following DNA concentrations: | ||
| + | * P 211 : 2,45 ng/µl | ||
| + | * P 250 : 11,15 ng/µl | ||
| + | <br> | ||
| + | [[File:Freiburg10_Digestion_of_pMA_RC_and _pAAV_RC_20_08_2010.jpg|400px]] | ||
| + | <br> | ||
| + | The marked bands were cut out and the DNA was extracted. The ligation was performed accodring to the standard prtocol with a total volume of 10 µl: 2,67 µl vector backbone(P250), 5,33 µl Insert following a transformation according to the standard protocol with XL1B cells. | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Mini-Preps of pSB1C3_Rep68_ex and pSB1C3_ZEGFR:1907</b></p>==== | ||
| + | <b>Investigator: Chris L.,Stefan</b><br> | ||
| + | <ul> | ||
| + | <b>Glycerol stocks were prepared:</b> | ||
| + | <li>B218 = pSB1C3_Rep68_ex clone1</li> | ||
| + | <li>B219 = pSB1C3_Rep68_ex clone2</li> | ||
| + | <li>B220 = pSB1C3_Rep68_ex clone3</li> | ||
| + | <br /> | ||
| + | |||
| + | <li>B221 = pSB1C3_ZEGFR:1907 clone 1</li> | ||
| + | <li>B222 = pSB1C3_ZEGFR:1907 clone 2</li> | ||
| + | <br /> | ||
| + | <b>Mini-Prep following the standard protocol</b> | ||
| + | <br /> | ||
| + | <li>B218 = pSB1C3_Rep68_ex clone1 c=286.72 ng/µl</li> | ||
| + | <li>B219 = pSB1C3_Rep68_ex clone2 c=277.06 ng/µl</li> | ||
| + | <li>B220 = pSB1C3_Rep68_ex clone3 c=298.18 ng/µl </li><br /> | ||
| + | <br /> | ||
| + | |||
| + | <li>B221 = pSB1C3_ZEGFR:1907 clone 1 c=174.20 ng/µl | ||
| + | <li>B222 = pSB1C3_ZEGFR:1907 clone 2 c=162.48 ng/µl <br /> | ||
| + | <br /> | ||
| + | </ul> | ||
| + | <b>Test digestion</b><br> | ||
| + | <ul> | ||
| + | <li>buffer used: 4 ; Restriction-enzymes used: XbaI ; SpeI ; EcoRI ; PstI </li> | ||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | align="left" | '''Components''' ||align="left"| '''P264 Volume/µL''' ||align="left"| '''P265 Volume/µL''' ||align="left"| '''P266 Volume/µL''' ||align="left"| '''P267 Volume/µL''' ||align="left"| '''P268 Volume/µL''' | ||
| + | |- | ||
| + | | align="left" | DNA (800 ng) ||align="left"| 2.8 ||align="left"| 2.9 ||align="left"| 2.7 ||align="left"| 4.6 ||align="left"| 4.9 | ||
| + | |- | ||
| + | | align="left" | BSA (10x) ||align="left"| 1 ||align="left"| 1 ||align="left"| 1 ||align="left"| 1 ||align="left"| 1 | ||
| + | |- | ||
| + | | align="left" | Buffer 4 (10x) ||align="left"| 1 ||align="left"| 1 ||align="left"| 1 ||align="left"| 1 ||align="left"| 1 | ||
| + | |- | ||
| + | | align="left" | SpeI ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| - ||align="left"| - | ||
| + | |- | ||
| + | | align="left" | XbaI ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| 0.5 ||align="left"| - ||align="left"| - | ||
| + | |- | ||
| + | | align="left" | PstI ||align="left"| - ||align="left"| - ||align="left"| - ||align="left"| 0.5 ||align="left"| 0.5 | ||
| + | |- | ||
| + | | align="left" | EcoRI ||align="left"| - ||align="left"| - ||align="left"| - ||align="left"| 0.5 ||align="left"| 0.5 | ||
| + | |- | ||
| + | | align="left" | H<sub>2</sub>O ||align="left"| 4.2 ||align="left"| 4.1 ||align="left"| 4.3 ||align="left"| 2.4 ||align="left"| 2.1 | ||
| + | |- | ||
| + | | align="left" | <b>Total volume</b> ||align="left"| <b>10</b> ||align="left"| <b>10</b> ||align="left"| <b>10</b> ||align="left"| <b>10</b> ||align="left"| <b>10</b> | ||
| + | |} | ||
| + | <br /> | ||
| + | Incubation time: 1 h; Incubation temperature: 37° | ||
| + | <br /> | ||
| + | 0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , running time:5 minutes at 90 Volt and 40 minutes at 115 Volt <br /> | ||
| + | 4µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas) | ||
| + | <br /> | ||
| + | <br /> | ||
| + | [[File:Freiburg10 pSB1C3 Rep68 ex and pSB1C3 ZEGFR1907.png|400px]] | ||
| + | <br /> | ||
| + | <br /> | ||
| + | <p style="font-size:13px; color:#003399;"><b>Comment</b>:pSB1C3_Rep68_ex clone 3 (P266) sent for sequencing with O162 (VF2) and O51 (VR2). pSB1C3_ZEGFR:1907 clone 1 (P267) sent for sequencing with O51 (VR2).</p> | ||
| + | </ul> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Hybridisation of the oligos for NLS and cloning NLS into pSB1C3</b></p>==== | ||
| + | <b>Investigator: Anissa </b><br> | ||
| + | <ul> | ||
| + | <li> '''Hybridisation''': program: Origami 1 <br/> | ||
| + | Hybridisation was peformed according the standard protocol </li> | ||
| + | <br/> | ||
| + | <li>Digestion of pSB1C3_CFP | ||
| + | <br/> | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''Vector /µl''' | ||
| + | |- | ||
| + | | DNA || align="right" |6,62 | ||
| + | |- | ||
| + | | BSA (10X) || align="right" |1,5 | ||
| + | |- | ||
| + | | Buffer 4 || align="right" |1,5 | ||
| + | |- | ||
| + | | Enzyme EcoRI|| align="right" |1 | ||
| + | |- | ||
| + | | Enzyme PstI|| align="right" |1 | ||
| + | |- | ||
| + | | H2O|| align="right" |3,38 | ||
| + | |- | ||
| + | | Endvolume|| align="right" |15 | ||
| + | |- | ||
| + | |}</li> | ||
| + | <br/> | ||
| + | <li>Gel:After digestion, the sample was loaded on a 1% agarose-gel.</li> | ||
| + | <br/> | ||
| + | <gallery widths=300px heights=300px caption="Digest pSB1C3_CFP"> | ||
| + | Image:Freiburg10 pSB1C3 NLS.png | ||
| + | </gallery> | ||
| + | |||
| + | |||
| + | <li> Gelextraction: After the gel run, the vector backbone pSB1C3 was cut and purified according to the standard protocol</li> | ||
| + | <br/> | ||
| + | <li>Ligation: the original concentration of the hybridisation was 209,7 ng/µl, for the ligation a 1:10 dilution was made | ||
| + | <br/> | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''Volume /µl''' || align="right" |'''Concentration/ ng/µl''' | ||
| + | |- | ||
| + | | Insert (NLS) || align="right" |6,79|| align="right" | 2,097 | ||
| + | |- | ||
| + | | Vector (pSB1C3_cut) || align="right" |1,21|| align="right" | 8,3 | ||
| + | |- | ||
| + | |} | ||
| + | <br/></li> | ||
| + | <li>Transformation: Trafo into XL1 blue was performed according the standard protocol</li> | ||
| + | </ul> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Seeding HT1080 cells</b></p>==== | ||
| + | <b>Investigator: Kerstin, Adrian </b><br> | ||
| + | |||
| + | * Seeding HT1080 cells for transduction with viral stocks from 18.08. | ||
| + | * We harvest four T75 flask (two of them were used three times (3 passages)) and seed them into eight 6er dishes (200.000 cells per well) | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''PCR and ligation for biobrick-production of pCerulean'''</p>==== | ||
| + | |||
| + | '''Investigator: Christian L. , Stefan'''<br> | ||
| + | <br /> | ||
| + | DNA template: pEGFP_C1 (P230) | ||
| + | C=367,4 ng/µl | ||
| + | <br /> | ||
| + | <br /> | ||
| + | '''PCR:''' | ||
| + | (was performed following the standard protocol) | ||
| + | <br> | ||
| + | {| border="1" | ||
| + | | '''Ingredients''' || align="right" |'''Volume / µl''' | ||
| + | |- | ||
| + | | 5X Phusion HF buffer || align="right" |10 | ||
| + | |- | ||
| + | | 10 mM dNTP mix|| align="right" |1 | ||
| + | |- | ||
| + | | forward primer: O163 (pEGFP-C1_rev) || align="right" |2.5 | ||
| + | |- | ||
| + | | reverse primer: O164 (pEGFP-C1_for) || align="right" |2.5 | ||
| + | |- | ||
| + | | DNA Template(1:1000) || align="right" |3 µl | ||
| + | |- | ||
| + | | DMSO (2%)|| align="right" | 1 | ||
| + | |- | ||
| + | | Phusion Polymerase|| align="right" |0,5 | ||
| + | |- | ||
| + | |H<sub>2</sub>O|| align="right" |29.5 | ||
| + | |- | ||
| + | |Total volume|| align="right" |50 | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | '''PCR program:''' | ||
| + | {| border="1" | ||
| + | |'''PCR Program'''|| align="right" |'''temperature/ °C'''|| align="right" |'''time/ sec.''' | ||
| + | |- | ||
| + | |1|| align="right" |98 || align="right" |60 | ||
| + | |- | ||
| + | |2|| align="right" |98 || align="right" |15 | ||
| + | |- | ||
| + | |8x|| align="right" |59 || align="right" |25 | ||
| + | |- | ||
| + | |3 || align="right" |72 || align="right" |60 | ||
| + | |- | ||
| + | |4 || align="right" |98 || align="right" |15 | ||
| + | |- | ||
| + | |17x|| align="right" |66 || align="right" |25 | ||
| + | |- | ||
| + | |5|| align="right" |72 || align="right" |60 | ||
| + | |- | ||
| + | |1x|| align="right" |72 || align="right" |5 | ||
| + | |- | ||
| + | |Hold|| align="right" |4 || align="right" | - | ||
| + | |} | ||
| + | <br> | ||
| + | <p style="font-size:13px; color:#FF0033;">'''''Comments''''': Errors in table, please correct!!! (TB)</p> <br> | ||
| + | '''Gelextraction of PCR product''' | ||
| + | <br /> | ||
| + | Gelextraction was performed according to the standard protocol of the PCR product. | ||
| + | <br> | ||
| + | [[File:Freiburg10 pEGFP C1.jpg|300px]] | ||
| + | <br> | ||
| + | <p style="font-size:13px; color:#68bbff;">'''''Comment''''': 0,4 g Agarose,50 ml TAE (0,8%), 3 µl GELRED , at 115 Volt, running time: 45 minutes</p> <br /> | ||
| + | '''Digestion of PCR product and vector:''' | ||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''PCR product /µl''' || align="right" |'''vector /µl''' | ||
| + | |- | ||
| + | | DNA || align="right" | 10 || align="right" | 29 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" | 2 || align="right" | 2 | ||
| + | |- | ||
| + | | Buffer 4 (10x)|| align="right" | 2 || align="right" |2 | ||
| + | |- | ||
| + | |Enzyme '''EcoRI || align="right" | 1 || align="right" | 1 | ||
| + | |- | ||
| + | |Enzyme '''PstI || align="right" | 1 || align="right" | 1 | ||
| + | |- | ||
| + | |H2O|| align="right" | 4 || align="right" | 5 | ||
| + | |- | ||
| + | |'''Total volume (e.g. 15,20,25,30 µl)'''|| align="right" | 20 || align="right" | 40 | ||
| + | |} | ||
| + | <br /> | ||
| + | Purification of the PCR product with PCR purification was made according to the standard protocol. | ||
| + | <br /> | ||
| + | '''Gelextraction of CFP''' | ||
| + | <br /> | ||
| + | Gelextraction was performed according to the standard protocol. | ||
| + | <br> | ||
| + | [[File:Freiburg10 pSB1C3 CFP.png|300px]]> | ||
| + | <br /> | ||
| + | <p style="font-size:13px; color:#68bbff;">'''''Comment''''': 0.5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time: 45 minutes</p> <br /> | ||
| + | <br /> | ||
| + | '''Ligation''' | ||
| + | <br /> | ||
| + | The Ligation was performed as following: | ||
| + | * Vector Volume: 4,1 µl | ||
| + | * Insert Volume: 3,9 µl | ||
| + | <br> | ||
| + | * 1µl T4-Ligase buffer (10x) | ||
| + | * 8µl (Vector + Insert) mix | ||
| + | * 1µl T4-Ligase | ||
| + | <br> Incubation for 45 minutes at room temperature | ||
| + | <br> | ||
| + | '''Transformation:''' | ||
| + | <br /> | ||
| + | performed according to the standard protocol (XL1B). The cells were plated on a agar plate with Kanamycin. | ||
| + | <br> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Biobrick production of VP1_up'''</p>==== | ||
| + | |||
| + | '''Investigator: Achim'''<br> | ||
| + | <br /> | ||
| + | DNA template: pKex_VP1 (P27) | ||
| + | C=99,3 ng/µl | ||
| + | <br /> | ||
| + | <br /> | ||
| + | '''PCR:''' | ||
| + | (was performed following the standard protocol) | ||
| + | <br> | ||
| + | {| border="1" | ||
| + | | '''Ingredients''' || align="right" |'''Volume / µl''' | ||
| + | |- | ||
| + | | 5X Phusion HF buffer || align="right" |10 | ||
| + | |- | ||
| + | | 10 mM dNTP mix|| align="right" |1 | ||
| + | |- | ||
| + | | forward primer: O89 (Präfix_VP1_ex_RFC25_for) || align="right" |2.5 | ||
| + | |- | ||
| + | | reverse primer: O167 (pEGFP-C1_for) || align="right" |2.5 | ||
| + | |- | ||
| + | | DNA Template(1:100) || align="right" |1 µl | ||
| + | |- | ||
| + | | DMSO (2%)|| align="right" | - | ||
| + | |- | ||
| + | | Phusion Polymerase|| align="right" |0,5 | ||
| + | |- | ||
| + | |H<sub>2</sub>O|| align="right" | | ||
| + | |- | ||
| + | |Total volume|| align="right" |50 | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | '''PCR program:''' | ||
| + | {| border="1" | ||
| + | |'''PCR Program'''|| align="right" |'''temperature/ °C'''|| align="right" |'''time/ sec.''' | ||
| + | |- | ||
| + | |1|| align="right" |98 || align="right" |60 | ||
| + | |- | ||
| + | |2|| align="right" |98 || align="right" |15 | ||
| + | |- | ||
| + | |8x|| align="right" |60 || align="right" |25 | ||
| + | |- | ||
| + | |3 || align="right" |72 || align="right" |15 | ||
| + | |- | ||
| + | |4 || align="right" |98 || align="right" |15 | ||
| + | |- | ||
| + | |17x|| align="right" |66 || align="right" |25 | ||
| + | |- | ||
| + | |1x|| align="right" |72 || align="right" |15 | ||
| + | |- | ||
| + | |Hold|| align="right" |4 || align="right" | - | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | <gallery widths=450px heights=300px perrow=2 caption=""> | ||
| + | File:Freiburg10 20082010achim2.jpg | ||
| + | File:Freiburg10 20082010achim.jpg | ||
| + | </gallery> | ||
| + | <br/> | ||
| + | |||
| + | |||
| + | '''Digestion of PCR product and vector:''' | ||
| + | |||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | '''components''' || align="right" |'''PCR product /µl''' || align="right" |'''vector /µl''' | ||
| + | |- | ||
| + | | DNA || align="right" | 20 || align="right" | 5,2 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" | 3 || align="right" | 2 | ||
| + | |- | ||
| + | | Buffer 4 (10x)|| align="right" | 3 || align="right" |2 | ||
| + | |- | ||
| + | |Enzyme '''NgOMIV|| align="right" | 1 || align="right" | 1 | ||
| + | |- | ||
| + | |Enzyme '''SpeI || align="right" | 1 || align="right" | 1 | ||
| + | |- | ||
| + | |H2O|| align="right" | 2 || align="right" | 8,8 | ||
| + | |- | ||
| + | |'''Total volume (e.g. 15,20,25,30 µl)'''|| align="right" | 30 || align="right" |20 | ||
| + | |} | ||
| + | <br /> | ||
| + | Ligation with T4-Ligase and transformation with XL1B were carried out. Clones have to be picked tomorrow. | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''BioBrick assembly for pSB1C3_Affibody_Linker and pSB1C3_YFP_His-Tag'''</p>==== | ||
| + | Investigators: Volker | ||
| + | |||
| + | |||
| + | The aim of this experiment is to assemble the affibodies with the the five linkers and the YFP with the His-Tag. All cloning is done according to RFC 25 and will be used to test the affibody if it really binds the EGF-receptor, for the N-terminal fusions and to supply it to other groups in the registry.<br> | ||
| + | |||
| + | This experiment went wrong on Saturday 21.08. and was repeated on Sunday 22.08. and gave the following results. Results from the first experiment are not shown but can be senn in the written Labjournal. | ||
| + | |||
| + | <p style="font-size:15px; font-weight: bold; color: blue;">Digestion</p> | ||
| + | The following digestions were performed: | ||
| + | {| border="1" | ||
| + | | components | ||
| + | | Affibody | ||
| + | | Affibody | ||
| + | | Affibody | ||
| + | | Short Linker | ||
| + | | Middle Linker | ||
| + | | Long Linker | ||
| + | | SEG Linker | ||
| + | | GSAT Linker | ||
| + | | YFP | ||
| + | | His-Tag | ||
| + | |- | ||
| + | | DNA | ||
| + | |8.6 µl | ||
| + | |8.6 µl | ||
| + | |8.6 µl | ||
| + | |7.7 µl | ||
| + | |8.2 µl | ||
| + | |8.0 µl | ||
| + | |8.1 µl | ||
| + | |8.4 µl | ||
| + | |5.4 µl | ||
| + | |14 µl | ||
| + | |- | ||
| + | | BSA (100x) | ||
| + | |2µl | ||
| + | |2µl | ||
| + | |2µl | ||
| + | |0µl | ||
| + | |0µl | ||
| + | |0µl | ||
| + | |0µl | ||
| + | |0µl | ||
| + | |2µl | ||
| + | |0µl | ||
| + | |- | ||
| + | | Buffer (10x) | ||
| + | |2µl | ||
| + | |2µl | ||
| + | |2µl | ||
| + | |2µl | ||
| + | |2µl | ||
| + | |2µl | ||
| + | |2µl | ||
| + | |2µl | ||
| + | |2µl | ||
| + | |2µl | ||
| + | |- | ||
| + | |Enzyme 1 (1µl each) | ||
| + | |PstI | ||
| + | |PstI | ||
| + | |PstI | ||
| + | |PstI | ||
| + | |PstI | ||
| + | |PstI | ||
| + | |PstI | ||
| + | |PstI | ||
| + | |EcoRI | ||
| + | |EcoRI | ||
| + | |- | ||
| + | |Enzyme 2 (1µl each) | ||
| + | |AgeI | ||
| + | |AgeI | ||
| + | |AgeI | ||
| + | |NgoMIV | ||
| + | |NgoMIV | ||
| + | |NgoMIV | ||
| + | |NgoMIV | ||
| + | |NgoMIV | ||
| + | |AgeI | ||
| + | |NgoMIV | ||
| + | |- | ||
| + | |H<sub>2</sub>O | ||
| + | |5.4 µl | ||
| + | |5.4 µl | ||
| + | |5.4 µl | ||
| + | |8.3 µl | ||
| + | |7.8 µl | ||
| + | |8.0 µl | ||
| + | |7.9 µl | ||
| + | |7.6 µl | ||
| + | |8.6 µl | ||
| + | |2.0 µl | ||
| + | |||
| + | |- | ||
| + | |'''Total volume ''' | ||
| + | |20µl | ||
| + | |20µl | ||
| + | |20µl | ||
| + | |20µl | ||
| + | |20µl | ||
| + | |20µl | ||
| + | |20µl | ||
| + | |20µl | ||
| + | |20µl | ||
| + | |20µl | ||
| + | |} | ||
| + | |||
| + | Incubation: 2 h | ||
| + | |||
| + | <p style="font-size:15px; font-weight: bold; color: blue;">Preparative Agarose-Gel</p> | ||
| + | <br /> | ||
| + | 1 g Agarose, 100 ml TAE (1%), 6 GELRED (3-6µl), at 120Volt, running time: 130 minutes | ||
| + | <br /> | ||
| + | <br /> | ||
| + | {| border="1" cellspacing="0" cellpadding="2" bordercolor="black" | ||
| + | !Lane | ||
| + | !Sample | ||
| + | !Sample/µl] | ||
| + | !Loading dye (5x/6x)/µl | ||
| + | !Expected size 1 (Geneious) | ||
| + | !Expected size 2 (Geneious) | ||
| + | |-- | ||
| + | |1 | ||
| + | |Affibody | ||
| + | |20 µl | ||
| + | |4 µl | ||
| + | | bp | ||
| + | | bp | ||
| + | |-- | ||
| + | |2 | ||
| + | |Affibody | ||
| + | |20 µl | ||
| + | |4 µl | ||
| + | | bp | ||
| + | | bp | ||
| + | |-- | ||
| + | |3 | ||
| + | |Affibody | ||
| + | |20 µl | ||
| + | |4 µl | ||
| + | | bp | ||
| + | | bp | ||
| + | |-- | ||
| + | |10 | ||
| + | |His-Tag | ||
| + | |20 µl | ||
| + | |4 µl | ||
| + | | bp | ||
| + | | bp | ||
| + | |-- | ||
| + | |||
| + | |} | ||
| + | {| align=right | ||
| + | |} | ||
| + | |||
| + | |||
| + | <br /> | ||
| + | 2 g Agarose, 100 ml TAE (2%), 6 GELRED (3-6µl), at 120 Volt, running time: 130 minutes | ||
| + | <br /> | ||
| + | <br /> | ||
| + | {| border="1" cellspacing="0" cellpadding="2" bordercolor="black" | ||
| + | !Lane | ||
| + | !Sample | ||
| + | !Sample/µl] | ||
| + | !Loading dye (5x/6x)/µl | ||
| + | |-- | ||
| + | |4 | ||
| + | |Short Linker | ||
| + | |20 µl | ||
| + | |4 µl | ||
| + | |-- | ||
| + | |5 | ||
| + | |Medium Linker | ||
| + | |20 µl | ||
| + | |4 µl | ||
| + | |-- | ||
| + | |6 | ||
| + | |Long Linker | ||
| + | |20 µl | ||
| + | |4 µl | ||
| + | |-- | ||
| + | |7 | ||
| + | |SEG Linker | ||
| + | |20 µl | ||
| + | |4 µl | ||
| + | |-- | ||
| + | |8 | ||
| + | |GSAT Linker | ||
| + | |20 µl | ||
| + | |4 µl | ||
| + | |-- | ||
| + | |9 | ||
| + | |YFP | ||
| + | |20 µl | ||
| + | |4 µl | ||
| + | |-- | ||
| + | |} | ||
| + | {| align=right | ||
| + | |} | ||
| + | |||
| + | <gallery widths=450px heights=300px perrow=2 caption="Sequencing results of pSB1C3_hTERT"> | ||
| + | Image:Freiburg10_Affi-Linker preparative gel 1.jpg | ||
| + | Image:Freiburg10_Affi-Linker preparative gel 2.jpg | ||
| + | </gallery> | ||
| + | |||
| + | |||
| + | <p style="font-size:15px; font-weight: bold; color: blue;">Gel extraction</p> | ||
| + | {| border="1" | ||
| + | | align="left" | '''Sample''' | ||
| + | | align="left" | '''Weight''' | ||
| + | | align="left" | '''Volume''' | ||
| + | | align="left" | '''Concentration''' | ||
| + | |- | ||
| + | | align="left" | Affibody | ||
| + | | align="left" | 0.4g | ||
| + | | align="left" | 1200µl | ||
| + | | align="left" | 29.78 | ||
| + | |- | ||
| + | | align="left" | Affibody | ||
| + | | align="left" | 0.44g | ||
| + | | align="left" | 1320µl | ||
| + | | align="left" | 24.86 | ||
| + | |- | ||
| + | | align="left" | Short-Linker | ||
| + | | align="left" | 0.27 g | ||
| + | | align="left" | 810µl | ||
| + | | align="left" | 0.22 | ||
| + | |- | ||
| + | | align="left" | Middle-Linker | ||
| + | | align="left" | 0.38 g | ||
| + | | align="left" | 1140µl | ||
| + | | align="left" | 3.67 | ||
| + | |- | ||
| + | | align="left" | Long-Linker | ||
| + | | align="left" | 0.35 g | ||
| + | | align="left" | 1050µl | ||
| + | | align="left" | 1.31 | ||
| + | |- | ||
| + | | align="left" | SEG-Linker | ||
| + | | align="left" | 0.34 g | ||
| + | | align="left" | 1020µl | ||
| + | | align="left" | 2.13 | ||
| + | |- | ||
| + | | align="left" | GSAT-Linker | ||
| + | | align="left" | 0.22 g | ||
| + | | align="left" | 660µl | ||
| + | | align="left" | 3.58 | ||
| + | |- | ||
| + | |} | ||
| + | <br /> | ||
| + | |||
| + | |||
| + | <p style="font-size:15px; font-weight: bold; color: blue;">Ligation</p> | ||
| + | {| border="1" | ||
| + | | Sample | ||
| + | | '''Insert''' | ||
| + | | '''V(Insert)''' | ||
| + | | '''Vector''' | ||
| + | | '''V(Vector)''' | ||
| + | |- | ||
| + | |pSB1C3_Affibody_Short-Linker | ||
| + | |Affibody | ||
| + | |1.06µl | ||
| + | |Short-Linker | ||
| + | |6.94µl | ||
| + | |- | ||
| + | |pSB1C3_Affibody_Middle-Linker | ||
| + | |Affibody | ||
| + | |5.25µl | ||
| + | |Middle-Linker | ||
| + | |2.75µl | ||
| + | |- | ||
| + | |pSB1C3_Affibody_Long-Linker | ||
| + | |Affibody | ||
| + | |2.93µl | ||
| + | |Long-Linker | ||
| + | |5.07µl | ||
| + | |- | ||
| + | |pSB1C3_Affibody_SEG-Linker | ||
| + | |Affibody | ||
| + | |2.46µl | ||
| + | |SEG-Linker | ||
| + | |5.54µl | ||
| + | |- | ||
| + | |pSB1C3_Affibody_GSAT-Linker | ||
| + | |Affibody | ||
| + | |3.41µl | ||
| + | |GSAT-Linker | ||
| + | |4.59µl | ||
| + | |- | ||
| + | |} | ||
| + | <br /> | ||
| + | |||
| + | ===96. labday 21.08.2010=== | ||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Transfection of AAV293, Harvest viral particles, Transduction of HT1080 and Fluorescent microscopy'''</p>==== | ||
| + | <b> Investigators: Adrian, Kerstin </b> | ||
| + | |||
| + | <b>The Motivation</b><br /> | ||
| + | Measure the Transduction efficiency from 19.8 | ||
| + | <br /> | ||
| + | |||
| + | <b>The Motivation</b><br /> | ||
| + | Transfection of AAV293 cells from 19.8 (ten 10cm<sup>2</sup> dishes with 2.000.000 cells each) <font size="4" color="#F0000">with original standard protocoll!!!</font> We want to try different 2xHBS Buffers (pH) and their influence on transfection efficiency!<br /><br /> | ||
| + | <b>The Plan</b><br /> | ||
| + | <br /> | ||
| + | <b>Requiered goods</b>: 0,3 M CaCl<sub>2</sub> , 2x HBS-Buffer , autoclaved millipore water, falcons, eppis | ||
| + | <br /> | ||
| + | <b>Working steps</b>: | ||
| + | <ol> | ||
| + | <li>Remove the three plasmids to be co-transfected (the recombinant pAAV expression plasmid or control plasmid, pAAV-RC, and pHelper) from storage at –20°C.</li> | ||
| + | <li>Pipet 10 µg of each of the three plasmid DNA solutions into an eppi.</li> | ||
| + | <li>Add 1ml of 0.3 M CaCl<sub>2</sub> and mix gently</li> | ||
| + | <li>Pipet 1ml of 2 × HBS into a 15-ml falcon.</li> | ||
| + | <li>pipet the DNA/CaCl<sub>2</sub> solution dropwise into the falcon (use the Pipetus).</li> | ||
| + | <li>Apply the DNA/CaCl<sub>2</sub>/HBS suspension to the plate of cells in a dropwise fashion, swirling gently to distribute the DNA suspension evenly in the medium.</li> | ||
| + | <li>Return the tissue culture plate to the 37°C room for 6 hours.</li> | ||
| + | <li>At the end of the incubation period, remove the medium from the plate and wash cells with warm PBS, replace it with 10 ml of fresh DMEM growth medium.</li> | ||
| + | <li>Return the plate to the 37°C incubator for an additional 72 hours.</li> | ||
| + | </ol> | ||
| + | <br /> | ||
| + | We try 5 different 2xHBS buffers: | ||
| + | <font size="4" color="#FF0000"> | ||
| + | <ul> | ||
| + | <li>pH: 7.06</li> | ||
| + | <li>pH: 7.08</li> | ||
| + | <li>pH: 7.10</li> | ||
| + | <li>pH: 7.12</li> | ||
| + | <li>pH: 7.14</li> | ||
| + | </ul></font> | ||
| + | *1. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.06 | ||
| + | *2. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.08 | ||
| + | *3. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.10 | ||
| + | *4. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.12 | ||
| + | *5. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.14 | ||
| + | *6. Plate: 10µg of Rep/Cap, pHelper and GOI=TKGMK; pH(2xHBS)= 7.08 | ||
| + | *7. Plate: 10µg of Rep/Cap, pHelper and GOI=TKGMK; pH(2xHBS)= 7.10 | ||
| + | *8. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.14 | ||
| + | <br /> | ||
| + | <br /> | ||
| + | The transfected cells were seeded at 19.8 (2.000.000 cells per 10 cm<sup>2</sup>) | ||
| + | <br /> | ||
| + | Used Plasmids: Standart protocol: 10µg of each plasmid | ||
| + | <ul> | ||
| + | <li>GOI: mVENUS => P262, conc:1148ng/µl; used amount: 8,71µl</li> | ||
| + | <li>GOI: TKGMK => P82b, conc:1500ng/µl; used amount: 6,67µl</li> | ||
| + | <li>pHELPER => P64, conc:503ng/µl; used amount: 20µl</li> | ||
| + | <li>RepCap => 10.8: 1348ng/µl; used amount: 7,41µl</li> | ||
| + | |||
| + | |||
| + | |||
| + | <li> <b> Harvest viral paricles from 18.08.2010 and transduction of HT1080 cells following the standard protocol </b></li> | ||
| + | |||
| + | |||
| + | <li> <b> Analysis via Fluorescent microscopy (Transduction from 19.08.2010)</b> </li> | ||
| + | *Results: | ||
| + | </ul> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Midi-Prep of pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR clone1</b></p>==== | ||
| + | |||
| + | '''Investigators: Chris W. <br> | ||
| + | <p style="font-size:13px; color:#003399;"><b>Comment</b>: Midi-Preps of P 242, B200</p> | ||
| + | The Midi-Preps were performed according to the standard protocol yielding the following concentrations: | ||
| + | |||
| + | {| border="1" | ||
| + | | plasmid-no. || align="right" |P269|| align="right" |p270 | ||
| + | |- | ||
| + | | concentration (ng/µl)|| align="right" |1035,10 || align="right" |1376,95 | ||
| + | |- | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Re-Trafo of pSB1C3_SDM_SspI_Bla14FM'''</p>==== | ||
| + | Investigator: Stefan<br /> | ||
| + | <br /> | ||
| + | <p style="font-size:13px; color:#68bbff;">'''''Comment''''': Mini-Prep of pSB1C3_SDM_SspI_Bla14FM showed extremely low plasmid concentrations. It was proposed that this could correspond to using not the standard lab cells but rather XL-10 Gold cells for trafo. Therefore a re-trafo was performed using BL21 cells. 2 µl of each clone was added to BL21 cells and trafo was performed. Clones can be picked tonight (Volker)!</p> <br /> | ||
| + | ===97. labday 22.08.2010=== | ||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Test-digestion of pCerulean (P273,4,5), pSB1C3_VP1up (P279, 280), pSB1C3_NLS (P281,282), pAAV_RC_final (P283) </b></p>==== | ||
| + | Investigator: Patrick | ||
| + | <br> | ||
| + | Digestions: | ||
| + | * 0,5 µl EcoRI, 0,5 µl Enzyme PstI, 1 µl BSA (10x), 1 µl Buffer 4 (10x), 1 µl pCerulean (P73), 6 µl H2O; total volume = 10 µl | ||
| + | * 0,5 µl EcoRI, 0,5 µl Enzyme PstI, 1 µl BSA (10x), 1 µl Buffer 4 (10x), 1 µl pCerulean (P74), 6 µl H2O; total volume = 10 µl | ||
| + | * 0,5 µl EcoRI, 0,5 µl Enzyme PstI, 1 µl BSA (10x), 1 µl Buffer 4 (10x), 1 µl pCerulean (P75), 6 µl H2O; total volume = 10 µl | ||
| + | * 1 µl NotI, 1 µl BSA (10x), 1 µl Buffer 4 (10x), 1 µl pSB1C3_VP1up (P279), 6 µl H2O; total volume = 10 µl | ||
| + | * 1 µl NotI, 1 µl BSA (10x), 1 µl Buffer 4 (10x), 1 µl pSB1C3_VP1up (P280), 6 µl H2O; total volume = 10 µl | ||
| + | * 0,5 µl EcoRI, 0,5 µl Enzyme PstI, 1 µl BSA (10x), 1 µl Bufer 4 (10x), 1 µl pSB1C3_NLS (P281) , 6 µl H2O; total volume = 10 µl | ||
| + | * 0,5 µl EcoRI, 0,5 µl Enzyme PstI, 1 µl BSA (10x), 1 µl Bufer 4 (10x), 1 µl pSB1C3_NLS (P282) , 6 µl H2O; total volume = 10 µl | ||
| + | * 0,5 µl Acc65I, 0,5 µl BspMI, 1 µl BSA (10x), 1 µl Buffer 3 (10x) , 1 µl pAAV_RC_final , 6 µl H2O; total volume = 10 µl | ||
| + | Incubation time: 37°C, 1h | ||
| + | |||
| + | <br> | ||
| + | |||
| + | Expected size of the fragments: | ||
| + | *pCerulean clone 1 : 771 & 3915 bp | ||
| + | *pCerulean clone 2 : 771 & 3915 bp | ||
| + | *pCerulean clone 3 : 771 & 3915 bp | ||
| + | *pSB1C3_VP1up clone 1 : 468 & 2030 bp | ||
| + | *pSB1C3_VP1up clone 2 : 468 & 2030 bp | ||
| + | *pSB1C3_NLS clone 1 : 78 & 2031 bp | ||
| + | *pSB1C3_NLS clone 2 : 78 & 2031 bp | ||
| + | *pAAV_RC_final : 615 & 6720 bp | ||
| + | |||
| + | |||
| + | [[File:Freiburg10_tesdigestions20100823.jpg|500px]] | ||
| + | |||
| + | <br> | ||
| + | pCerulean clone 1 (P273), pSB1C3_VP1up clone 2 (P280), pSB1C3_NLS clone 2 (P282) and pAAV_RC_final (P283) were sent for sequencing although the results don't look very promising. <br> | ||
| + | Labelling: | ||
| + | * pCerulean clone 1 (P273) : iGEM2-CMV-F | ||
| + | * pSB1C3_VP1up clone 2 (P280) : iGEM1-VF2 | ||
| + | * pSB1C3_NLS clone 2 (P282) : iGEM276-O51_VR2 and iGEM3-O51_VR2 (was ordered twice due to an ordering mistake) | ||
| + | * pAAV_RC_final (P283) : iGEM4-SK | ||
| + | * pAAV_RC_final (P283) : iGEM4-VP1 pKEX rev | ||
| + | * pAAV_RC_final (P283) : iGEM4-Cap2800for | ||
| + | * pAAV_RC_final (P283) : iGEM4-Cap2800rev | ||
| + | * pAAV_RC_final (P283) : iGEM4-Cap3500for | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Mini-Prep of pCerulean</b></p>==== | ||
| + | <b>Investigator: Stefan</b><br> | ||
| + | <ul> | ||
| + | <b>Glycerol stocks were prepared:</b> | ||
| + | <li>B225 = pCerulean clone1</li> | ||
| + | <li>B226 = pCerulean clone2</li> | ||
| + | <li>B227 = pCerulean clone3</li> | ||
| + | <br /></ul> | ||
| + | <ul> | ||
| + | <b>Mini-Prep following the standard protocol</b> | ||
| + | <br /> | ||
| + | <li>P273 = pCerulean clone1 c= 419,52 ng/µl</li> | ||
| + | <li>P274 = pCerulean clone2 c= 383,37 ng/µl</li> | ||
| + | <li>P275 = pCerulean clone3 c= 382,72 ng/µl </li><br /> | ||
| + | <br /></ul> | ||
| + | Test digestion will be performed tomorrow. | ||
| + | |||
| + | ===98. labday 23.08.2010=== | ||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Primer design: Sequencing primer for BioBrick Assembly of vectorplasmid and phTERT qPCR primer</b></p>==== | ||
| + | <b>Investigator: Bea</b><br> | ||
| + | <p style="font-size:13px; color:#003399;"><b>Comment</b>: Two goals will be accomplished with this primers. | ||
| + | <ol> | ||
| + | <li>The first primer pair (left picture) serve as sequencing primers for the pSB1C3 vector plasmid. Problem was that the designed VR-2/VF-2 primers cannot be used for sequencing because the ITRs flank several assembled parts, mostly CMV, beta globin, GOI and hGH. These primers were designed to bind in the flanked region, directly after/before left and right ITR repsectivel and can therfore be used to sequence most of the flanked sequence inbetween the ITRs.</li> | ||
| + | <li>The second primer pair (right picture) will be used for performing the qPCR with th econtruct containing the phTERT promoter. The qPCR primers used for the other qRT-PCR cannot be used because these two primers anneal in the CMV region. The new designed primers bind in the phTERT region and can therefore bee used for determinining the infectious titers.</li> | ||
| + | </ol> | ||
| + | </p> | ||
| + | |||
| + | <gallery widths=450px heights=400px perrow=2 caption="Primer design. Ordered at Sigma Aldrich"> | ||
| + | Image:Freiburg10 Sequencingprimer pSB1C3 Bba 23 08 2010.jpg | ||
| + | Image:Freiburg10 phTERT qPCRprimer 23 08 2010.jpg | ||
| + | </gallery> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Mini-Prep of pSB1C3_CFP_middlelinker and pSB1C3_ZEGFR:1907 / testdigestion of pSB1C3_CFP_middlelinker and pSB1C3_lITR_pHTERT</b></p>==== | ||
| + | <b>Investigator: Jessica</b><br> | ||
| + | |||
| + | |||
| + | '''Mini-Prep following the standard protocoll''' | ||
| + | <br /> | ||
| + | *P276 PSB1C3_CFP_middlelinker clone1 c=207,7 ng/µl | ||
| + | *P277 PSB1C3_CFP_middlelinker clone2 c=203,0 ng/µl | ||
| + | *P278 PSB1C3_CFP_middlelinker clone3 c=227,9 ng/µl | ||
| + | *P279 pSB1C3_VP1up clone1 c=183,0 ng/µl | ||
| + | *P280 pSB1C3_VP1up clone2 c=178,1 ng/µl | ||
| + | *P281 pSB1C3_NLS clone1 c=155,7 ng/µl | ||
| + | *P282 pSB1C3_NLS clone2 c=142,6 ng/µl | ||
| + | *P283 pAAV_RC_final c=147,7 ng/µl | ||
| + | *P284 pSB1C3_ZEGFR:1907 clone 1 c=146,5 ng/µl | ||
| + | *P285 pSB1C3_ZEGFR:1907 clone 1 c=161,9 ng/µl<br> | ||
| + | <br /> | ||
| + | [[File:Cartoons48.jpg|thumb|right| pSB1C3_Zegfr:1907 wurde bereits von Stefan geprept... was hast du hier gemacht Jessy? Retrafo vom Glycerolstock???]] | ||
| + | <br> | ||
| + | <br> | ||
| + | <ul> | ||
| + | <b>Test digestion:</b> | ||
| + | <li>Perfomed with 10 µL of total volume</li> | ||
| + | </ul> | ||
| + | <table border=1 cellpadding=0 cellspacing=0 width=621 style='border-collapse: | ||
| + | collapse;table-layout:fixed;width:466pt'> | ||
| + | <tr height=29 style='mso-height-source:userset;height:21.75pt'> | ||
| + | <td height=29 class=xl6724191 width=143 style='height:21.75pt;width:107pt'> </td> | ||
| + | <td class=xl6824191 width=131 style='border-left:none;width:98pt'>Mastermix/µL</td> | ||
| + | <td class=xl6824191 width=87 style='border-left:none;width:65pt'>P256/µL</td> | ||
| + | <td class=xl6824191 width=84 style='border-left:none;width:63pt'>P257/µL</td> | ||
| + | <td class=xl6824191 width=90 style='border-left:none;width:68pt'>P276/µL</td> | ||
| + | <td class=xl6824191 width=86 style='border-left:none;width:65pt'>P277/µL</td> | ||
| + | <td class=xl6824191 width=86 style='border-left:none;width:65pt'>P278/µL</td> | ||
| + | </tr> | ||
| + | <tr height=29 style='mso-height-source:userset;height:21.75pt'> | ||
| + | <td height=29 class=xl6624191 width=143 style='height:21.75pt;border-top: | ||
| + | |||
| + | none;width:107pt'><span lang=EN-US style='mso-ansi-language:EN-US'>DNA</span></td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>-</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>5</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>5,2</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>4,3</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>4,4</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>4,0</td> | ||
| + | </tr> | ||
| + | <tr height=29 style='mso-height-source:userset;height:21.75pt;mso-yfti-irow: | ||
| + | |||
| + | 1'> | ||
| + | <td height=29 class=xl6624191 width=143 style='height:21.75pt;border-top: | ||
| + | |||
| + | none;width:107pt'><span lang=EN-US style='mso-ansi-language:EN-US'>BSA (10x)</span></td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>6</td> | ||
| + | <td rowspan=4 class=xl6524191 style='border-top:none'>1</td> | ||
| + | <td rowspan=4 class=xl6524191 style='border-top:none'>1</td> | ||
| + | <td rowspan=4 class=xl6524191 style='border-top:none'>1</td> | ||
| + | <td rowspan=4 class=xl6524191 style='border-top:none'>1</td> | ||
| + | <td rowspan=4 class=xl6524191 style='border-top:none'>1</td> | ||
| + | </tr> | ||
| + | <tr height=29 style='mso-height-source:userset;height:21.75pt;mso-yfti-irow: | ||
| + | |||
| + | 2'> | ||
| + | <td height=29 class=xl6624191 width=143 style='height:21.75pt;border-top: | ||
| + | |||
| + | none;width:107pt'><span lang=EN-US style='mso-ansi-language:EN-US'>Buffer | ||
| + | 4<span style='mso-spacerun:yes'> </span>(10x)</span></td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>6</td> | ||
| + | </tr> | ||
| + | <tr height=29 style='mso-height-source:userset;height:21.75pt;mso-yfti-irow: | ||
| + | |||
| + | 3'> | ||
| + | <td height=29 class=xl6624191 width=143 style='height:21.75pt;border-top: | ||
| + | |||
| + | none;width:107pt'><span lang=EN-US style='mso-ansi-language:EN-US'>Enzyme | ||
| + | XbaI<span style='mso-spacerun:yes'> </span></span></td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>3</td> | ||
| + | </tr> | ||
| + | <tr height=29 style='mso-height-source:userset;height:21.75pt;mso-yfti-irow: | ||
| + | |||
| + | 4'> | ||
| + | <td height=29 class=xl6624191 width=143 style='height:21.75pt;border-top: | ||
| + | |||
| + | none;width:107pt'><span lang=EN-US style='mso-ansi-language:EN-US'>Enzyme | ||
| + | PstI</span></td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>3</td> | ||
| + | </tr> | ||
| + | <tr height=29 style='mso-height-source:userset;height:21.75pt;mso-yfti-irow: | ||
| + | |||
| + | 5;mso-yfti-lastrow:yes'> | ||
| + | <td height=29 class=xl6624191 width=143 style='height:21.75pt;border-top: | ||
| + | |||
| + | none;width:107pt'><span lang=EN-US style='mso-ansi-language:EN-US'>H2O</span></td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>-</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>2</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>1,8</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>2,7</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>2,6</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>3,0</td> | ||
| + | </tr> | ||
| + | <tr height=29 style='mso-height-source:userset;height:21.75pt'> | ||
| + | <td height=29 class=xl6624191 width=143 style='height:21.75pt;border-top: | ||
| + | |||
| + | none;width:107pt'>Total volume</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>18</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>10</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>10</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>10</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>10</td> | ||
| + | <td class=xl6524191 style='border-top:none;border-left:none'>10</td> | ||
| + | </tr> | ||
| + | <tr height=0 style='display:none'> | ||
| + | <td width=143 style='width:107pt'></td> | ||
| + | <td width=131 style='width:98pt'></td> | ||
| + | <td width=87 style='width:65pt'></td> | ||
| + | <td width=84 style='width:63pt'></td> | ||
| + | <td width=90 style='width:68pt'></td> | ||
| + | <td width=86 style='width:65pt'></td> | ||
| + | </tr> | ||
| + | </table> | ||
| + | <br /> | ||
| + | *incubation time: 50' | ||
| + | *incubation temparatur: 37°C | ||
| + | <br /> | ||
| + | [[File:Freiburg10 04 08 2010 Test digestion of pSB1C3_lITR_phTERT and pSB1C3_CFP_middlelinker.jpg|600px]] | ||
| + | <br /> | ||
| + | *5µl new ladder mix | ||
| + | <b> Sequencing</b>: | ||
| + | <ul> | ||
| + | <li> Plasmid used: P276: pSB1C3_CFP_middlelinker</li> | ||
| + | <li> Primer used: VR-2</li> | ||
| + | <li> Tube name: JG 276</li> | ||
| + | </ul> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Continuation of BioBrick assembly for pSB1C3_Affibody_Linker'''</p>==== | ||
| + | |||
| + | <b> Investigator: Anna</b> | ||
| + | |||
| + | <p style="font-size:13px; color:#003399;"><b>Comment</b>: Transformation was done of pSB1C3_Affibody with Short-Linker, Middle-Linker, Long-Linker, SEG-Linker and GSAT-Linker. Unfortunately, the digestion of pSB1C3_YFP_His-Tag didn't work (see 20.08).</p> | ||
| + | |||
| + | Transformation was done following the standard protocol using XL1 blue cells. | ||
| + | |||
| + | <p style="font-size:13px; color:#003399;"><b>To do:</b> Repetition of BioBrick assembly for pSB1C3_YFP_His-Tag.</p> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;">'''Cloning of pSB1C3_ß-globin_YFP and pSB1C3_hGH_rITR'''</p>==== | ||
| + | |||
| + | <b> Investigator: Anna</b> | ||
| + | |||
| + | <p style="font-size:13px; color:#003399;"><b>Comment</b>: pSB1C3_ß-globin_mVenus (p178) was digested with SpeI and PstI and pSB1C3_hGH_rITR (p186) was digested with XbaI and PstI .</p> | ||
| + | |||
| + | Vector: <br> | ||
| + | *pSB1C3_ß-globin_YFP (p178): c= 237,2 ng/µl<br> | ||
| + | Insert: <br> | ||
| + | *pSB1C3_hGH_rITR (p186): c= 105,2 ng/µl<br> | ||
| + | |||
| + | |||
| + | |||
| + | {| border="1" | ||
| + | | components || align="right" |Vector (p178)|| align="right" |Insert (p186) | ||
| + | |- | ||
| + | | DNA || align="right" |4,2 || align="right" | 14 | ||
| + | |- | ||
| + | | BSA (10x) || align="right" |1,5|| align="right" | 2,5 | ||
| + | |- | ||
| + | | Buffer 4 (10x)|| align="right" |1,5 || align="right" | 2,5 | ||
| + | |- | ||
| + | |Enzyme EcoRI || align="right" |1 || align="right" | 1,5 | ||
| + | |- | ||
| + | |Enzyme PstI || align="right" |1|| align="right" | 1,5 | ||
| + | |- | ||
| + | |H<sub>2</sub>O|| align="right" |5,8 || align="right" | 3 | ||
| + | |- | ||
| + | |'''Total volume '''|| align="right" |15 || align="right" | 25 | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | *Gelextraction of pSB1C3_RFC25_CFP: | ||
| + | 0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 120 Volt, running time:55 <br /> | ||
| + | Marker: GeneRuler ladder mix (Fermentas) | ||
| + | <br /> | ||
| + | <br /> | ||
| + | <br /> | ||
| + | <br /> | ||
| + | |||
| + | [[file:Freiburg10_.jpg]] | ||
| + | <br /> | ||
| + | |||
| + | |||
| + | |||
| + | *c(pSB1C3_ß-globin_YFP)= 16.9 ng/µl; size: 3293 bp<br /> | ||
| + | *c(hGH_rITR)= 7,5 ng/µl; size: 657 bp | ||
| + | |||
| + | <br /> | ||
| + | |||
| + | *Ligation of PCR products and vector: | ||
| + | |||
| + | For the Ligation 1µl T4 buffer (2x) and 1µl T4 ligase were used. Incubation time: 30 min | ||
| + | <br /> | ||
| + | {| border="1" | ||
| + | | ''' ''' || align="right" |'''vector /µl''' || align="right" |'''insert /µl''' | ||
| + | |- | ||
| + | | pSB1C3_ß-globin_mVenus + hGH_rITR || align="right" |3,41 || align="right" |4,59 | ||
| + | |- | ||
| + | |} | ||
| + | <br /> | ||
| + | |||
| + | *Transformation: | ||
| + | |||
| + | The transformation was done following the standard protocol using XL1 blue cells.<br /> | ||
| + | <br /> | ||
| + | |||
| + | <p style="font-size:13px; color:#003399;"><b>To do:</b> Picking clones of pSB1C3_ß-globin_mVenus_hGH_rITR.</p> | ||
| + | |||
| + | ====<p style="font-size:15px; background-color:#66bbff;"><b>Repetition: Quickchange site-directed mutagenesis of pSB1C3_SDM_SspI_Bla14FM</b></p>==== | ||
| + | <b>Investigator: Stefan</b><br> | ||
| + | |||
| + | '''Comment:''' To delete a PvuII restriction site in the CAT marker, a SDM will be performed.<br /> | ||
| + | |||
| + | *'''PCR reaction:''' | ||
| + | |||
| + | {| border="1" | ||
| + | | '''Ingredients''' || align="right" |'''v / µl''' | ||
| + | |- | ||
| + | | 10x Pfu Ultra II buffer || align="right" |2,5 | ||
| + | |- | ||
| + | | dNTP|| align="right" |0,5 | ||
| + | |- | ||
| + | | forward primer: O109 (pSB1C3 PvuII for)|| align="right" |0,59 | ||
| + | |- | ||
| + | | reverse primer: O108 (pSB1C3 PvuII rev) || align="right" |0,58 | ||
| + | |- | ||
| + | | DNA Template 10ng (1:100 dilution) || align="right" |4 | ||
| + | |- | ||
| + | | DMSO (1%)|| align="right" | 0,5 | ||
| + | |- | ||
| + | | PfuUltra II fusion || align="right" |0,5 | ||
| + | |- | ||
| + | |H<sub>2</sub>O|| align="right" |15,83 | ||
| + | |- | ||
| + | |Total volume|| align="right" |25 | ||
| + | |} | ||
| + | <br> | ||
| + | |||
| + | *'''PCR program:''' | ||
| + | |||
Revision as of 11:39, 6 September 2010

76.Labortag 01.08.2010
Picking clones of pSB1C3_CMV
Investigator: Bea
Comments: Trafo was performed friday, and trafo plate were incubated over night. Clones need to be picked in order to perform Mini-Prep
- Bacterial strain used: XL1-B
- Two clones were picked of trafo plate
- Inoculating of 10 mL DYT medium containing 10µL chloramphenicol
- Put in 37°C room on rotary shaker
77.Labortag 02.08.2010
Sequenzing of pSB1C3_RFC25_longlinker, pSB1C3_RFC25_SEG, pSB1C3_RFC25_GSAT
Investigator: Jessica
Comments: Linkers (we got from Gerrit) we cloned in the RFC25-standard (pSB1C3_RFC25). Result looks good, linkers are in the vector pSb1C3_RFC25
- pSB1C3_RFC25_longlinker P105 und P106
- pSB1C3_RFC25_SEG P107 und P108
- pSB1C3_RFC25_GSAT P109 und P110
Sequencing of pGA14_Prefix-leftITR
Investigator: Hanna
Eventually GATC partially managed to sequence pGA14_Prefix-left ITR clone 1 and 2.
The sequencing results were better than before but in the ITR region still not evaluable due to the strong secondary structures.
It seemed that, despite of successful insertion of the RFC10-Prefix, something went wrong because there were 2 bases missing in the NotI restriction site. But by having a closer look, it became obvious that the Geneious misinterpreted the chromatogram and overlaid the peaks of the missing bases.

For the first time sequencing of parts of the 3' end worked: The remaining NotI restriction site, which was already eliminated this week and also the "backbone-PstI" can be seen:

Sequencing of very GC-rich regions :) :) :)

Sequencing of clone 4 delivered that the ITR and the RFC10-Prefix were not inserted in the vector. The RFC25 standard of the multiple cloning site is still in the plasmid:

Therefore clone 4 was dicarded from the plasmid- and glycerol stock.
Repetition of test digestion pSB1C3_CFP SDM SspI and PvuII
Investigator: Jessica
- buffer used: 2; Restriction-enzymes used: Enzyme 1: SspI ; Enzyme 2: PvuII
- Plasmids (all: ~ 700 ng/µL):
- pSB1C3_CFP P51.1
- pSB1C3_SDM_SspI P125
- pSB1C3_SDM_PvuII P129
- pSB1C3_SDM_PvuII P131
| Components | Mastermix | P125/µL | P51.1.1/µL | P129/µL | P131/µL | P51.1.2/µL |
| DNA | - | 2,4 | 2,3 | 1,6 | 5,3 | 5,3 |
| BSA (10x) | 6 | 1 | 1 | 1 | 1 | 1 |
| Buffer 2 (10x) | 6 | 1 | 1 | 1 | 12 | 1 |
| Enzyme 1 SspI (68) | - | 1 | 1 | - | - | - |
| Enzyme 2 PvuII (50) | - | - | - | 1 | 1 | 1 |
| H2O | - | 4,6 | 4,7 | 5,4 | 1,7 | 1,7 |
| Total volume | 12 | 10 | 10 | 10 | 10 | 10 |
- Incubation: 65 minutes
Agarosegel
0.45 g Agarose, 50 ml TEB (0,5 %), 3 µL GELRED, at 115 Volt, running time: 45 minutes
- Marker: GeneRuler ladder mix (6x)
| Marker | P125 | P51.1 SspI | - | P129 | P131 | P51.1 PvuII | |
|---|---|---|---|---|---|---|---|
| Lane | 7 µL | 10 µL | 10 µL | - | 10 µL | 10 µL | 10 µL |
- P125 isn't cut, therefore you see the bands for supercoiled, nicked and relaxed
- P51.1 is cut once, therefore you can see a faster running band
- P129 and P131 should be cut once because of the PvuII in the CFP but there is another band we can't identify
- P51.1 should be cut two times but there is also another unidentified band
- perhaps there is another PvuII in the vector we can't find (???)
Comments: SDM of SspI is ready! SDM of PvuII looks confusing because of the lower band. P 129 will be sequenced but this doesn't solve this problem. there has to be one more rs we don't know, perhaps. we will check it...anywise
Mini-Prep and Test digestion of pSB1C3_CMV
Investigator: Kerstin
Nanodrop
- pSB1C3_CMV clone1: 157,6 ng/µl P145
- pSB1C3_CMV clone2: 155,6 ng/µl P146
Test digestion 900ng DNA
| components | P145 /µl | P146 /µl |
| DNA | 5,7 | 5,8 |
| BSA (10x) | 1,5 | 1,5 |
| Buffer 4 (10x) | 0,5 | 0,5 |
| Enzyme 1: XbaI | 0,5 | 1,5 |
| Enzyme 2: PstI | 0,5 | 0,5 |
| H2O | 5,3 | 5,2 |
| Total volume (e.g. 15,20,25,30 µl) | 15 | 15 |
agarose gel: 1,5%, digestion: 90 minutes, 37°C
P145 is sent for sequencing with Reverse Primer (VR2)
Comments:Results look good. CMV has the expected size (~ 662bp)
</p>
Sent for sequencing
Investigator: Jessica
Hanna
- P145 left ITR
- Primer: GATC_std_pQE-FP and GATC_std_m13-FP
- P150 right ITR
- Primer: GATC_std_pQE-FP and GATC_std_m13-FP
Anissa
- P136 pSB1C3_hgh
- Primer: O51_Reverse Primer (VR2)
- P139 pAAV_BamHI
- Primer: O36_Rep_1250_rev
- P143 pAAV_RC_SalI
- Primer: O53_Rep_1250_for
- P141 pGL3_QTERT
- Primer: GATC_std_pTal-FP
- P133 pSB1C3_betaGlobin
- Primer: O51_ReversePrimer (VR2)
- P145 pSB1C3_CMV
- Primer: O51_ReversePrimer (VR2)
Jessica
- P129 pSB1C3_SDM_PvuII
- Primer: GATC_std_pTeSp-2
Preparation of competent E.coli
Investigator: Jessica
- 15ml DYT was prepared with 15µl tetracycline and inoculate with XL1B
- 10ml DYT w/o antibiotics was prepared and inoculate with BL21 (just for glycerol stock, no competent cells)
both were incubate over night in 37°C room
Last Mini-Prep and test digestion of ITRs
Investigator: Hanna
Plasmid Mini-Prep
- new vector name: pGA14_RFC10_leftITR and pGA14_RFC10_rightITR
Glycerol Stocks
pGA14_RFC10_leftITR
| Clone 1 | Clone 2 | Clone 3 | |
| Bacteria strain | XL1blue | XL1blue | XL1blue |
| Plasmidname | pGA14_RFC10_leftITR | pGA14_RFC10_leftITR | pGA14_RFC10_leftITR |
| Date | 2.8.10 | 2.8.10 | 2.8.10 |
| given glycerol-stock no. | B115 | B116 | B117 |
| given plasmid no. | P147 | P148 | P149 |
pGA14_RFC10_rightITR
| Clone 1 | Clone 2 | Clone 3 | |
| Bacteria strain | XL1blue | XL1blue | XL1blue |
| Plasmidname | pGA14_RFC10_rightITR | pGA14_RFC10_rightITR | pGA14_RFC10_rightITR |
| Date | 2.8.10 | 2.8.10 | 2.8.10 |
| given glycerol-stock no. | B112 | B113 | B114 |
| given plasmid no. | P150 | P151 | P152 |
Test digestion
- buffer used: 4 ; Restriction-enzymes used: Enzyme 1 XbaI ; Enzyme 2 SpeI
- Plasmid: pGA14_RFC10_leftITR:
- Given Plasmid-Number: P147; DNA concentration: 240.22 ng/µL ;
- Given Plasmid-Number: P148; DNA concentration: 268.97 ng/µL ;
- Given Plasmid-Number: P149; DNA concentration: 234.16 ng/µL ;
- Plasmid: pGA14_RFC10_rightITR:
- Given Plasmid-Number: P150; DNA concentration: 221.71 ng/µL ;
- Given Plasmid-Number: P151; DNA concentration: 191.13 ng/µL ;
- Given Plasmid-Number: P152; DNA concentration: 215.14 ng/µL ;
| Components | P147 Volume/µL | P150 Volume/µL |
| DNA | 4.2 | 4.5 |
| BSA (10x) | 1 | 1 |
| Buffer no. 4 (10x) | 1 | 1 |
| Enzyme 1 XbaI | 0.5 | 0.5 |
| Enzyme 2 SpeI | 0.5 | 0.5 |
| H2O | 2.8 | 2.5 |
| Total volume | P147 | P150 |
- Incubation: 1 h
Agarose-Gel:
0.83 g Agarose, 53 mL TBE (1.57%), 3 µL GELRED, at 115 Volt, running time: 45 minutes
| Sample | Sample/µl] | Loading dye (5x)/µl | Expected size 1 (Geneious) | Expected size 2 (Geneious) |
|---|---|---|---|---|
| P147 | 10 µl | 2 µl | 157 bp | 2902 bp |
| P150 | 10 µl | 2 µl | 156 bp | 2903 bp |
- Marker: GeneRuler ladder mix
| Marker /µL | Sample P147 /µL | Sample P150 /µL | |
|---|---|---|---|
| Lane | 6.5 | 12 | 12 |
Test digestion delivered positive results for both ITRs: The expected fragment sizes (156 resp. 157 bp) could be detected in the referring range between the 100 and 200 bp marker nucleotides:

Comment: In order to verify the results, P147 and P150 were sent for sequencing. Because sequencing never worked before, they will be sequenced under special conditions this time.
BioBrick production: mGMK in pSB1C3
Investigator: Stefan
Backbone taken from:
- pSB1C3_CFP (P51.2): 151,1 ng/µl
Insert taken from:
- pAAV_RFC25_mGMK (P100): 411,3 ng/µl
| components | pSB1C3_CFP(P51.2) /µl | pAAV_RFC25_mGMK (P100) /µl |
| DNA | 8 | 5 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| XbaI | 1 | 1 |
| AgeI | 1 | 1 |
| H2O | 6 | 9 |
| Total volume | 20 | 20 |
- 1% Agarose gel
- 3 µl Gelred
- 7 µl DNA-Ladder-Mix
- 115 Volt, running time: 50 minutes
Afer 50 minutes a picture (see below) was taken. Because there was no clear pattern (two bands) for the pSB1C3 backbone, the band of the insert mGMK was cut out and the gel ran another 20 minutes. After that, there were still two bands. Both were cut out and it was continued with two possible backbones. They were labeled vector (lower) and vector (upper).
| Sample/µl | Expected size/bp |
| Vector pSB1C3 | 2082 |
| Insert mGMK | 603 |
- weight of insert mGMK gel extract: 0,17 g
- weight of vector (lower) pSB1C3 gel extract: 0,08 g
- weight of vector (upper) pSB1C3 gel extract: 0,09 g
Nanodrop
- insert mGMK : 4,13 ng/µl
- vector (lower) pSB1C: 5,47 ng/µl
- vector (upper) pSB1C: 0,60 ng/µl => for calculation of ligation approach calculated with 1 ng/µl
Ligation
- vector (lower) : insert - 4,16 µl : 4,84 µl
- vector (upper) : insert - 7,43 µl : 1,57 µl
Transformation
Two approaches were prepared, one for each vector and put in 37 °C room overnight.
Sequencing results of SDM Rep 68 & 78
Investigator: Hanna and Volker
The sequencing results delivered that the PstI restriction sites of Rep 68 and Rep 78 were successfully deleted via side-directed mutageneis:
78.Labortag 03.08.2010
Results of sequencing pSB1C3_CFP_SDM_PvuII
Investigator: Jessica
SDM of PvuII:
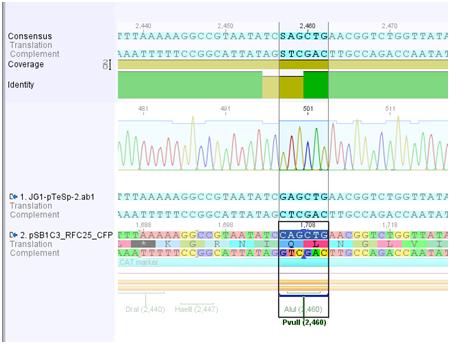
PvuII is succesfully deleted in the vector pSB1C3_RFC25_CFP
Repetition of the quickchange site-directed mutagenesis of pAAV_RC_1.1_SalI
Investigator: Kerstin,Anissa
Comments: Sequenzing revealed no mutagenesis of SalI in pAAV_RC_1.1_SalI, so SDM now will be repeated.
PCR-reaction:
| Volume / µl | ingredients | recommended /µl |
| 2,5 | 10X Pfu Ultra II buffer | 2,5 |
| 4,18 | template (~10 ng) | 4,18 of 1:100 dilution |
| 0,58 | forward primer: O68 | 62,5 ng |
| 0,58 | reverse primer: O69 | 62,5 ng |
| 0,5 | dNTP | 250 µM each dNTP |
| 16,16 | H2O | |
| 0,5 | PfuUltra II fusion (1.25) |
PCR program:
| Rounds | temperature/ °C | Time |
| 1 | 95 | 2 minutes |
| 20 | 95 | 30 seconds |
| 20 | 80,4 | 1 minute |
| 20 | 68 | 7,5 minutes |
Plasmid was transformed into BL21 cells. Clones have to be picked tomorrow.
Comments:NO CLONES, trafo didn't work, because annealing temperature was totally to high! (80,4 °C instead of 55°C) annealing temperature should always be 55°C (in case of troubleshooting temperature can be increased up to MAX. 68°C !)
PCR for Biobrickproduction of Rep 40, 52, 68, 78 and APP
Investigators: Volker, Anna
Aim of the experiment:
We wanted to produce biobricks from the expression plasmids Rep40ex, Rep52 ex, the Rep 68 and 78 in which the PstI at position 310 was silenced and the AAP. This was performed by a PCR with primes that annealed to the ends of the later biobrick but also contained restriction sites that can be cut afterwards and cloned into pSB1C3.
- Plasmids used as template:
Rep_68_ex (p119): c = 470,6 ng/µl
Rep_78_(p122): c = 201,08 ng/µl
Rep_40_(p22.2): c = 673,1 ng/µl
Rep_52_(p23.2): c = 532,22 ng/µl
pAAV-RC containing AAP ORF (p50): c = 378,5 ng/µl
- Primer used:
For Rep_40_ex: Praefix_40_52_ex & Suffix_40_68_ex
For Rep_52_ex: Praefix_40_52_ex & Suffix_52_78_ex
For Rep_68_ex: Praefix_68_78_ex & Suffix_40_68_ex
For Rep_78_ex: Praefix_68_78_ex & Suffix_52_78_ex
For AAP_ex: Praefix_AAP_ex & Suffix_AAP_ex
- PCR:
(was performed following the standard protocol)
| Ingredients | Volume / µl | Rep68 | Rep78 | Rep40 | Rep52 | AAP |
| 5X Phusion HF buffer | 10 | |||||
| 10 mM dNTP mix | 1 | |||||
| forward primer: | 2,5 | |||||
| reverse primer: | 2,5 | |||||
| DNA Template | *** | 4,2 µl | 1 µl | 2,9 µl | 3,8 µl | 5,3 µl |
| Phusion Polymerase | 0,5 | |||||
| H2O | *** | 28,3 µl | 31,5 µl | 29,6 µl | 28,7 µl | 27,2 µl |
| Total volume | 50 |
PCR program:
| PCR Program | temperature/ °C | Time | Rep40 | Rep50 | Rep68 | Rep78 | AAP |
| 1 | 98 | 1min | |||||
| 2 | 98 | 15s | |||||
| 8x | *** | 25s | 63°C | 62°C | 63°C | 62°C | 64°C |
| 3 | 72 | *** | 15s | 18s | 24s | 27s | 10s |
| 4 | 98 | 15s | |||||
| 17x | *** | 25s | 68°C | 66°C | 68°C | 64°C | 68°C |
| 5 | 72 | *** | 15s | 18s | 24s | 27s | 10s |
| 6x | 72 | 5min | |||||
| Hold | 4 |
Five µl of PCR-product were used for an analytical gel to see if the PCR worked.
Comment: The result is that the PCR worked for the longer Rep proteins (68&78) with which a Quick-change was performed to remove the PstI(310) but not with the constructs for the smaller Rep variants (40&52) that were recieved from PD Kleinschmidt. As the longer Rep variants the AAP PCR Reaction also resulted in a PCR-product of the expected size. Unfortunately there were a secondary band of ~90bp for Rep_68_ex. For this reason we performed a praeparative gel and decided to cut the bands of all PCR products.
The bands marked in the gel picture were cut and a gel extraction was performed.
Gel extraction
Gel measurement:
| Sample | Weight | Volume | Concentration |
| Rep_68_ex | 0,1 g | 20 µl | 17,6 ng/µl |
| Rep_78_ex | 0,15g | 20 µl | 80,6 ng/µl |
| AAP_ex | 0,19 g | 20µl | 77,78 ng/µl |
Continuation of preparation of competent E.coli
Investigator: Jessica
preparation of competent XL1blue was finished according to the standard protocol
- aliquots of 60µl (to use for 1 trafo) are stored in -80°C freezer
Sequencing of pSB1C3
Investigator: Jessica
Comments: We sent for sequencing pSB1C3_RFC25_longlinker (P105) to check the strange result of the test digestion of pSB1C3_SDM_PvuII(01.08.10.
pSB1C3_SDM_PvuII is too long for sequencing wherefore we sent pSB1C3_RFC25_longlinker because the longlinker just have 54kb (bp, nicht kb (Volker) (necessary is just the backbone we don't have.
- Primer:
- RESgen-241698
- Reverse primer (VR2)
- o_F1-fw (from Gerrit)
Design of Primer for p5 Promoter WT and TATA-less BioBrick production
Investigator: Hanna
79.Labortag 04.08.2010
Cloning of right ITR and left ITR into pSB1C3_RFC25_CFP
Intention: Get biobricks ready.
Investigator: Patrick
pGA14_lITR_RFC10 (P147), pGA14_rightITR (P150) and pSB1C3_RFC25_CFP (P51.1) were digested with EcoRI and PstI (Buffer 4) according to the standard protocol. Digestion Time: 80 minutes.
GelRun: 1% Agarose Gel. Expected results (from left to right):
- pSB1C3_RFC25_CFP: about 2100 bp and 800 bp.
- pGA14_leftITR_RFC10: the size of the insert should be 135 bp.
- pGA14_rightITR_RFC10: the size of the insert should be 138bp.
The mutual vector (now without CFP) and inserts (left ITR and right ITR) were cut out. The gelextraction was performed according to the standard protocol. DNA concentration of the extracts:
- pSB1C3_RFC25_CFP: 2,9 ng/µl
- left ITR: 1,4 ng/µl
- right ITR: 2,0 ng/µl
The Quick Ligation was not performed according to the standard protocol:
- left ITR + pSB1C3: 5µl Buffer, 1 µl Quick-Ligase, 2,5 µl left ITR, 1,5 µl pSB1C3_RFC25.
- right ITR + pSB1C3: 5µl Buffer, 1 µl Quick-Ligase, 2,5 µl right ITR, 1,5 µl pSB1C3_RFC25.
Transformation: performed according to the standard protocol (BL21). The cells were plated on a agar plate with chloramphenicol. The clones will be picked tomorrow.
Cloning of SDM SspI and SDM PvuII
Investigator: Jessica
- Vector: name: pSB1C3_SDM_SspI P125
- Insert: name: pSB1C3_SDM_PvuII P129
- new vector name: pSB1c3_SDM_SspI/PvuII P157
- buffer used: 4 ; Restriction-enzymes used: Enzyme 1 (no. Lab: 141) BstI ; Enzyme 2 (no.Lab: 144) BpmI
- DNA concentration (vector): 321,9 ng/µl ; DNA concentration (insert): 308,4 µg/µl
| components | volume of pSB1C3_SDM_SspI /µl | volume of pSB1C3_SDm_PvuII /µl |
| DNA | 4,66 | 4,87 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| Enzyme BstI (no.Lab:141) | 1 | 1 |
| Enzyme BpmI (no.Lab:144) | 1 | 1 |
| H2O | 9,34 | 9,13 |
| Total volume (e.g. 15,20,25,30 µl) | 20 | 20 |
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 130 Volt, running time:45
Loading plan for agarose gel:
Marker used: GeneRuler ladder mix (Fermentas)
| Marker | Sample 125, 20µl | Sample 129, 20µl | |
|---|---|---|---|
| Lane | 1 | 3 | 5 |
Results: digesttemperature of BstI is 55°C, plasmid was just digested at 37°C... :( (
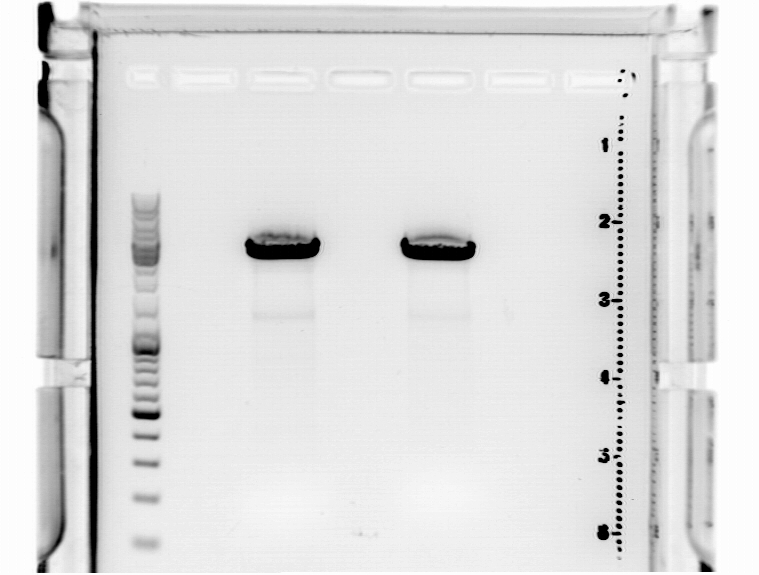
2nd Repetition of Quickchange site-directed mutagenesis of pAAV_RC_1.1_SalI
Investigator: Kerstin, Anissa
Comments: Last trafo didn't work. Annealing temperature was to high (80,4°C instead of 55°C)
Two approaches were made (short and long PCR:
PCR-reaction:
| Volume / µl | ingredients | recommended /µl |
| 2,5 | 10X Pfu Ultra II buffer | 2,5 |
| 1,83 | template (~10 ng): p139 (c: 546,97 ng/µl) | 1,83µl of 1:100 dilution |
| 0,58 | forward primer: O68 | 62,5 ng |
| 0,58 | reverse primer: O69 | 62,5 ng |
| 0,5 | dNTP | 250 µM each dNTP |
| 18,51 | H2O | |
| 0,5 | PfuUltra II fusion (1.25) |
PCR program (long):
| Cycles | temperature/ °C | Time |
| 1 | 95 | 2 minutes |
| 20 | 95 | 30 seconds |
| 20 | 55 | 1 minute |
| 20 | 68 | 7,5 minutes |
PCR program (short):
| Cycles | temperature/ °C | Time |
| 1 | 95 | 2 minutes |
| 20 | 95 | 30 seconds |
| 20 | 55 | 1 minute |
| 20 | 68 | 4 minutes |
Plasmids were transformed into BL21 cells. Clones have to be picked tomorrow.
Mini-Prep of pSB1C3_mGMK
Investigator: Chris W., Bea
Vector name: pSB1C3_mGMK upper band 1.1 and pSB1C3_mGMK upper band 1.2 and pSB1C3_mGMK lower band 2.1 and pSB1C3_mGMK lower band 2.2
Mini-Prep following the standart Protokoll
Test digestion:
- Perfomed with 15 µL of total volume with all four clones
| Mastermix/µL | P153/µL | P154/µL | P155/µL | P156/µL | |
| DNA | - | 3,5 | 3,5 | 3,5 | 3,5 |
| BSA (10x) | 7,5 | 4 | 4 | 4 | 4 |
| Buffer 4 (10x) | 7,5 | ||||
| Enzyme XbaI | 2,5 | ||||
| Enzyme AgeI | 2,5 | ||||
| H2O | - | 7,5 | 7,5 | 7,5 | 7,5 |
| Total volume | 15 | 15 | 15 | 15 | 15 |
- All four samples were loaded on a 1% agarose gel
- P153 = pSB1C3_mGMK upper band 1.1
- P154 = pSB1C3_mGMK upper band 1.2
- P155 = pSB1C3_mGMK lower band 1.1
- P156 = pSB1C3_mGMK lower band 1.2
- Plasmid used: P156: pSB1C3_mGMK lower band 1.2
- Primer used: VR-2
- Tube name: Bea_1
cell culture
Investigator: Adrian
Plan for the next virus production procedures
The Motivation: investigation of the influence of different GOI amounts.
The Plan: keep rep/cap and pHelper amount stable (3,3 µg each => 6,6 µg), differ the GOI (YFP) amount in each stock.
Viral Stocks:
- 3,3 µg YFP + 3,3 µg pHelper + 3,3 rep/cap
- 10 µg YFP + 3,3 µg pHelper + 3,3 rep/cap
- 20 µg YFP + 3,3 µg pHelper + 3,3 rep/cap
- 3,3 µg pHelper + 3,3 rep/cap (without GOI !!!)
- one GMK_TK clone (with the confirmed sequence) + 3,3 µg pHelper + 3,3 rep/cap
Transduction plan
| A | 150µl AAV stock 1 | 300µl AAV stock 1 | control no Virus |
|---|---|---|---|
| B | 150µl AAV stock 2 | 300µl AAV stock 2 | 150µl AAV without GOI |
Repetition of cloning of SDM SspI and SDM PvuII
Investigator: Jessica
- Vector: name: pSB1C3_SDM_SspI P125
- Insert: name: pSB1C3_SDM_PvuII P129
- new vector name: pSB1C3_RFC25_SDM_SspI/PvuII P157
- buffer used: 4 ; Restriction-enzymes used: Enzyme 1 (no. Lab: 141) BtsI ; Enzyme 2 (no.Lab: 144) BpmI
- DNA concentration (vector): 321,9 ng/µl ; DNA concentration (insert): 308,4 µg/µl
| components | volume of pSB1C3_SDM_SspI /µl | volume of pSB1C3_SDm_PvuII /µl |
| DNA | 4,66 | 4,87 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| Enzyme ClaI (no.Lab:152) | 1 | 1 |
| Enzyme BpmI (no.Lab:144) | 1 | 1 |
| H2O | 9,34 | 9,13 |
| Total volume (e.g. 15,20,25,30 µl) | 20 | 20 |
incubation time: 1,5h
0,5 g Agarose,50 ml TBE (1%), 3 µl GELRED (gel was shared with Anna) , at Volt, running time:
Loading plan for agarose gel:
Marker used: GeneRuler ladder mix (Fermentas)
| Marker | Sample 125, 20µl | Sample 129, 20µl | |
|---|---|---|---|
| Lane | 1 | 3 | 5 |
This appproach also didn't work, same result. the guess is that BpmI doesn't work anymore. will be repeated by Chris W. on 05.08.10. the result will show that the guess is right and BpmI from labstock is changed out with a new one.
Test transformation of XL1 competent cells
Investigator: Kira
Test transformation was performed in order to test the efficiency of produced chemical competent XL1 blue cells.
2 plates were prepared: one with 50 pg and the second with 100 pg pUC 18 plasmid (50 pg/㎕) 50 pg plate: 1 ㎕ pUC + 50 ㎕ XL1B cells 100 pg plate: 2 ㎕ pUC + 50 ㎕ XL1B cells
Transformation was performed according to the standard protocol and the plates were incubated at 37 C.
Sequencing results of ITRs
Investigator: Hanna
8 sequencing files were analyzed "per hand" base by base. Alignments (will be inserted soon) delivered that the right ITR is 100% OK and was successfully converted into the RFC10 BioBrick standard.
The sequencing and alignments of the left ITR delivered that a 15 bp fragment in the middle of the sequence is lacking. Therefore further test digestions have to be performed. Nevertheless also this ITR was successfully converted into the RFC10 BioBrick standard. Secondary structure analysis showed that the stemloop-structure will be nevertheless forming. We will try to test whether the referring fragment is also missing in the pAAV_MCS. If it's also lacking there we will continue with this ITR and test whether it functions.
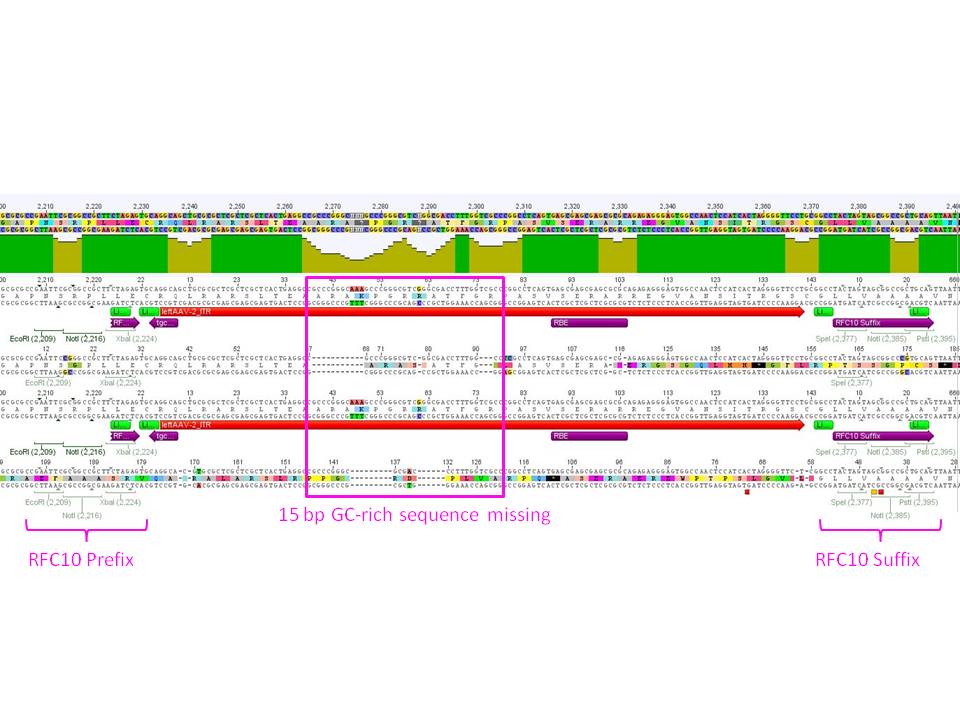
Cloning of Rep68, Rep78 and AAP into pSB1C3
Investigator: Anna
Comments: The PCR of Rep 40/52 didn't work (see 03.08), whereas the PCR of Rep 68/78 and AAP was succesful. Ligation was done with two different samples of pSB1C3 (see agarose gel). Samples from digestion and from ligation are stored in the 4°C freezer.
- Digestion of PCR products and vector:
| components | PCR product /µl | vector /µl |
| DNA | 19 | 10 |
| BSA (10x) | 3 | 3 |
| Buffer 4 (10x) | 3 | 3 |
| Enzyme XbaI | 1 | 1 |
| Enzyme SpeI | 1 | 1 |
| H2O | 3 | 12 |
| Total volume (e.g. 15,20,25,30 µl) | 30 | 30 |
Comments: Digestion was done with XbaI and SpeI, it has to be checked if the inserts are cloned into the vector in the right orientation.
- Purification of Rep68, 78 and AAP:
For the purification 95 µl of buffer PBI was used.
c(Rep68)= 17,6 ng/µl
c(Rep78)= 80,60 ng/µl
c(AAP)= 77,78 ng/µl
- Gelextraction of pSB1C3_RFC25_CFP:
0,5 g Agarose,50 ml TBE (1%), 3 µl GELRED , at 115 Volt, running time:55
5µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas)
c(pSB1C3)= 2,87 ng/µl
c(pSB1C3_2)= 9,46 ng/µl
- Quickligation of PCR products and vector:
For the Ligation 10µl buffer (2x) and 1µl Quickligase were used.
| Sample-no. | vector /µl | insert /µl | |
| pSB1C3 + Rep68 | 1.1 | 6,51 | 2,4 |
| pSB1C3_2 + Rep68 | 1.2 | 3,9 | 5,02 |
| pSB1C32 + Rep78 | 2.1 | 8,2 | 0,8 |
| pSB1C3_2 + Rep78 | 2.2 | 6,82 | 2,18 |
| pSB1C3 + AAP | 3.1 | 8,71 | 0,92 |
| pSB1C3_2 + AAP | 3.2 | 8,1 | 0,9 |
| Total volume (e.g. 15,20,25,30 µl) | 9 |
- Transformation:
The transformation was done following the standard protocol using B21 cells.
80.Labortag 05.08.2010
New LB Agar was prepared.
Investigator: Patrick
Sequenc analysis of BioBricks: CMV, hGH and beta globin
Investigator: Bea
Comments: Sequence analysis of three BioBricks which were cloned into the iGEM standard plasmid pSB1C3. Cloning of this three plasmids were performed at:
Sequencing results of pSB1C3_CMV:
- Plasmid sent for sequencing:
- Primer used: VR-2
- Results: Sequence read looks good. The incorporation of the PCR product can be confirmed. It can be seen that the insert is in the RFC10 standard and prefix and suffix have the right sequence. Therefore, this BioBrick can be send to the registry and used for BioBrick assembly.But: there is a mutation in the termintor (sequence picture do not show this mutation)
<img src="http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/images/7/74/Freiburg10_Sequence_analysis_of_PSB1C3_CMV.jpg" />
Sequencing results of pSB1C3_hGH:
- Plasmid sent for sequencing:
- Primer used: VR-2
- Results: Sequence read looks good. The incorporation of the PCR product can be confirmed. It can be seen that the insert is in the RFC10 standard and prefix and suffix have the right sequence. Therefore, this BioBrick can be send to the registry and used for BioBrick assembly.
http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/images/e/e1/Freiburg10_Sequence_analysis_of_PSB1C3_betaglobin.jpg
Sequencing results of pSB1C3_hGH:
- Plasmid sent for sequencing:
- Primer used: VR-2
- Results: Sequence read looks good. The incorporation of the PCR product can be confirmed. It can be seen that the insert is in the RFC10 standard and prefix and suffix have the right sequence. Therefore, this BioBrick can be send to the registry and used for BioBrick assembly.
http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/images/4/44/Freiburg10_Sequence_analysis_of_PSB1C3_hGH.jpg
Cellculture
Investigator: Adrian
The Motivation: Transfection
The Action: HEK cells were split into four T75 flasks
The Plan: The cells should be ready at saturday for seeding and monday for transfection
We recived new HT1080 cells, they'll be split tomorrow (thx to Sven)
We recived new tumor cells which overexpress EGFR (thx to barbara)
Picking clones of pSB1C3_AAP_2, pSB1C3_AAP, pSB1C3_Rep78_2, pSB1C3_Rep78, pSB1C3_Rep68_2 and pSB1C3_Rep68
Investigator: Anissa
Of each construct two clones were picked. plates are still stored in the cold-room, tomorrow mini-prep will be done.
PCR and ligation for biobrick-production of pSB1C3_hTERT
Investigator: Anissa,Kerstin
Comments: PCR was performed one time without DMSO and one time with DMSO. Only the approach with DMSO showed bands in the analytic gel after PCR. That's why only this approach will be noted.
- PCR:
(was performed following the standard protocol)
| Ingredients | Volume / µl |
| 5X Phusion HF buffer | 10 |
| 10 mM dNTP mix | 1 |
| forward primer: O111 (phTERT_prefix_for_RFC10) | 2,5 |
| reverse primer: O112 (phTERT_suffix_rev_RFC10) | 2,5 |
| DNA Template | 0,35 µl |
| DMSO (2%) | 1 |
| Phusion Polymerase | 0,5 |
| H2O | 32,15 |
| Total volume | 50 |
PCR program:
| PCR Program | temperature/ °C |
| 1 | 98 |
| 2 | 98 |
| 8x | 58 |
| 3 | 72 |
| 4 | 98 |
| 17x | 70 |
| 5 | 72 |
| 6x | 72 |
| Hold | 4 |
Gel extraction
Gel measurement:
| Sample | Weight | Volume | Concentration |
| hTERT | 0,15g | 20 µl | 37,49 ng/µl |
| pSB1C3_cut | 0,08g | 20 µl | 4,56 ng/µl |
- Digestion of PCR product and vector:
| components | PCR product /µl | vector /µl |
| DNA | 18 | 7,58 |
| BSA (10x) | 0,3 (100X used) | 2 |
| Buffer 4 (10x) | 2,5 | 2 |
| Enzyme XbaI | 1 | 1 |
| Enzyme PstI | 1 | 1 |
| H2O | 2,2 | 6,42 |
| Total volume (e.g. 15,20,25,30 µl) | 25 | 20 |
Ligation with T4-Ligase and transformation with BL 21 was made. Clones have to be picked tomorrow.
Transformation evaluation of XL1B cells and repetition of transformation
Investigator: Kira
Against our expectations both agar plates contain very few colonies. In order to figure out if the lack of colonies is due to the XL1 blue cells or loss of pUC activity, test transformation will be repeated this evening with recently prepared XL1 blue cells as well as with Lab-XL1B cells.
50 pg pUC-plate contains 8 colonies
100 pg pUC-plate contains 18 colonies
Transformation will be performed again with 100 pg pUC (= 2 ㎕ pUC).
Cloning of SDM SspI and SDM PvuII
Investigator: Chris W.
- Vector: name: pSB1C3_SDM_SspI P125
- Insert: name: pSB1C3_SDM_PvuII P129
- new vector name: pSB1c3_SDM_SspI/PvuII P157
- buffer used: 4 ; Restriction-enzymes used: Enzyme 1 (no. Lab: 144) BpmI ; Enzyme 2 (no.Lab: 152) ClaI
- DNA concentration (vector): 321,9 ng/µl ; DNA concentration (insert): 308,4 µg/µl
| components | volume of pSB1C3_SDM_SspI /µl | volume of pSB1C3_SDm_PvuII /µl |
| DNA | 4,66 | 4,87 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| Enzyme BpmI (no.Lab:144) | 1 | 1 |
| Enzyme ClaI (no.Lab:152) | 1 | 1 |
| H2O | 9,34 | 9,13 |
| Total volume (e.g. 15,20,25,30 µl) | 20 | 20 |
Comment: I guess enzyme 144 was BpmI and enzyme 152 was ClaI ? (Jessica)
Comment: u r right, changed (Chris)
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 130 Volt, running time:45
Loading plan for agarose gel:
Marker used: GeneRuler ladder mix (Fermentas)
| Marker | Sample 125, 20µl | Sample 129, 20µl | |
|---|---|---|---|
| Lane | 1 | 5 | 7 |
The mutual vector (now without SspI) and insert (now without PvuII) were cut out. The gelextraction was performed according to the standard protocol. DNA concentration of the extracts:
- P125 Sspl: 12,8 ng/µl
- P129 PvuII: 11,6 ng/µl
The Ligation was performed as following:
- Vector Volume: 2,81 µl
- Insert Volume: 5,19 µl
- 1µl T4-Ligase buffer (10x)
- 8µl (Vector + Insert) mix
- 1µl T4-Ligase
Incubate for 30min
Transformation: performed according to the standard protocol (BL21). The cells were plated on a agar plate with chloramphenicol
Miniprep of p158 and p159
Plasmid Mini-Prep
Investigator: Bea & Volker
For the tripple mutant of pAAV-RC a Quick-change reaction was carried out to remove the last restriction site (SalI) that is required for the Viral Brick standard. For this construct glycerol stocks and minipreps were carried out for two clones.
Glycerol Stocks
| Clone 1 | Clone 2 | |
| Bacteria strain | BL-21 | BL-21 |
| Plasmidname | pAAV_RC_1.2 SDM SalI | align="left"| pAAV_RC_1.2 SDM SalI |
| Date | 05.08.2010 | 05.08.2010 |
| given glycerol-stock no. | B126 | B127 |
| given plasmid no. | p158 | p159 |
TO do: Test digestion and sequencing!!!
81.Labortag 06.08.2010
Cloning Rep40 & Rep52 into pSB1C3
Intention: get biobrick ready
Investigator: Patrick
PCR of pKEX-2XL.Rep 40 (P22) and pKEX-2XL.Rep 52 (P23) following gelrun (1%) and gelextraction.
Used Primers:
- Praefix Rep40_52ex (O94)
- Suffix Rep 40_68ex (O96)
- Suffix Rep 52_78ex (O97)
PCR programm of Rep40:
- 98°C 1 min
- 98°C 15 sec
- 63°C 25 sec
- 72°C 15 sec Repeat this cycle 8 times.
- 98°C 15 sec
- 68°C 25 sec
- 72°C 15 sec Repeat this cycle 17 times.
- 72°C 5 min
- Hold 4°C
PCR programm of Rep52:
- 98°C 1 min
- 98°C 15 sec
- 62°C 25 sec
- 72°C 20 sec Repeat this cycle 8 times.
- 98°C 15 sec
- 66°C 25 sec
- 72°C 20 sec Repeat this cycle 17 times.
- 72°C 5 min
- Hold 4°C
| ingredients | Rep 40 | Rep40 + DMSO | Rep52 | Rep52 + DMSO |
| 5x Phusion HF buffer | 10 µl | 10 µl | 10 µl | 10 µl |
| 10 mM dNTP mix | 1 µl | 1 µl | 1 µl | 1 µl |
| for Primer O94 (1:10 dilution, 05 µM) | 2,5 µl | 2,5 µl | 2,5 µl | 2,5 µl |
| rev Primer (1:10 dilution, 0,5 µM) | 2,5 µl O96 | 2,5 µl O96 | 2,5 µl O97 | 2,5 µl O97 |
| DNA template | 1 µl, 112,7 ng/µl | 1 µl, 112,7 ng/µl | 0,9 µl, 142,3 ng/µl | 0,9 µl, 142,3 ng/µl |
| DMSO | 0 µl | 0,5 µl | 0 µl | 0,5 µl |
| Phusion Polymerase | 0,5 µl | 0,5 µl | 0,5 µl | 0,5 µl |
| H2O | 32,5 µl | 32 µl | 32,6 µl | 32,1 µl |
| Total volume | 50 µl | 50 µl | 50 µl | 50 µl |
Ecpected size of Rep40: 940 bp, expected size of Rep52:1198 bp. There are two samples of Rep40 and Rep52 and the PCR of one of each was run with 1% DMSO because the Rep40 Praefix has a strong secondary structure and was used as a primer for Rep 40 and Rep 52

Digestion of pSB1C3 (P51.1) with EcoRI-HF and SpeI following gelrun (0.8%) and gelextraction.
Expected size of the fragments: about 2000 bp and 800bp

All marked fragments were extracted. The Gelextraction was performed according to the standard protocol following a digestion of the PCR products with EcoRI HF and SpeI and a purification of the PCR product. The Ligation was not performed according to the standard protocol: 1 µl T4 DNA Ligase, 1 µl (10x) Buffer, 3 µl vector pSB1C3 (60 ng/µl) and 5 µl insert (all concentrations about 100 ng/µl). The Transformation (with BL21) was performed according to the standard protocol. Tomorrow there will hopefully be some clones on the agar plates.
BioBrick Assembly of pSB1C3_beta globin_mVenus
Investigator: Bea
Comments: BioBricks are ready to use. The sequenced pSB1C3_beta globin (P133) and pGA14_mVenus (P60) will be digested with different enzymes and will be assembled in order to produce the first step in assembling the whole vector together.
- First the plasmids P133 and P60 were digested:
| pSB1C3_betaglobin/µL | pGA14_mVenus/µL | |
| DNA | 7 | 9 |
| BSA (100x) | 0,2 | 0,2 |
| Buffer 4 (10x) | 2 | 2 |
| Enzyme XbaI | 1 | 1 |
| Enzyme AgeI | 1 | 1 |
| H2O | 8,8 | 6,8 |
| Total volume | 20 | 20 |
- Incubation of plasmids at 37°C for 2 hours
- Load samples on preparative 1% agarose gel and run at 110 V, 45 minutes
Results: As it can be seen in the picture below, digestion of the vector pSB1C3_betaglobin (left lane) revealed a band at around 2500 - 3000 bp. The expected size after digetsing with SpeI and PstI was: 2555 bp. The band ran a little bit higher than expected, anyhow the band was cut out of the gel. The right lane belongs to the pGA14_mVenus which was digested with XbaI and PstI which resulted in two fragments. mVENUS was digested. Expected sizes were: mVenus = 756bp and pGA14 = 2900bp. The expected sizes correspond to the bands seen in the picture below. The smaller fragment belonging to mVenus was cut out of the gel as well.

After gel extraction has been performed, the concentrations of the two fragments were measured and we obtained following concentrations:
- c(pSB1C3_betaglobin) = 15,62 ng/µL
- c(mVenus) = 6,83 ng/µL
For ligation the T4-Ligase were used.
- v(pSB1C3_betaglobin) = 2,64 µL
- v(mVenus) =5,36 µL
- v(10xT4 Ligase buffer) = 1 µL
- v(T4-Ligase) = 1 µL
- Total volume = 10 µL
Additionally a control ligation was performed.
- v(pSB1C3_betaglobin) = 2,64 µL
- v(H20) =5,36 µL
- v(10xT4 Ligase buffer) = 1 µL
- v(T4-Ligase) = 1 µL
- Total volume = 10 µL
- Ligation duration was 45 minutes on room temperature.
After ligation a transformation was performed.
- Used bacterial strain: XL1-B
- LB-agar plates containing chloramphenicol were plated and put in the 37°C room over night
To do: Mini-Prep and test digestion in order to verify assembly of beta globin and YFP. After verification of the fusion of the two fragments this construct is ready for the next BioBrick assembly step.
Test digestion of SalI SDM
Investigator: Hanna
Test digestion
- buffer used: 3 ; Restriction-enzymes used: Enzyme 1 (no. Lab: 53): SalI ; Enzyme 2: XbaI
- Plasmid
- Given Plasmid-Number: P158; DNA concentration: 398.8 ng/µL ;
- Given Plasmid-Number: P159; DNA concentration: 437.4 ng/µL ;
| Components | P158 Volume/µL | P159 Volume/µL |
| DNA | 2.5 | 2.3 |
| BSA (10x) | 1 | 1 |
| Buffer 3 (10x) | 1 | 1 |
| SalI (no. Lab: 53) | 0.5 | 0.5 |
| XbaI | 0.75 | 0.75 |
| H2O | 4.25 | 4.45 |
| Total volume | 10 | 10 |
Incubation: 1 h
Agarose-Gel:
0.5 g Agarose, 50 mL TBE (1 %),3 µL GELRED, at 115 Volt, running time: 45 minutes
| Sample | Sample/µl] | Loading dye (5x)/µl | Expected size 1 (if SDM didn't work) | Expected size 2 (if SDM didn't work) | Expected size 3 (if SDM didn't work) |
|---|---|---|---|---|---|
| P158 | 10 µl | 2 µl | 6129 bp | 1143 bp | 61 bp |
| P159 | 10 µl | 2 µl | 6129 bp | 1143 bp | 61 bp |
- Marker: GeneRuler ladder mix
| Marker /µL | Sample P158 /µl | Sample P159 /µl | Marker /µL | |
|---|---|---|---|---|
| Lane | 6.5 | 10 | 10 | 7 |
Comments: Test digestion looked good: No 1143 bp fragment was detectable in neither test digestion.
P158 was sent for sequencing to GATC.
ITR test digestion of pAAV_MCS
Investigator: Hanna
Comment: Because sequencing of the right ITR delivered that perhaps a 15 bp fragment is missing in the sequence, we wanted to check, whether this fragment is also not present in the original pAAV_MCS plasmid or whether the loss of these 15 bp happened during cloning.
Test digestion
- buffer used: 4 ; Restriction-enzymes used: Enzyme 1: NotI; Enzyme 2: PstI
- Plasmid
- Plasmid-Number: from Sven; DNA concentration: 259 ng/µL ;
| Components | Volume/µL |
| DNA | 3.8 |
| BSA (10x) | 1.5 |
| Buffer 4 (10x) | 1.5 |
| NotI-HF | 0.75 |
| PstI-HF | 0.75 |
| H2O | 3.2 |
| Total volume | 15 |
Incubation: 1 h
Agarose-Gel:
0.5 g Agarose, 50 mL TBE (1 %),3 µL GELRED, at 115 Volt, running time: 45 minutes
| Sample | Sample/µl] | Loading dye (5x)/µl | Expected size ITRs | Expected size 2 | Expected size 3 | Expected size 4 |
|---|---|---|---|---|---|---|
| pAAV_MCS | 15 µl | 2 µl | 145 bp | 1216 bp | 555 bp | 2608 bp |
- Marker: GeneRuler ladder mix
| Marker /µL | Sample /µl | Marker /µL | |
|---|---|---|---|
| Lane | 6.5 | 12 | 6.5 |
Comments: The test digestion showed that there're two bands between the 100 and 200 bp marker nucleotides! By comparing the gel picture with the last mini-prep digestion of the fancy method we found out that the space between the two bands are similar. Because of that we assumed that the 15 pb are already missing in the original Stratagene plasmid!!!
Conclusion: Because we already tested the Stratagene Kit via YFP expression, we can conclude that the usage of the "wrong" rigth ITR works. Therefore we decided to continue working with this "mutated" = ENGINEERED :) right ITR as BioBrick.
Miniprep of pSBC13_Rep78, pSB1C3_Rep68, pSB1C3_AAP, pSB1C3_rightITR and psB1C3_leftITR
Plasmid Mini-Prep
Investigator: Kerstin
Glycerol Stocks
pSB1C3_Rep78
| Clone 1 | Clone 2 | |
| Bacteria strain | BL-21 | BL-21 |
| given glycerol-stock no. | B128 | B129 |
| given plasmid no. | P160 | p161 |
| DNA-concentration ng/µl | 139,1 | 71,0 |
pSB1C3_2_Rep78
| Clone 1 | Clone 2 | |
| Bacteria strain | BL-21 | BL-21 |
| given glycerol-stock no. | B130 | B131 |
| given plasmid no. | P162 | p163 |
| DNA-concentration ng/µl | 112,1 | 79,7 |
pSB1C3_Rep68
| Clone 1 | Clone 2 | |
| Bacteria strain | BL-21 | BL-21 |
| given glycerol-stock no. | B132 | B133 |
| given plasmid no. | P164 | p165 |
| DNA-concentration ng/µl | 80,2 | 72,8 |
pSB1C3_2_Rep68
| Clone 1 | Clone 2 | |
| Bacteria strain | BL-21 | BL-21 |
| given glycerol-stock no. | B134 | B135 |
| given plasmid no. | P166 | p167 |
| DNA-concentration ng/µl | 85,6 | 69,1 |
pSB1C3_AAP
| Clone 1 | Clone 2 | |
| Bacteria strain | BL-21 | BL-21 |
| given glycerol-stock no. | B136 | B137 |
| given plasmid no. | P168 | p169 |
| DNA-concentration ng/µl | 63,6 | 56,3 |
pSB1C3_2_AAP
| Clone 1 | Clone 2 | |
| Bacteria strain | BL-21 | BL-21 |
| given glycerol-stock no. | B138 | B139 |
| given plasmid no. | P170 | p171 |
| DNA-concentration ng/µl | 135,1 | 114,2 |
pSB1C3_rightITR
| Clone 1 | Clone 2 | |
| Bacteria strain | BL-21 | BL-21 |
| given glycerol-stock no. | B140 | B141 |
| given plasmid no. | P172 | p173 |
| DNA-concentration ng/µl | 73,7 | 66,4 |
pSB1C3_leftITR
| Clone 1 | Clone 2 | |
| Bacteria strain | BL-21 | BL-21 |
| given glycerol-stock no. | B142 | B143 |
| given plasmid no. | P174 | p175 |
| DNA-concentration ng/µl | 74,9 | 80,3 |
Test digestion:
pSB1C3_Rep78, pSB1C3_Rep68 and pSB1C3_AAP were digested with XbaI and SpeI
pSB1C3_rightITR and pSB1C3_leftITR were digested with EcoRI and PstI.
| components | volume of P160/µl | volume of P161/µl | volume of P162/µl | volume of P163/µl | volume of P164/µl | volume of P165/µl | volume of P166µl | volume of P167/µl | volume of P168µl | volume of P169/µl | volume of P170/µl | volume of P171/µl | volume of P172/µl | volume of P173µl | volume of P174/µl | volume of P175/µl |
| DNA | 5,8 | 11,3 | 7,1 | 10 | 9,9 | 10,9 | 9,3 | 11,6 | 12,5 | 14,2 | 5,9 | 7,0 | 10,9 | 12,0 | 10,7 | 9,7 |
| BSA (10x) | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| Enzyme 1 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 |
| Enzyme 2 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 |
| H2O | 9,2 | 3,7 | 7,9 | 5,0 | 5,1 | 4,1 | 5,7 | 3,4 | 2,5 | 0,8 | 9,1 | 8 | 4,1 | 3 | 4,3 | 5,3 |
| Total volume /µl | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 | 20 |
Expected sizes:
- Rep78: 1,8 kb
- Rep68: 1,6kb
- AAP: 650bp
- left ITR: 150bp
- right ITR: 150bp
Results of samples loaded 1,5% agarose gel and ran ~50 minutes at 115 V:
Results of samples loaded 0,8% agarose gel and ran ~50 minutes at 115 V:
Sequencing
P160, P165 and P170 were send for sequencing.
82.Labortag 07.08.2010
Cellculture
HEK 293 cells were split and seeded according to the standard protocol: 2x T75 flask and 10x10cm cellculture dishes. Investigator: Patrick
Continuation: Cloning Rep40 & Rep52 into pSB1C3
Investigator: Patrick
Results: no clones could be picked because there were no clones. Maybe the PCR purification or one of the following steps did not work (the PCR purifications were quite bad) so i purified the PCR products again yielding very low DNA concentrations (0-13,28 ng/µl) and still very obvious pollution of the sample. Nonetheless a ligation with the insert Rep40+DMSO, Rep52, Rep52+DMSO and the vector pSB1C3 was carried out following a transformation and according to the standard protocols. Hopefully there will be some clones tomorrow.
Mini-Prep and test digestion of pSB1C3_SDM_SspI/PvuII
Investigator: Jessica
Plasmid Mini-Prep
- new vector name: pSB1C3_SDM_SspI/PvuII
Glycerol Stocks
pSB1C3_SDM_SspI/PvuII
| Clone 1 | Clone 2 | Clone 3 | |
| Bacteria strain | BL21 | BL21 | BL21 |
| Plasmidname | pSb1C3_SDM_SspI/PvuII | pSb1C3_SDM_SspI/PvuII | pSb1C3_SDM_SspI/PvuII |
| Date | 7.8.10 | 7.8.10 | 7.8.10 |
| given glycerol-stock no. | B144 | B145 | B146 |
| given plasmid no. | P157.1 | P157.2 | P157.3 |
Test digestion
- buffer used: 2 ; Restriction-enzymes used: Enzyme 1 SspI ; Enzyme 2 PvuII
- Plasmid: pSB1C3_SDM_SspI/PvuII:
- Given Plasmid-Number: P157.1; DNA concentration: 246,3 ng/µL ;
- Given Plasmid-Number: P157.2; DNA concentration: 231,7 ng/µL ;
- Given Plasmid-Number: P157.3; DNA concentration: 219,6 ng/µL ;
| Components | P157.1 Volume/µL | P157.2 Volume/µL | P157.3 Volume/µL |
| DNA | 2,8 | 3,0 | 3,19 |
| BSA (10x) | - | - | - |
| Buffer no. 2 (10x) | 1 | 1 | 1 |
| Enzyme 1 SspI | 0.5 | 0.5 | 0.5 |
| Enzyme 2 PvuII | 0.5 | 0.5 | 0.5 |
| H2O | 5,2 | 5 | 4,81 |
| Total volume | 10 | 10 | 10 |
- Incubation: 45 min
Agarose-Gel:
0.45 g Agarose, 50 mL TAE, 3 µL GELRED, at 115 Volt, running time: 45 minutes
| Sample | Sample/µl] | Loading dye (6x)/µl |
|---|---|---|
| P157.1 | 10 µl | 1,6 µl |
| P157.2 | 10 µl | 1,6 µl |
| P157.3 | 10 µl | 1,6 µl |
- Marker: GeneRuler ladder mix
| Marker /µL | Sample P157.1 /µL | Sample P157.2 /µL | Sample P157.3 /µL | |
|---|---|---|---|---|
| Lane | 7 | 10 | 10 | 10 |
I expect one cut from the PvuII RS in the CFP --> linearized
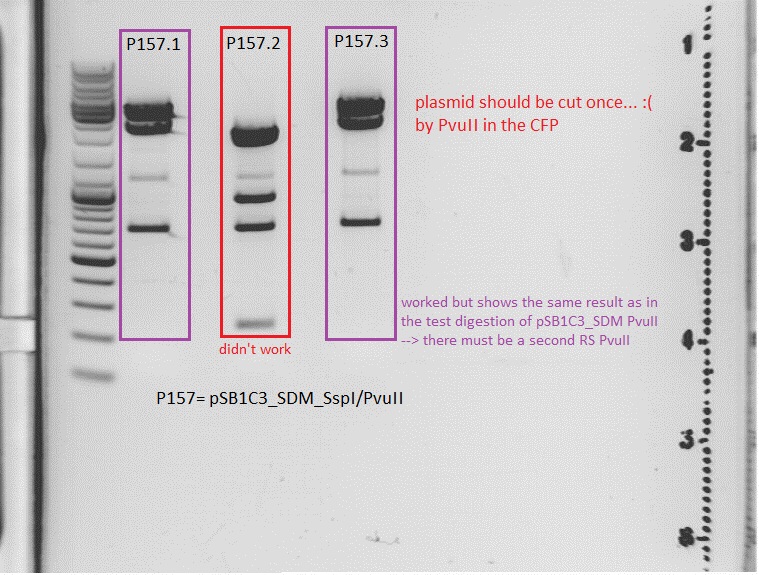
Comment:
Comments:Oh f... it makes no sense... if the cloning didn't work, there can just be 3 bands (one time SspI and twice PvuII). The results of the test digestion from aug 1th showed that there can be one RS PvuII more. but then you have a maximum of 4 bands. P157.2 show 5 bands. i can't interpret this result at the moment. interpretation will follow
Evaluation of test transformation with XL1 blue cells and pUCI
Investigator: Kira
Transformation was performed with lab XL1B and iGEM XL1B cells. iGEM plate contained just 25 colonies, while lab plate contained around 50 colonies. Conclusion: the production of competent cells has to be repeated.
Biobrick production of p5 promoter from pTAV2 and pAAV_RC
Investigator: Kira
- PCR:
DNA samples were diluted 1:100
| Ingredients | p 32 | p 50 |
| 5X Phusion HF buffer | 10 µl | 10 µl |
| 10 mM dNTP mix | 1µl | 1 µl |
| forward primer: O114 | 2,5µl | 2,5 µl |
| reverse primer: O115 | 2,5 µl | 2,5 µl |
| DNA Template | 0,3 µl | 0,4 µl |
| DMSO | 0 µl | 0 µl |
| Phusion Polymerase | 0,5 µl | 0,5 µl |
| H2O | 33,2 µl | 28,6 µl |
| Total volume | 50 µl | 50 µl |
PCR program:
| Cycles | Temperature | Time |
| 98°C | 1 | |
| 8x | 98°C | 30" |
| 60°C | 25" | |
| 72°C | 6" | |
| 17x | 98°C | 30" |
| 70°C | 25" | |
| 72°C | 6" | |
| 1x | 72°C | 5' |
| Hold 4°C |
Digestion of plasmid backbone:
c (pSB1C3) = 151, 1 ng/ µl
| Components | vector Volume/µL |
| DNA 1 µg | 6,0 µl |
| BSA (100x) | 0,2 µl |
| Buffer no. 4 (10x) | 2,0 µl |
| Enzyme 1 XbaI | 1 µl |
| Enzyme 2 PstI HF | 1 µl |
| H2O | 9,8 µl |
| Total volume | 20 |
incubation @ 37 C for approx. 2 h
1% agarose gel with PCR products:
Concentrations after gel extraction:
c(p32) = 67,95 ng/µl
c(p50) = 32,45 ng/µl
c(vector) = 7,54 ng/µl
Digestion of PCR products:
| Components | PCR product Volume/µL |
| DNA 1 µg | 23,0 µl |
| BSA (100x) | 0,3 µl |
| Buffer no. 4 (10x) | 3,0 µl |
| Enzyme 1 XbaI | 1,5 µl |
| Enzyme 2 PstI HF | 1,5 µl |
| H2O | 0,7 µl |
| Total volume | 30 |
Concentrations after PCR purification:
c(p32) = 21, 63 ng/µl
c(p50) = 19,48 ng/µl
Ligation:
Quickligase was used for ligation.
p32 DNA-Mix: 8 µl vector + 1 µl insert
p50 DNA-Mix: 7,9 µl vector + 1,1 µl insert
Transformation:
Transformation was performed according to the standard protocol with DYT and BL21 cells. The cells were spread on the agar plates containing chloramphenicol.
Sequencing results of SalI SDM
Investigator: Hanna
Comment: The test digestion already showed that the SDM of the SalI restriction site in pAAV_RC worked. This could be validated by sequencing the plasmid with the Rep-1250_for primer:
P58 can be used for further RepCap-experiments.
83. Labortag 08.08.2010
HT 1080 cells were split and seeded: 3x10^6 cells per into each well of a 6-well plate.
A431 cells were washed and received new medium.
There were no clones on the agar plates so they were put back into the 37°C room. Maybe there will be some clones tomorrow.
84. Labortag 09.08.2010
Midi-Prep Inoculation
- pAAV_RC
- pAAV_RC_1.2 SDM SalI (P158)
- pAAV_RC_mGMK_TK30 clone 1 (P81)
- pAAV_RC_mGMK_TK30 clone 2 (P82)
Investigators: Chris W., Patrick
Continuation: Cloning Rep40 & Rep52 into pSB1C3
Investigator: Patrick
Two Rep52+DMSO clones grew. They were picked and tomorrow there will be a midi prep and a test digestion.
Simultaneously a new approach for cloning Rep40 and Rep52 into pSB1C3 was performed. (see next header)
Repetition: BioBrick production of Rep40 & Rep52
Investigator: Anna
Comments: Another approach was done because there were no clones for Rep 40 and only two for Rep 52 (Cloning see 06.08.2010). PCR was performed again with DMSO. Samples from gel extraction are stored in 4°C freezer.
PCR
| Ingredients | Rep 40 | Rep52 |
| 5X Phusion HF buffer | 10 µl | 10 µl |
| 10 mM dNTP mix | 1µl | 1 µl |
| forward primer: O094 | 2,5µl | 2,5 µl |
| reverse primer: O0096/ O097 | 2,5 µl | 2,5 µl |
| DNA Template (1ng) | 1 µl | 1 µl |
| DMSO | 1 µl | 1 µl |
| Phusion Polymerase | 0,5 µl | 0,5 µl |
| H2O | 31,5 µl | 31,5 µl |
| Total volume | 50 µl | 50 µl |
PCR programm
| PCR Program | temperature/ °C | Time | PCR products |
| 1 | 98 | 1min | |
| 2 | 98 | 15s | |
| 8x | *** | 15s | 63°C |
| 3 | 72 | *** | 15s |
| 4 | 98 | 15s | |
| 19x | *** | 15s | 68°C |
| 5 | 72 | *** | 15s |
| 6x | 72 | 5min | |
| Hold | 4 |
Agarose gel
0.5 g Agarose, 50 mL TAE (1 %),3 µL GELRED, at 120 Volt, running time: 45 minutes
| Sample | Sample / µl] | Loading dye (6x) / µl | Expected size |
|---|---|---|---|
| p22 (Rep40) | 44 µl | 7 µl | 940 bp |
| p23 (Rep52) | 44 µl | 7 µl | ~1200 bp |
- Marker: GeneRuler ladder mix
| Marker /µL | Sample Rep40 /µl | Sample Rep52 /µl | |
|---|---|---|---|
| Lane | 7 | 51 | 51 |
Gel extraction
PCR products were eluted with 20 µl BE puffer following the standard protocol.
c(Rep40)= 28,03 ng/µl
c(Rep52)= 121,16 ng/µl
Comments: Digestion was done with the wrong enzymes (EcoRI HF and SpeI), it will be repeated on 11.08. with XbaI and SpeI
Sequence analysis of pSB1C3_mGMK
Investigator: Bea
Comments: mGMK was cloned into the pSB1C3 backbone in order to send it to the parts registry!. The sequence analysis revealed that the mGMK was successfully cloned into pSB1C3 plasmid. This polasmid is in the RFC25 standard.
-
Results sequencing: Sequence read ok!
Primer used: VR-2
Comment: A mutation has been found in the pSB1C3 backone which was found generally in the backbone which we received from the iGEM HQ.
Mini-Prep of pSB1C3_hTERT, pSB1C3_beta-globin_mVenus, pSB1C3_pTAV2(P5), pSB1C3_pAAV_RC(P5TATAles)
Investigator: Jessica
Bold text
Mini-Prep following the standart Protokoll
Test digestion:
- Perfomed with 10 µL of total volume with all 9 clones
| Mastermix/µL | P176/µL | P177/µL | P178/µL | P179/µL | P180/µL | P181/µL | P182/µL | P183/µL | P184/µL | |
| DNA | - | 4,3 | 2,3 | 3,0 | 2,4 | 6,4 | 5,0 | 4,2 | 4,0 | 4,0 |
| BSA (10x) | 10 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 | 3 |
| Buffer 4 (10x) | 10 | |||||||||
| Enzyme XbaI | 5 | |||||||||
| Enzyme AgeI | 5 | |||||||||
| H2O | - | 2,7 | 4,7 | 4,0 | 4,6 | 0,6 | 2,0 | 2,8 | 3,0 | 3,0 |
| Total volume | 30 | 10 | 10 | 10 | 10 | 10 | 10 | 10 | 10 | 10 |
- All 9 samples were loaded on a 1% agarose gel
- P176 = pSB1C3_hTERT clone1
- P177 = pSB1C3_hTERT clone2
- P178 = pSB1C3_betaglobin clone1
- P179 = pSB1C3_betaglobin clone2
- P180 = pSB1C3_pTAV2(P5) clone1
- P181 = pSB1C3_pTAV2(P5) clone2
- P182 = pSB1C3_pAAV_RC(P5TATAless) clone1
- P183 = pSB1C3_pAAV_RC(P5TATAless) clone2
- P184 = pSB1C3_pAAV_RC(P5TATAless) clone3
| Sample | Sample/µl] | Loading dye (6x)/µl | Expected size 1 (Geneious) |
|---|---|---|---|
| P176 | 10 µl | 1,6 µl | 474 bp |
| P177 | 10 µl | 1,6 µl | 474 bp |
| P178 | 10 µl | 1,6 µl | 509 bp |
| P179 | 10 µl | 1,6 µl | 509 bp |
| P1780 | 10 µl | 1,6 µl | 164 bp |
| P181 | 10 µl | 1,6 µl | 164 bp |
| P182 | 10 µl | 1,6 µl | 164 bp |
| P183 | 10 µl | 1,6 µl | 164 bp |
| P184 | 10 µl | 1,6 µl | 164 bp |
Comments: hTERT and betaglobin_mVenus is inserted in pSB1C3, pTAV(P5) and pAAV_RC (P5TATAless) will be sequenced
Primer: Reverse Primer VR2
Cell culture
Investigators: Adrian, Kerstin, Patrick
Transfection on HEK293
The motivation: first we want to create new stock of AAV without GOI, the next viral stock is with correct TK/GMK, the last stocks are with different YFP-amounts to check if the amount of GOI is a critical step for transduction effency (transgene expression).
| Plate | pAAV_RC/µg | pHelper/µg | GOI/µg |
| 1 | 3,3 | 3,3 | no GOI |
| 2 | 6,6 | 3,3 | no GOI |
| 3 | 3,3 | 3,3 | 3,3 TK_GMK clone1 |
| 4 | 3,3 | 3,3 | 3,3 TK_GMK clone1 |
| 5 | 3,3 | 3,3 | 3,3 TK_GMK clone2 |
| 6 | 3,3 | 3,3 | 3,3 YFP |
| 7 | 3,3 | 3,3 | 10 YFP |
| 8 | 3,3 | 3,3 | 20 YFP |
| 9 | 3,3 | 3,3 | 40 YFP |
| 10 | 3,3 | 3,3 | 60 YFP |
Transduction of HT1080
Motivation
The Motivation: checking survivalrate of Cells, with either no Virus, virus without GOI and virus with YFP (10µg stock)
The Plan:
Plate I
| control, no cells | no GOI 150µl | 10µg 500µl| |
| control, no virus | no GOI 300µl | 10µg 750µl| |
Plate II
| control, no cells | no GOI 150µl | 10µg 500µl| |
| control, no virus | no GOI 300µl | 10µg 750µl| |
Plate III
| control, no cells | no GOI 150µl | 10µg 500µl| |
| control, no virus | no virus | 10µg 750µl| |
Passaging and seeding of A431
one 6er dish was prepared for making pictures and one T75 Flask.
BioBrick Assembly of pSB1C3_leftITR_CMV and pSB1C3_hGH_rightITR
Investigator: Bea, Achim, Jo
Comments: In order to proceed with the BioBrick assembly the pasmids pSB1C3_left ITR and pSB1C3_CMV were used and pSB1C3_rigthITR and pSB1C3_hGH were used in order to obatin plasmids with both sequences.
Digestion:
| components | pSB1C3_left ITR | pSB1C3_CMV | pSB1C3_right ITR | pSB1C3_hGH |
| DNA | 13,4 | 9,5 | 13,6 | 6,6 |
| BSA (10x) | 2 | 2 | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 | 2 | 2 |
| Enzyme1 | 1 (SpeI) | 1 (Xba1) | 1 (EcoRI) | 1 (EcoRI) |
| Enzyme2 | 1 (PstI) | 1 (PstI) | 1 (XbaI) | 1 (SpeI) |
| H2O | 0,6 | 4,5 | 0,4 | 7,4 |
| Total volume | 20 | 20 | 20 | 20 |
Preparation of gel:
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time: 45 minutes
Oh, nice picture.. I can recognize many details :)*g* (Hanna)
Expected sizes of constructs:
- pSB1C3_CMV: 650 bp
- pSB1C3_lITR: 2200 bp
- pSB1C3_hGH: 500 bp
- pSB1C3_rITR: 2200 bp
The correspnding bands were cut out and Gel-Extraction was performed according to protocol.
concentrations measured via NanoDrop:
- pSB1C3_CMV: 13,51 ng/µl
- pSB1C3_lITR: 18,49 ng/µl
- pSB1C3_hGH: 7,22 ng/µl
- pSB1C3_rITR: 20,24 ng/µl
Ligation:
For ligation, T4 ligase was used.
- 1µl T4-Ligase buffer (10x)
- 8µl (Vector + Insert) mix
- 1µl T4-Ligase
Ligation was carried out as following:
- Ligation 1:
- pSB1C3_CMV: 4,4 ng/µl
- pSB1C3_lITR: 3,6 ng/µl
- Ligation 2:
- pSB1C3_hGH: 5,25 µl
- pSB1C3_rITR: 2,75 µl
Incubation for 40 minutes.
Transformation:
Transformation carried out according to standard protocol (BL21). Cells were plated on an agar plate with chloramphenicol.
Repetition: Cloning of SDM SspI and SDM PvuII
Investigator: Stefan
Comments: Last week's cloning of SDM_SspI and SDM_PvuII didn't work out, so this is the repetition. In this approach the same restriction enzymes and amounts of DNA were used.
- Vector: name: pSB1C3_SDM_SspI P125
- Insert: name: pSB1C3_SDM_PvuII P129
- buffer used: 4
- Restriction-enzymes used:
- BpmI (no. Lab: 144)
- ClaI (no.Lab: 152)
- DNA concentration (P125): 321,9 ng/µl
- DNA concentration (P129): 308,4 ng/µl
Digestion:
| components | volume of pSB1C3_SDM_SspI /µl | volume of pSB1C3_SDm_PvuII /µl |
| DNA | 4,66 | 4,87 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| BpmI (no.Lab:144) | 1 | 1 |
| ClaI (no.Lab:152) | 1 | 1 |
| H2O | 9,34 | 9,13 |
| Total volume | 20 | 20 |
Gel extraction:
Preparation of gel:
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time: 45 minutes
Expected sizes of constructs:
- pSB1C3_SDM_SspI: 1819 bp
- pSB1C3_SDM_PvuII: 981 bp
The correspnding bands were cut out and Gel-Extraction was performed according to protocol.
concentrations measured via NanoDrop:
- Sspl: 19,85 ng/µl
- PvuII: 10,57 ng/µl
Ligation:
For ligation, T4 ligase was used.
- 1µl T4-Ligase buffer (10x)
- 8µl (Vector + Insert) mix
- 1µl T4-Ligase
The Ligation was performed as following:
- SDM_SspI Volume: 4,03 µl
- SDM_PvuII Volume: 3,97 µl
- 1µl T4-Ligase buffer (10x)
- 8µl (Vector + Insert) mix
- 1µl T4-Ligase
Incubating for 45 minutes.
Transformation:
Transformation performed according to the standard protocol (BL21). The cells were plated on a agar plate with chloramphenicol.
Design and ordering of primer for P40 Promoter BioBrick production
Investigator: Hanna
In order to generate a P40 promoter BioBrick = "Cap-Promoter", oligos were designed and ordered at Sigma Aldrich:
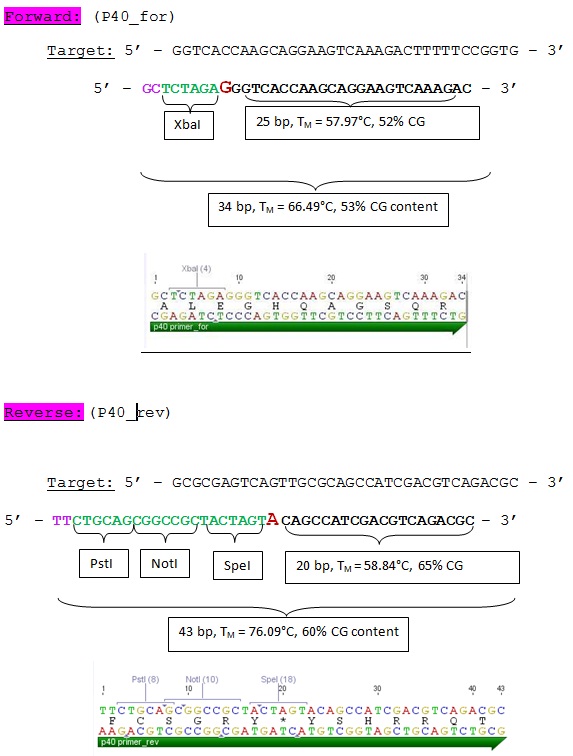
85. labday 10.08.2010
Retrafo with RepCap gene (Mr.Gene)
Investigator: Bea
Comments: We received the rep_cap insert ordered from Mr. Gene today. In order to used and clone the insert into the desired sequences a retrafo has to be performed.
Re-transformation:
- Construct used: pMA_REP_CAP
- We reveived 5 µg (lyophilzied). These were resuspended in 100 µL H2O and vortexted.
- c = 50 ng/µL
- c = Plasmid need to be diluted in order to obatain the final 500pg
- Dilution: 1:100
- 100 µL aliquot of XL1-B were thawed on ice. The proper amount of 1µL was added to the cells.
- Transformed cells were plated on LB plates containing ampicillin
Continuation: Cloning Rep40 & Rep52 into pSB1C3
A midi prep of the two Rep52+DMSO clones was performed yielding concentrations of 261,35 ng/µl (clone 1) and 240,47 ng/µl (clone 2). Unfortunately we noticed today that we should not use EcoRI to clone Rep52 into pSB1C3 because EcoRI cuts two times within the sequence of Rep52.
Midi-Prep of pAAV_RC, pAAV_RC_1.2 SDM SalI (P158), pAAV_RFC25_mGMK_TK30 clone 1 (P81) and 2 (P82)
Investigators: Patrick, Chris
The Midi-Preps were performed according to the standard protocol yielding the following concentrations:
| pAAV_RC | P81a | P81b | P82a | P82b | P158a | P158b |
| 1348,49 ng/µl | 1783,86 ng/µl | 2096,25 ng/µl | 1915 ng/µl | 1469,35 ng/µl | 1834,25 ng/µl | 1771,27 ng/µl |
The annotations a and b were added because two midi-preps of P81, P82 and P158 were carried out.
Results of sequencing pSB1C3_pTAV2(P5) and Psb1C3_pAAV_RC(P5TATAless)
Investigator: Jessica
@Jessi: Which clones are ok, which not?? text does not correspond to pictures! (Bea)
sequencing of P180 was successful, P5 with TATAbox is in pSB1C3
sequencing of P183 was successful but there is a G missing --> plamid is thrown away
sequencing of P184 was successful, P5 is in without TATAbox. this plasmid can be used.
P181 and P180 are thrown away because of the result of the test digestion.
Repetition: BioBrick production of Rep40 and Rep 52
Investigator: Anna
Comments: PCR was repeated once again (that was bad!), because digestion was done with the wrong enzymes (EcoRI HF and SpeI). There are some differences in the PCR programm in comparison to the last (see 09.10.)
PCR
| Ingredients | Rep 40 | Rep52 |
| 5X Phusion HF buffer | 10 µl | 10 µl |
| 10 mM dNTP mix | 1µl | 1 µl |
| forward primer: O094 | 2,5µl | 2,5 µl |
| reverse primer: O0096/ O097 | 2,5 µl | 2,5 µl |
| DNA Template 1:100 (1ng) | 1 µl | 1 µl |
| DMSO | 1 µl | 1 µl |
| Phusion Polymerase | 0,5 µl | 0,5 µl |
| H2O | 31,5 µl | 31,5 µl |
| Total volume | 50 µl | 50 µl |
PCR programm
| PCR Program | temperature/ °C | Time | PCR products |
| 1 | 98 | 1min | |
| 2 | 98 | 15s | |
| 8x | *** | 15s | 62,5°C |
| 3 | 72 | *** | 25s |
| 4 | 98 | 15s | |
| 19x | *** | 15s | 67°C |
| 5 | 72 | *** | 25s |
| 6x | 72 | 5min | |
| Hold | 4 |
Agarose gel
0.5 g Agarose, 50 mL TAE (1 %),3 µL GELRED, at 120 Volt, running time: 45 minutes
| Sample | Sample / µl] | Loading dye (6x) / µl | Expected size |
|---|---|---|---|
| p22 (Rep40) | 50 µl | 8 µl | 940 bp |
| p23 (Rep52) | 50 µl | 8 µl | ~1200 bp |
- Marker: GeneRuler ladder mix
| Marker /µL | Sample Rep40 /µl | Sample Rep52 /µl | |
|---|---|---|---|
| Lane | 7 | 58 | 58 |
Comments: Digestion of the samples will be done tomorrow with XbaI and SpeI.
Gelextraction
Elution was done with 20 µl Pe buffer following the standard protocol.
c(Rep40)= 23,1 ng/µl
c(Rep52)= 78,0 ng/µl
Preparation of competent XL1blue and testtrafo
Investigator: Jessica
preparation of competent E.coli XL1blue was made according to the standardprotocol (91 eppis with 60µl) they are in the green boxes in -80°C
testtrafo was made
- one plate with 50pg/µl
- one plate with 100pg/µl
...trafo is failed because amp was missing on the plates (resistence in pUC). trafo will be repeated on aug 11th by Tobi...
Repetition: Mini-Prep and test digestion of pSB1C3_SDM_SspI/PvuII
Investigator: Adrian, Stefan
Plasmid Mini-Prep
- vector name: pSB1C3_SDM_SspI/PvuII
Glycerol Stocks
pSB1C3_SDM_SspI/PvuII
Only clone 1 grew, therefore there is just one stock = B159
Test digestion
- buffer used: 2
- Restriction-enzymes used:
- Enzyme 1 SspI
- Enzyme 2 PvuII
- Plasmid: pSB1C3_SDM_SspI/PvuII:
- Plasmid-Number: P185; DNA concentration: 47,9 ng/µL
- Plasmid-Number: P185; DNA concentration: 47,9 ng/µL
comment: DNA was eluted into previously used tube, therefore contamination is possible.
| Components | P185 Volume/µL | P51.1 Volume/µL |
| DNA | 18,1 | 6,8 |
| BSA (10x) | 2,5 | 2,5 |
| Buffer no. 2 (10x) | 2,5 | 2,5 |
| Enzyme 1 SspI | 0.5 | 0.5 |
| Enzyme 2 PvuII | 0.5 | 0.5 |
| H2O | 0,9 | 12,2 |
| Total volume | 10 | 10 | |
- Incubation: 45 min
Agarose-Gel:
0.4 g Agarose, 50 mL TAE, 3 µL GELRED, at 115 Volt, running time: 45 minutes
- Marker: GeneRuler ladder mix
| Marker /µL | Sample P185 /µL | Sample P51.1 /µL | |
|---|---|---|---|
| Lane | 7 | 17 | 17 |
One cut is expected (PvuII in CFP).

Comment: There are 6 bands now. This test digestion again looks very different from that of P157.2 done by Jessica (lab day 82). Results should be discussed tomorrow.
Picking clones of pSB1C3_SDM_SspI/PvuII
Investigator: Stefan
Comments: Test digestion didn't work out, three additional clones were picked from the plate we get from yesterday's Trafo to retry tomorrow.
- Bacterial strain used: BL21
- Inoculating of 10 mL DYT medium containing 10µL chloramphenicol
- Put in 37°C room on shaker at 20.45hours.
86. labday 11.08.2010
Biobrick Assembly:Picking Clones, Miniprep, Testdigestion of CMV&lITR/ hGH & rITR
Investigator: Chris L., Achim
Test Digestion showed that the cloning was succesful.

NanoDrop concentrations:
- hGH_rITR1:105,15 ng/µl
- hGH_rITR2:96.97 ng/µl
- CMV_lITR1:231.61 ng/µl
- CMV_lITR2:129.19 ng/µl
Biobrick Assembly: bSB1C3_lITR_CMV & pSB1C3_betaglobin_mVenus
Investigator: Chris L., Achim
Comments: In order to proceed with the BioBrick assembly the plasmids pSB1C3_leftITR_CMV was cut with SpeI and PstI, pSB1C3_betaglobin_mVenus was cut with XbaI and PstI. The insert fragment containing betaglobin_mVenus was ligated to the vector fragment containing the leftITR_CMV promotor.
Digestion:
| components | pSB1C3_leftITR_CMV | pSB1C3_betaglobin_mVenus |
| DNA | 5,2 | 8,4 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| Enzyme1 | 1 (SpeI) | 1 (Xba1) |
| Enzyme2 | 1 (PstI) | 1 (PstI) |
| H2O | 8,8 | 5,6 |
| Total volume | 20 | 20 |
Preparation of gel:
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time: 45 minutes
Expected sizes of constructs:
- pSB1C3_leftITR_CMV: 2800 bp
- pSB1C3_betaglobin_mVenus: 1200 bp, 2000 bp
The corresponding bands were cut out and Gel-Extraction was performed according to protocol.
concentrations measured via NanoDrop:
- pSB1C3_leftITR_CMV: 55,5 ng/µl
- pSB1C3_betaglobin_mVenus:: 10,4 ng/µl
Continuation of Cloning of Rep40/52
Investigator: Anna
- Digestion of PCR products and vector:
| components | PCR product /µl | vector /µl |
| DNA | 19 | 10 |
| BSA (10x) | 3 | 3 |
| Buffer 4 (10x) | 3 | 3 |
| Enzyme XbaI | 1 | 1 |
| Enzyme SpeI | 1 | 1 |
| H2O | 3 | 12 |
| Total volume (e.g. 15,20,25,30 µl) | 30 | 30 |
Comments: This time digestion was done with XbaI and SpeI (means it has to be checked if the inserts are cloned into the vector in the right orientation).
- Purification of Rep40 and Rep52:
For the purification 95 µl of buffer PBI was used.
c(Rep40)= 12,5 ng/µl
c(Rep52)= 51,8 ng/µl
- Gelextraction of pSB1C3_RFC25_CFP:
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 120 Volt, running time:55
5µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas)
c(pSB1C3)= 10,59 ng/µl
c(pSB1C3_2)= 4,25 ng/µl
- Ligation of PCR products and vector:
For the Ligation 1µl T4 buffer (2x) and 1µl T4 ligase were used.
| Sample-no. | vector /µl | insert /µl | |
| pSB1C3 + Rep40 | 1 | 3,65 | 4,35 |
| pSB1C3 + Rep52 | 2 | 5,87 | 2,13 |
| Total volume (e.g. 15,20,25,30 µl) | 8 |
- Transformation:
The transformation was done following the standard protocol using B21 cells.
PCR of bla_14FM and ligation into pSB1C3_SDM_SspI and pSB1C3_SDM_PvuII for virobrick-production T
Investigator: Anissa,Kerstin
Comments: Because it still is unclear if SDM of pSB1C3_SspI_PvuII has worked, we decided to use the one time mutagenisised vector pSB1C3_SDM_SspI, to clone bla into it and to make the second mutation of PvuII after bla is inserted. .
- PCR:
(was performed following the standard protocol)
| Ingredients | v(pTB106_bla14FM) - DMSO / µl | v(pTB106_bla14FM) + DMSO / µl |
| 5X Phusion HF buffer | 10 | 10 |
| 10 mM dNTP mix | 1 | 1 |
| forward primer: O116(P extreme f) | 2,5 | 2,5 |
| reverse primer: O117 (P extreme r) | 2,5 | 2,5 |
| DNA Template 1ng | 5,3 µl | 5,3 µl |
| DMSO (1%) | 0,5 | 0,5 |
| Phusion Polymerase | 0,5 | 0,5 |
| H2O | 28,2 | 27,7 |
| Total volume | 50 | 50 |
- PCR program:
| PCR Program | temperature/ °C |time | |
| 1 | 98 | 1' |
| 2 | 98 | 15 |
| 8x | 61 | 25 |
| 3 | 72 | 15 |
| 4 | 98 | 15 |
| 5 | 72 | 15 |
| 1x | 72 | 5' |
| Hold | 4 |
- Digestion PCR product:
- DNA-Concentration: 4,2 ng/µl
| components | volume / µl |
| DNA | 28,5 |
| BSA 100x | 0,4 |
| Buffer: 4 10x | 4 |
| Enzyme 1 (XbaI) | 1 |
| Enzyme 2 (PstI) | 1 |
| H2O | 5,1 |
| Total volume | 40 |
- Purification of PCR product: (QlAquick Gel Extraction Kit)
preparation of purification was made according to the standardprotocol
Comments: The wrong enzymes were used. Experiment has to be done tomorrow.
Test digestion pSB1C3_CFP_SDM SspI and PvuII
Investigator: Stefan
- buffer used: 2; Restriction-enzymes used: Enzyme 1: ClaI ; Enzyme 2: BpmI ; Enzyme 3: SspI ; Enzyme 4: PvuII
- Plasmids:
- pSB1C3_SDM_SspI/PvuII P185 (first tree lanes, cut with different enzymes)
- pSB1C3_RFC25_longlinker P105
- pSB1C3_CFP P51.1
| Components | P185/µL | P185/µL | P185/µL | P105/µL | P51.1/µL |
| DNA | 18 | 18 | 18 | 4,8 | 6,8 |
| BSA (10x) | 2,5 | 2,5 | 2,5 | 2,5 | 2,5 |
| Buffer 2 (10x) | - | - | - | 2,5 | 2,5 |
| Buffer 4 (10x) | 2,5 | 2,5 | 2,5 | - | - |
| Enzyme 1 ClaI (152) | 0,5 | 0,5 | 0,5 | - | - |
| Enzyme 2 BpmI (144) | 0,5 | 0,5 | 0,5 | - | - |
| Enzyme 3 SspI (68) | - | 0,5 | - | 0,5 | 0,5 |
| Enzyme 4 PvuII (50) | - | - | 1 | 0,5 | 0,5 |
| H2O | 1 | - | 0,5 | 14,2 | 12,2 |
| Total volume | 25 | 25 | 25 | 25 | 25 |
- Incubation: 75 minutes
Agarosegel
0.5 g Agarose, 50 ml TAE, 3 µL GELRED, at 115 Volt, running time: 60 minutes
comments: Test digestion of pSB1C3_SDM_SspI/PvuII with ClaI, BpmI and SspI looks the same as with ClaI and BpmI. Therefore we can assume that there is no additional restriction site within the vector. But still the bands above 3000 bp cannotz be explained. A new test digestion will be performed tomorrow to check the size of the vector.
The approach cut with ClaI, BpmI and PvuII looks strange. Propably, PvuII does not work correctly. New PvuII-HF was ordered and delivered so this can be verified tomorrow.
pSB1C3_RFC25_longlinker digested with SspI and PvuII seems to be not completely digested This could correspond to non-working PvuII.
pSB1C3_CFP looks like expected.
pSB1C3_SDM_Ssp1 (P125) sent for sequenzing
Investigator: Stefan
comment: To verify the correct deletion of the SspI restriction site it was sent for sequenzing.
Primer used: GATC_std_pTeSp-2
Picking clones of pSB1C3_SDM_SspI/PvuII (P185)
Investigator: Adrian
comment: pSB1C3_SDM_SspI/PvuII (P185) is empty. For plasmid production DYT + Cm was inoculated from the glycerol stock (B159).
87. labday 12.08.2010
XL1blue are ready to use
Investigator: Jessica
there are 60µl aliquots to use for one trafo
Mini-Prep of pSB1C3_SDM_SspI/PvuII and pMA_RC_insert
Investigator: Stefan
Gycerol stock was prepared for pMA_RC_insert and labeled B167. No glyerol stock for pSB1C3_SDM_SspI/PvuII because cells from earlier stock werer used for Mini-Prep.
Miniprep was performed according to protocol.
Samples were labeled:
- pSB1C3_SDM_SspI/PvuII = P185.1
- pMA_RC_insert = P190
Concentrations:
- pSB1C3_SDM_SspI/PvuII (P185.1): 257,2 ng/µl
- pMA_RC_insert (P190): 339,5 ng/µl
No test digestion was performed.
Biobrick Assembly: pSB1C3_lITR_CMV & pSB1C3_betaglobin_mVenus
Investigator: Chris L., Achim
Comments: In order to proceed with the BioBrick assembly the plasmids pSB1C3_leftITR_CMV was cut with SpeI and PstI, pSB1C3_betaglobin_mVenus was cut with XbaI and PstI. The insert fragment containing betaglobin_mVenus was ligated to the vector fragment containing the leftITR_CMV promotor.
Ligation:
For ligation, T4 ligase was used.
- 1µl T4-Ligase buffer (10x)
- 8µl (Vector + Insert) mix
- 1µl T4-Ligase
Ligation was carried out as following:
- pSB1C3_leftITR_CMV: 6,98 µl
- pSB1C3_betaglobin_mVenus: 1,02 µl
Incubation for 40 minutes.
- Transformation:
The transformation was done following the standard protocol using XL1B cells.
Repetition of cloning of bla14FM_viralbrick_MCS into pSB1C3_SDM_SspI
Investigator: Anissa
Comments:Yesterday the pcr-product of bla14FM was cut with the wrong enzymes, so no overhangs could result. Today it will be repeated with the rihgt ones (SpeI and NgomIV) .
- Digestion of the PCR product bla14FM for 2 h :
- DNA-Concentration: 10,2 ng/µl
| components | volume / µl |
| DNA | 18 |
| BSA 100x | 3 |
| Buffer: 4 10x | 3 |
| Enzyme 1 (NgomIV) | 1 |
| Enzyme 2 (SpeI) | 1 |
| H2O | 4 |
| Total volume | 30 |
- Purification of PCR product: (QlAquick Gel Extraction Kit)
preparation of purification was made according to the standardprotocol
- Digestion of vector backbone pSB1C3_SDM_SspI for 2 h:
- 1 µg vector has been used
| components | volume / µl |
| DNA | 3,12 |
| BSA 100x | 1,5 |
| Buffer: 4 10x | 1,5 |
| Enzyme 1 (NgomIV) | 1 |
| Enzyme 2 (SpeI) | 1 |
| H2O | 6,88 |
| Total volume | 15 |
- the cut vector was loaded on a preperative 1% agarose gel and the gel was running for 45 minutes
- gel was cut and gelextraction was performed according the qiagen standard protocol.
- Ligation of pSB1C3_SDM_SspI and bla14FM:
| components | volume / µl |
| t4 buffer (10X) | 1 |
| t4 Ligase | 1 |
| vector DNA | 2,72 |
| Insert | 5,28 |
- Transformation: pSB1C3_SDM_SspI_bla14FM was transformed into BL21 following the stanard protocol
Quickchange of the EaglII restriction site in the PMA_RepCap vector
Investigator: Chris L., Achim
Comments: We performed a QuikChange mutagenesis on the synthesized PMA_RepCap vector (P190). The EaglII restriction enzyme also cuts NotI (IGEM standard) therefore EaglII has to be changed into a different recognition site. Primers were designed to change it into PvuII. The new QuikChange Lighting Site-Directed Mutagenesis Kit from Stratagene was used, therefore the mutagenesis was carried out as described in the QuikChange Manual. The PCR product was transformed into XL10-Gold cells, we used the IGEM standard protocol, except for the addition of ß-ME to the XL10-Gold cells. :
PCR-reaction:
| Volume / µl | ingredients | recommended /µl |
| 2,5 | 10X reaction buffer | 2,5 |
| 2,9 | P190 template (~10 ng) | 2,9 of 1:100 dilution |
| 0,51 | forward primer: O122 | 62,5 ng |
| 0,51 | reverse primer: O123 | 62,5 ng |
| 0,5 | dNTP | 250 µM each dNTP |
| 17,6 | H2O | |
| 0,5 | QuikChange Lightning ennzyme(1.25) |
PCR program:
| Rounds | temperature/ °C | Time |
| 1 | 95 | 2 minutes |
| 18 | 95 | 20 seconds |
| 18 | 60 | 10 seconds |
| 18 | 68 | 2 minutes (4 kb * 30 sec) |
| 1 | 68 | 5 minutes |
The PCR product was digestet with Dpn I for 10 minutes at 37°C
Plasmid was transformed into XL10-Gold cells.
Sequencing results of Rep68, Rep78 and AAP BioBricks
Investigator: Hanna
Comment: pSB1C3_Rep78 (P160), pSB1C3_Rep68 (P165) and pSB1C3_AAP (P170) were sent for sequencing twice. No sequencing made sense. Paradoxically the Rep78 sequencing result fit to the p5 promoter BioBrick...
By having a closer look to the BioBrick PCR we found out that the fragment sizes didn't fit to the expected sequences. In addition to that the test digestion also showed some discrepancies.
Because we weren't able to find out in which steps of the BioBrick production things went wrong, we decided to discard all referring glycerol stocks and plasmids and to perform the Rep68, Rep78 and AAP BioBrick PCR again!!!
Comment 13.08.2010 (Hanna): We wondered how the whole Rep stuff could be sequenced reverse AND forward without any forward sequencing primer... By checking the "sequencing book" we found out that always the wrong primers were used: the GOI primers can be used in order to sequence the Gene of interest = GOI (!) in pAAV_MCS and NOT the pSB1C3!!!!!!!!!
Order oligos for CD-biobrick
Investigator: Jessica and Kira
ordered on aug 12th
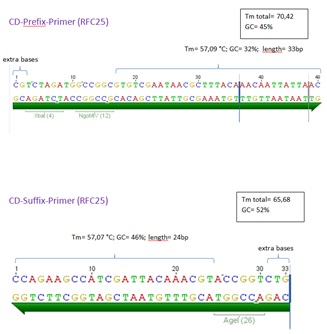
BioBrick assembly of pSB1C3_leftITR_CMV_mVenus
Investigator: Stefan
comment: In order to assemble our complete constructs mVenus has to be included into the pSB1C3_leftITR_CMV vector.
Digestion:
| components: | pSB1C3_leftITR_CMV P188(Vector) | pGA14_mVenus P60 (Insert) |
| DNA | 6 | 8 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| Enzyme1 | 1 (SpeI) | 1 (Xba1) |
| Enzyme2 | 1 (PstI) | 1 (PstI) |
| H2O | 8 | 6 |
| Total volume | 20 | 20 |
Preparation of gel:
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 110 Volt, running time: 45 minutes
Expected sizes of constructs:
- pSB1C3_leftITR_CMV: 2869 bp
- pGA14_mVenus: 2896 bp, 756 bp
The corresponding bands were cut out and Gel-Extraction was performed according to protocol.
concentrations measured via NanoDrop:
- pSB1C3_leftITR_CMV: 39,2 ng/µl
- pGA14_mVenus: 8,2 ng/µl
Ligation:
T4 ligation was performed according to protocol.
Volumes used:
- vector: 1,67 ng/µl
- insert: 6,33 ng/µl
Transformation:
bacterial strain used: XL1bB
antibiotic used for plate: chlorampenicol
Transformation was performed according to protocol.
Plate was prepared and put in 37°C room.
Opened a BioBrick box
Investigator: Bea
In order to "save" the plasmids/BioBricks which has been prepared and confirmed for submitting it to the parts registry a new box have been prepared. The box should contain 30 µL of the desired construct cloned into the pSB1C3 vector and the BBa number given from the headquarter.
The first BioBricks which are in this box are:
- pSB1C3_hGH (P136): Bba_K404002
- pSB1C3_betaglobin (P133: Bba_K404001
- pSB1C3_CMV (P145): Bba_K404003
- pSB1C3_leftITR (P175): Bba_K404006
- pSB1C3_rightITR (P172): Bba_K404005
- pSB1C3_mGMK (P156): Bba_K404004
In our nomenclature plasmid excel list the constructs which we aliquoted should be highlighted in blue and commented with the given Bba number. Additionally it should be taken in consideration that enough DNA should be in the "working" plasmid box. Therefore, if necessary, inoculate DYT medium for Mini-Preps.
In this case ALL the glycerol stocks were used for incoluating 10 mL DYT medium containing 10 µL chloramphenicol respectively.
To do: Mini-Preps of the over night cultures.
88. labday 13.08.2010
Continuation:pSB1C3_SDM_SspI_bla14FM
Investigator: Patrick
Intention: receive enough plasmid for a sequencing and maybe futher working steps.
Three clones were picked in the morning to inoculate DYT and perform a mini-prep in the evening and to send them for sequencing. The three clones were picked and prepped according to the standard protocol yielding the following DNA concentrations:
- pSB1C3_SDM_SspI_bla14FM clone 1: 65,15 ng/µl , Sequencing labeled: PS3.1
- pSB1C3_SDM_SspI_bla14FM clone 2: 22,3 ng/µl , Sequencing labeled: PS3.2
- pSB1C3_SDM_SspI_bla14FM clone 3: 62,3 ng/µl , Sequencing labeled: PS3.3
Used primer: seq_pSB1C3_VR2_rev (O51)
Results: see http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/index.php/Laborjournal#Sequence_analysis_of_pSB1C3_SDM_SspI_BLA
Mini-preps of pSB1C3_hgh,pSB1C3_beta globin,pSB1C3_left ITR,pSB1C3_right ITR,pSB1C3_mGMK,pSB1C3_CMV
Comments:new mini-preps of the glycerol-stocks ofpSB1C3_hgh(=B103),pSB1C3_beta globin(=B100),pSB1C3_left ITR (=B143),pSB1C3_right ITR (=B140),pSB1C3_mGMK (=B125),pSB1C3_CMV :
| components | concentration in ng/µl |
| P191 (pSB1C3_right ITR) | 100,74 |
| P192 (pSB1C3_beta-globin) | 240,08 |
| P193 (pSB1C3_CMV) | 333,89 |
| P194 (pSB1C3_hgh) | 328,42 |
| P195 (pSB1C3_mGMK) | 258,63 |
| P196 (pSB1C3_left ITR) | 173,98 |
Design and ordering of oligos for VP fusion targeting approaches
Investigator: Hanna
Comment: In order to target the EGFR receptor of our A431 cells different approaches have been figured out: Besides loop insertions, N-terminal fusion to the VP2 protein will be performed. In addition to that a VP1 insertion approach (after Grieger et al., 2007) was planned.
Fusion and insertion molecules will be e.g. His-tag, Affibody, Darpin, GFP. These approaches can be combinated with the loop insertions.
For this purpose the following oligos were ordered today:
Mini-Prep and Test digestion of Rep40 and Rep52 of the cloning experiment ("Cloning of Rep40/52" 11.08.2010 Investigator: Anna)
Investigator: Chris L., Achim
Comments: There are small and big colonies on the plate. Maybe problems with the agar plate. Picked colonies of pSB1C3_Rep40 sample 1 and pSB1C3_Rep52 sample 1 are small colonies. Picked colonies of pSB1C3_Rep40 sample 2 + 3 and pSB1C3_Rep52 sample 2 + 3 are standard size colonies.
Nanodrop
- Rep 40 Sample 1: 156,26 ng/µl P197
- Rep 40 Sample 2: 200,05 ng/µl P198
- Rep 40 Sample 3: 144,06 ng/µl P199
- Rep 52 Sample 1: 173,36 ng/µl P200
- Rep 52 Sample 2: 160,75 ng/µl P201
- Rep 52 Sample 3: 295,77 ng/µl P202
Test digestion 800ng DNA
| components | P145 /µl | P146 /µl | P145 /µl | P146 /µl | P145 /µl | P146 /µl |
| DNA | 5,1 | 4 | 5,6 | 4,6 | 5 | 2,7 |
| BSA (10x) | 2 | 2 | 2 | 2 | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 | 2 | 2 | 2 | 2 |
| Enzyme 1: XbaI | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 |
| Enzyme 2: SpeI | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 |
| H2O | 9,9 | 11 | 9,4 | 10,4 | 10 | 12,3 |
| Total volume (e.g. 15,20,25,30 µl) | 20 | 20 | 20 | 20 | 20 | 20 |
Incubation time: 45 minutes, Incubation temperature: 37°
0,5 g Agarose, 50ml x TAE, 3µl GELRED, at 115 Volt, running time: 45 minutes
| Sample | Sample/µl] | Loading dye (6x)/µl | Expected size 1 (Geneious) | Expected size 2 (Geneious) |
|---|---|---|---|---|
| Rep 40 1 | 20 µl | 4 µl | ____ bp | 940 bp |
| Rep 40 2 | 20 µl | 4 µl | ____ bp | 940 bp |
| Rep 40 3 | 20 µl | 4 µl | ____ bp | 940 bp |
| Rep 52 1 | 20 µl | 4 µl | ____ bp | 1198 bp |
| Rep 52 2 | 20 µl | 4 µl | ____ bp | 1198 bp |
| Rep 52 3 | 20 µl | 4 µl | ____ bp | 1198 bp |
| Marker | Sample Rep40 1 /20µl | Sample Rep40 2 /20µl | Sample Rep40 3 /20µl | Sample Rep52 1 /20µl | Sample Rep52 2 /20µl | Sample Rep52 3 /20µl | |
|---|---|---|---|---|---|---|---|
| Lane | 1 | 2 | 3 | 4 | 5 | 6 | 7 |
agarose gel: 1%, digestion: 45 minutes, 37°C
Comments:Results look bad. Rep52 clone 3 looks good, send for sequencing.
</p>
Repetition of Rep 40 and Rep 52 ligation and transformation with the vector pSB1C3_CFP
Investigator: Chris L.
- Ligation of PCR products and vector:
For the Ligation 1µl T4 buffer (10x) and 1µl T4 ligase were used.
| Sample-no. | vector /µl | insert /µl | |
| pSB1C3 + Rep40 | 1 | 4,92 | 3,08 |
| pSB1C3 + Rep52 | 2 | 6,47 | 1,53 |
| Total volume (e.g. 15,20,25,30 µl) | 10 | 10 |
- Transformation:
The transformation was done following the standard protocol using XL1B cells.
Strategy for VP2 N-terminal fusion and VP1 insertion of targeting moelcules
Investigator: Hanna
Comment: In order to target the EGFR receptor of tumor cells via Affibody or Darpin (antibody fragment is also possible!!!), or in order to expose e.g. His-Tags on the virus surface, two approaches have been figured out.


pSB1C3_Rep52 (P202) sent for sequencing
Investigator: Chris L.
Primer used: Reverse Primer VR2
<p style="font-size:15px; background-color:#66bbff;">Repetition: Biobrick production of Rep 68, 78 and AAP
Investigator: Stefan
Aim of the experiment:
We want to produce biobricks from Rep 68, Rep 78 and the AAP.
- Plasmids used as template:
Rep_68_ex (p119): c = 470,6 ng/µl
Rep_78_(p122): c = 566,8 ng/µl
pAAV-RC containing AAP ORF (p50): c = 378,5 ng/µl
- Primer used:
For Rep_68_ex: Praefix_68_78_ex & Suffix_40_68_ex
For Rep_78_ex: Praefix_68_78_ex & Suffix_52_78_ex
For AAP_ex: Praefix_AAP_ex & Suffix_AAP_ex
- PCR:
(was performed following the standard protocol)
| Ingredients | Volume / µl | Rep68 | Rep78 | AAP |
| 5X Phusion HF buffer | 10 | |||
| 10 mM dNTP mix | 1 | |||
| forward primer: | 2,5 | |||
| reverse primer: | 2,5 | |||
| DNA Template | *** | 2,5 µl | 2 µl | 3 µl |
| DMSO (2%) | - | - | 1 µl | |
| Phusion Polymerase | 0,5 | |||
| H2O | *** | 31 µl | 31,5 µl | 29,5 µl |
| Total volume | 50 |
PCR program:
| PCR Program | temperature/ °C | Time | Rep68 | Rep78 | AAP |
| 1 | 98 | 1min | |||
| 2 | 98 | 15s | |||
| 3 | *** | 25s | 63°C | 62°C | 64°C |
| 4 (step 2-4 8x) | 72 | *** | 24s | 27s | 10s |
| 5 | 98 | 15s | |||
| 6 | *** | 25s | 68°C | 64°C | 68°C |
| 7 | 72 | *** | 24s | 27s | 10s |
| 8 (step 6-8 17x) | 72 | 5min | |||
| Hold | 4 |
Used agarose gel: 0,5 g Agarose,50 ml TBE (1%), 3 µl GELRED , at 115 Volt, running time:45 minutes
Gel extraction
Gel extraction was perfomed according to protocol.
- Digestion of PCR products and vector:
| components | PCR products /µl | vector /µl |
| DNA | 29 | 11 |
| BSA (10x) | 4 | 2 |
| Buffer 4 (10x) | 4 | 2 |
| Enzyme XbaI | 1 | 1 |
| Enzyme SpeI | 1 | 1 |
| H2O | 1 | 3 |
| Total volume (e.g. 15,20,25,30 µl) | 40 | 20 |
Comments: Digestion was done with XbaI and SpeI, it has to be checked if the inserts are cloned into the vector in the right orientation.
- Purification of Rep68, 78 and AAP:
For the purification 200 µl of buffer PBI was used.
c(Rep68)= 14,3 ng/µl
c(Rep78)= 12,0 ng/µl
c(AAP)= 25,1 ng/µl
- Gelextraction of pSB1C3_RFC25_CFP:
0,5 g Agarose,50 ml TBE (1%), 3 µl GELRED , at 115 Volt, running time:50 minutes
4µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas)
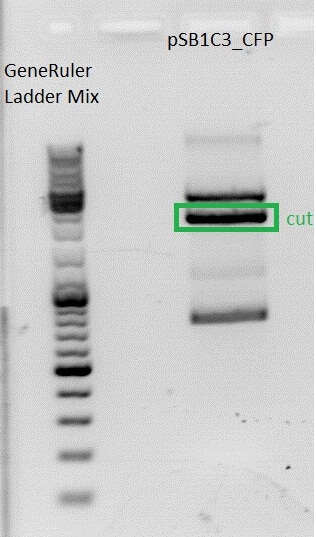
c(pSB1C3)= 6,9 ng/µl
- T4 ligation of PCR products and vector:
For the Ligation 1µl buffer (10x) and 1µl T4 ligase were used.
| vector /µl | insert /µl | |
| pSB1C3 + Rep68 | 3,78 | 4,24 |
| pSB1C32 + Rep78 | 3,13 | 4,87 |
| pSB1C3 + AAP | 6,42 | 1,58 |
- Transformation:
The transformation was done following the standard protocol using XL1b cells.
89. labday 14.08.2010
Sequence analysis of pSB1C3_SDM_SspI_BLA
Investigator: Patrick, Bea
Comments: Sequence analysis of three clones revealed that all sequences contain some mutations in the regions where the primer annealed. This was due to the extreme primer lentgh. Primers that long should be ordered as HPLC purified in order to avoid mutations in the primer sequence.
- Clone 1: Mutation in the SalI site of the loop insertion site in the Viral Brick MCS lead to the removal of the recognition site. CANNOT be used for the loop insertions!!!!
- Clone 2: Mutation in the region of the primer upstream of the loop insertion sites SalI and PvuII.
- Clone 3: Mutation in the region of the primer downstream of the loop insertion sites SalI and PvuII.
Next steps: Two additional clones will be picked and sent for sequencing (clones 4 and 5). In order to obtain a construct without any mutations, clone 2 and clone 3 couldbe cloned together. Each clone will be digested with SalI and BamHI. Clone 2 will be the "insert", clone 3 is going to be the "vector". Therefore the fragments will (hopefully) not contain anymore mutations in the relevant sequence.
Mini Preps of PMA_RepCap vector and pSB1C3_left ITR_CMV_betaglobin_mVenus
Investigator: Chris L., Bea
Comments:new mini-preps of the glycerol-stocks of pSB1C3_leftITR_CMV_betaglobin_mVenus clone 1 (=B179), pSB1C3_leftITR_CMV_betaglobin_mVenus clone 2 (=B180), pSB1C3_leftITR_CMV_betaglobin_mVenus clone 3 (=B181), pMA_RepCap Vector_SDM_InsPvuII clone 1, (=B182), pMA_RepCap Vector_SDM_InsPvuII clone 2 (=B183) and pMA_RepCap Vector_SDM_InsPvuII clone 3 (=B184)
Comment: The plamsid numbers in the picture (P179-P184) correspond to the glycerol stocks B179-B184. The corresponding plasmid numbers are: P208-P213.
Nanodrop
- pSB1C3_leftITR_CMV_betaglobin_mVenus clone 1: 256,64 ng/µl B179
- pSB1C3_leftITR_CMV_betaglobin_mVenus clone 2: 274,86 ng/µl B180
- pSB1C3_leftITR_CMV_betaglobin_mVenus clone 3: 217,20 ng/µl B181
- pMA_RepCap Vector_SDM_InsPvuII clone 1: 253,71 ng/µl B182
- pMA_RepCap Vector_SDM_InsPvuII clone 2: 195,64 ng/µl B183
- pMA_RepCap Vector_SDM_InsPvuII clone 3: 144,61 ng/µl B184
Test digestion 800ng DNA
| components | B179 /µl | B180 /µl | B181 /µl | B182 /µl | B183 /µl | B184 /µl | pMA_RepCap Vector /µl |
| DNA | 3,1 | 2,9 | 3,7 | 3,1 | 4 | 5,5 | 2,4 |
| BSA (10x) | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| Enzyme 1: NgoMIV | 0,5 | 0,5 | 0,5 | 0 | 0 | 0 | 0 |
| Enzyme 2: EcoRI | 0,5 | 0,5 | 0,5 | 0 | 0 | 0 | 0 |
| Enzyme 2: PvuII | 0 | 0 | 0 | 0,5 | 0,5 | 0,5 | 0,5 |
| H2O | 11,9 | 12,1 | 11,3 | 12,4 | 11,5 | 10 | 13,1 |
| Total volume (e.g. 15,20,25,30 µl) | 20 | 20 | 20 | 20 | 20 | 20 | 20 |
Incubation time: 45 minutes, Incubation temperature: 37°
0,5 g Agarose, 50ml x TAE, 3µl GELRED, at 115 Volt, running time: 45 minutes
Results of test digestion
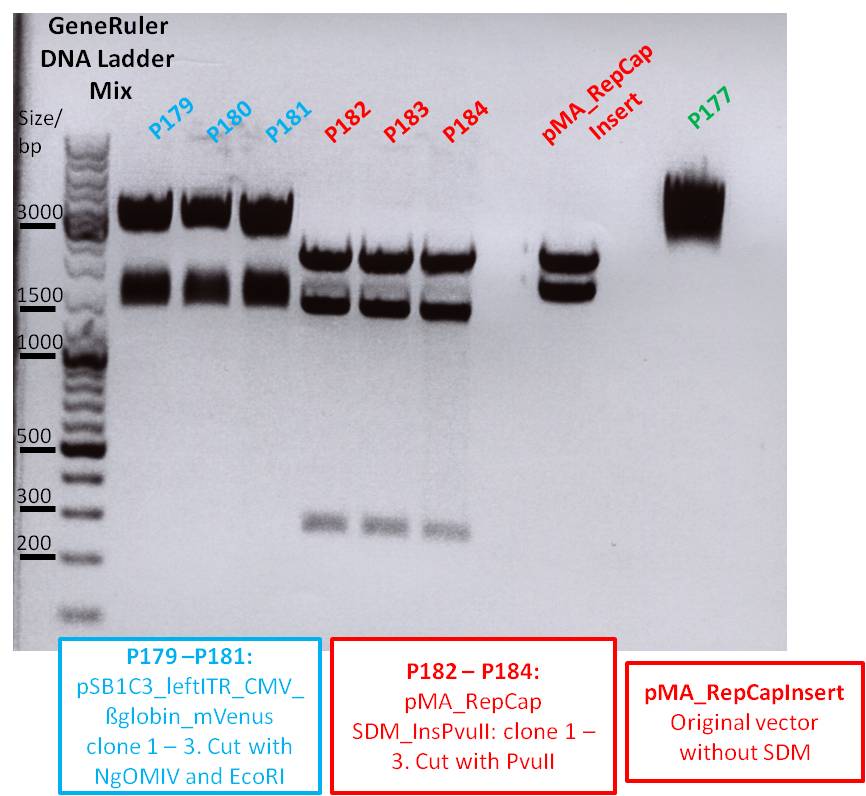
Results: Two (three) different approaches can be seen on the agarose gel.
- P178-P181 (blue): The plasmid pSB1C3_leftITR_CMV_betaglobin_mVenus was digested with NgoMIV and EcoRI. Expected bands were: 2873bp and 1343 . As it can be seen in the picture, the two fragments ran too high (~300bp). As it is almost normal for the pSB1C3 plasmid, we should figure out why this is always the problem for this vector.
- P182-P183 (red): The plasmid pMA_RepCap_SDM_insPvuII was digested with PvuII as well as the control vetor pMA_RepCap where no site-directed mutagenesis was performed in order to see the difference. As it can be seen in the picture above, the expected band sizes of 263bp,1384bp and 2133bp correspond to the bands detected in the gel. The control vector instead reveals only two bands detectable. Therefore site-directed mutagenesis was successfully performed.
- P177 (green): pSB1C3_hTERT. For further details see extra part below (Test digestion of pSB1C3_hTERT.
Mini-Prep of pSB1C3_leftITR_CMV_mVenus
Investigator: Stefan
Glycerol stocks were prepared:
Test digestion: or test digestion, pSB1C3_leftITR_CMV (P188) was digested as well.
| Ingredients | clone 1/µl | clone 2/µl | P188/µl |
| DNA: | 3,5 | 3,5 | 4 |
| BSA (10x): | 2 | 2 | 2 |
| Buffer 4: | 2 | 2 | 2 |
| Enzyme 1 EcoRI: | 0,5 | 0,5 | 0,5 |
| Enzyme 2 AgeI | 0,5 | 0,5 | 0,5 |
| H2O | 6,5 | 6,5 | 6 |
-
All samples were loaded on a 1% agarose gel:
- GeneRuler Ladder Mix (7µl)
- P206 = pSB1C3_leftITR_CMV_mVenus clone 1
- P207 = pSB1C3_leftITR_CMV_mVenus clone 2
- P188 = pSB1C3_leftITR_CMV
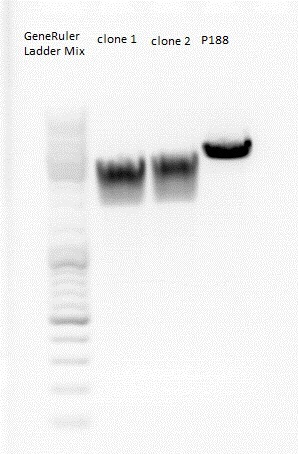
comment: Through digestion, fragments would have similar sizes. Therefore, a new digestion will be performed tomorrow.
Test digestion of pSB1C3_pHTERT
Investigator: Bea
Comments: pSB1C3_hTERT was sent for sequencing and revealed that the PCR and the inserting of the PCR product : promoter for hTERT was sucessfully performed. But the sequencing results showed a new AgeI restriciton site in the plasmid backbone. We could not see if the new recognition site was due to the bad sequence read or that there is really a new AgeI site. Therefore, a test digestion will be perfomed with PstI and AgeI in order to figure our wheter AgeI is present in the backbone or not.
Sequence analysis of pSB1C3_hTERT:
First the sequenced plasmid P177 was digested:
| P177/µL | |
| DNA | 3 |
| BSA (10x) | 1,5 |
| Buffer 4 (10x) | 1,5 |
| Enzyme PstI | 0,5 |
| Enzyme AgeI | 0,5 |
| H2O | 8 |
| Total volume | 15 |
- Incubation time: 45minutes
- Incubation tempereature: 37°C
After the incubation,the samples were loaded on the 1% agarose gel, and run at 110 V for 45 minutes.
Result of the test digestion:
Results: As it can be seen in the sequencing results of pSB1C3_hTERT there could have been an additional AgeI site in the vector. If AgeI would have been in the vector, two fragments have been expected by digetsing the plasmid with AgeI and PstI. As it can be seen in the agarose gel, lane 11 (labeled with the green P177) only reveals one lane instead of two. Therefore it can be assumed that NO AgeI site is in the pSB1C3 backbone.
Picking clones ...
Investigator: Patrick 15 Clones were picked saturday night so that a mini-prep could be performed on sunday.
90. labday 15.08.2010
Sequence analysis of pSB1C3_Rep52
Investigator: Bea
Comments: Clone 3 was sent for sequencing because this clone revealed the right band sizes of the test digestion performed friday. For more details see: [http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/index.php/Laborjournal#Mini-Prep_and_Test_digestion_of_Rep40_and_Rep52_of_the_cloning_experiment_.28.22Cloning_of_Rep40.2F52.22_11.08.2010_Investigator:_Anna.29 Test digestion]
Sequence assembly showed that the suffix of the Rep protein 52 is ok and that PCR worked well.
There can be seen that a silent mutation from AGG to AGA was inserted, but it will have no effect on the protein because codon usage in H. sapiens is nearly the same for both codons.
The prefix could not be analysed because of bad sequence read quality. For further verification, another sequencing with the forward primer has to be performed.
Next steps: Send plasmid for another sequencing round with forward primer in order to analyse the prefix.
[http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/index.php/Laborjournal top of page]
Mini-Prep of pSB1C3_Rep68, pSB1C3_Rep78, pSB1C3_RepAAP, pSB1C3_Rep40, pSB1C3_Rep52
Investigator: Stefan
-
Glycerol stocks were prepared:
- B185 = pSB1C3_Rep68 clone 1
- B186 = pSB1C3_Rep68 clone 2
- B187 = pSB1C3_Rep68 clone 3
- B188 = pSB1C3_Rep78 clone 1
- B189 = pSB1C3_Rep78 clone 2
- B190 = pSB1C3_Rep78 clone 3
- B191 = pSB1C3_AAP clone 1
- B192 = pSB1C3_AAP clone 2
- B193 = pSB1C3_AAP clone 3
- B194 = pSB1C3_Rep40 clone 1
- B195 = pSB1C3_Rep40 clone 2
- B196 = pSB1C3_Rep52 clone 1
- B197 = pSB1C3_Rep52 clone 2
- B198 = pSB1C3_SDM_SspI_Bla14FM clone 4
- B199 = pSB1C3_SDM_SspI_Bla14FM clone 5
Mini-Prep following the standard protokol
- P208 = pSB1C3_Rep68 clone 1 c=230,0
- P209 = pSB1C3_Rep68 clone 2 c=160,5
- P210 = pSB1C3_Rep68 clone 3 c=128,6
- P211 = pSB1C3_Rep78 clone 1 c=150,8
- P212 = pSB1C3_Rep78 clone 2 c=108,6
- P213 = pSB1C3_Rep78 clone 3 c=131,3
- P214 = pSB1C3_AAP clone 1 c=142,1
- P215 = pSB1C3_AAP clone 2 c=148,3
- P216 = pSB1C3_AAP clone 3 c=163,0
- P217 = pSB1C3_Rep40 clone 1 c=164,5
- P218 = pSB1C3_Rep40 clone 2 c=<145,1br />
- P219 = pSB1C3_Rep52 clone 1 c=156,0
- P220 = pSB1C3_Rep52 clone 2 c=164,0
- P221 = pSB1C3_SDM_SspI_Bla14FM clone 4 c=220,4
- P222 = pSB1C3_SDM_SspI_Bla14FM clone 5 c=252,2
Test digestion:
No test digestion for pSB1C3_SDM_SspI_Bla14FM clone 4 and 5.
For test digestion, pSB1C3_leftITR_CMV_mVenus (P206-207) and pSB1C3_leftITR_CMV (P188) was digested as well.
Samples P208-220 were digested with XbaI and SpeI.
Samples P207, P207 and P188 were digested with XbaI and NgoMIV.
Gel used:
100 ml TAE (1x), 1 g agarose, 6 µl GELRED
115 V, 50 minutes
| Ingredients | P208/µl | P209/µl | P210/µl | P211/µl | P212/µl | P213/µl | P214/µl | P215/µl | P216/µl | P217/µl | P218/µl | P219/µl | P220/µl | P206/µl | P207/µl | P188/µl |
| DNA: | 4 | 6 | 7 | 6 | 9 | 7 | 6 | 6 | 6 | 6 | 6 | 6 | 6 | 3,5 | 3,5 | 4 |
| BSA (10x): | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| Buffer 4: | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 | 2 |
| Enzyme 1 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 |
| Enzyme 2 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 | 0,5 |
| H2O | 11 | 9 | 8 | 9 | 6 | 8 | 9 | 9 | 9 | 9 | 9 | 9 | 9 | 11,5 | 11,5 | 11 |

comment: Rep- and AAP approaches have to be repeated. We did not realize that the vector can religate! Plasmids and glycerol stocks were thrown away.
Inoculation of CD colony
Investigator: Kira
One colony was picked from agar plate and dispersed into 10 ml DYT+ Amp and incubated over night @ 37 C.
91. labday 16.08.2010
Minipreps of P119, P122, P204, P205
Investigator: Achim
Cloning of pSB1C3_SspIdel_BLA clone 2 & 3 to get rid of mutations
Investigator: Achim
- Sequencing showed that clone 2 had a mutation in the vector, clone 3 had one in the BLA sequence. Therefore, both were cut with BamHI and SalI, then the BLA fragment of clone 2 and the vector fragment of clone 3 were ligated. The final construct shouldn't contain any more mutations.
Nice picture, but where's the protocol? Results? (Hanna) ;)
New working solutions were prepared
Investigator: Patrick
- DMEM
- Amp
- H2O millipore
- DYT
Further pSB1C3_SDM_SspI_Bla14FM sequencing
Two additional Clones were picked (clones 4 & 5) and sent for sequencing, labeled: iGEM4.1 and iGEM4.2.
Used Primer: seq_pSB1C3_VR2_rev (O51)
Investigator: Patrick
Mini prep of pKS-CD
Investigator: Kira
c(pKS-CD) = 291, 39 ng/µl
Yielded DNA was used for site directed mutagenesis
SDM of PstI in pKS-CD
Investigator: Jessica, Kira
Motivation: the desired gen contains 2 iGEM restrictions sites, thus in oder to use it for further cloning, these site have to be deleted. The first SDM was performed on PstI restriction site.
- P223 pKS_CD
- O154 CD_PstI FP
- O155 CD_PstI RP
| component | Volume/µl |
| DNA: (1:20) | 0,5 |
| 10x reaction buffer: | 2,5 |
| primer forward PstI (1:10) | 0,56 |
| primer reverse PstI (1:10) | 0,56 |
| dNTP | 0,5 |
| DMSO | 0,5 |
| H2O | 14,38 |
| PfuTurbo DNA Polymerase | 0,5 |
PCR
| segment | cycles | temperature | time |
| 1 | 1 | 95 | 2min |
| 2 | 20 | 95 | 30sec |
| 55 | 1min | ||
| 68 | 4 min30 sec | ||
| 3 | 4 | hold |
sample was digested with DpnI for 1h at 37°C
Transformation was performed according to the standard protocol.
Biobrick production of pSB1C3_lITR_CMV_beta-globin_mVENUS_hgh_rITR and pSB1C3_lITR_CMV_mVENUS_hgh_rITR
Investigator: Anna, Anissa
Comments: hgh_rITR was cloned one time into pSB1C3_lITR_CMV_beta-globin_mVENUS and one time into pSB1C3_lITR_CMV_mVENUS to receive a control for the improtance of beta-globin
- Digestion:
| components | volume of pBS1C3_hgh_rITR INSERT = p186 /µl | volume of pBS1C3_lITR_CMV_beta-globin_mVENUS VECTOR = p208 /µl | volume of pBS1C3_lITR_CMV_mVENUS VECTOR = p207 /µl |
| DNA | 19 | 6 | 3,84 |
| BSA (10x) | 3 | 3 | 1,5 |
| Buffer 4 (10x) | 3 | 3 | 1,5 |
| SpeI° /XbaI* | 1* | 1° | 1° |
| PstI | 1 | 1 | 1 |
| H2O | 3 | 6 | 6,16 |
| Total volume | 30 | 20 | 15 |
The samples were digested for 2 h at 37°C
- Gel: samples were loaded on a 1% agarose-gel. running time: 45 minutes
- gelextraction was performed according with the standard-protocol
- Ligation:
| components | concentration/ ng/µl |
| p186 | 8,97 |
| p207 | 25,57 |
| p208 | 17,73 |
| components | amount for ligation /µl |
| p186 (for p207) | 4,2 |
| p207 | 3,8 |
| p186 (for p208) | 4,7 |
| p208 | 3,3 |
| T4 ligation buffer | 1 |
| T4 Ligase | 1 |
- Transformation: was performed according with the standard-protocol
Preparation for Midi-Preps of B24 and B34
Investigator: Anna
- 40 ml DYT was prepared with 40 µl Ampicillin and inoculated with B24 and B34, both were incubated over night in 37°C room.
What is B24 and B34? Can you please be more precise - I also want to understand what's happening also when I am not able to check up in the excel sheets ;) (Hanna)
Cloning of pAAV_RC (P50) with pMA_RC_insert (P190)
Investigator: Chris L.
- Vector: name: pAAV_RC P50
- Insert: name: pMA_RC_insert P190
- buffer used: 3
- Restriction-enzymes used:
- BstEII (no. Lab: 17)
- SwaI (no.Lab: 135)
- DNA concentration (P50): 378,5 ng/µl
- DNA concentration (P190): 339,5 ng/µl
Digestion:
| components | volume of pMA_RC_insert /µl | volume of pAAV_RC /µl |
| DNA | 4,4 | 2,6 |
| BSA (10x) | 2 | 2 |
| Buffer 3 (10x) | 2 | 2 |
| SwaI (no.Lab:135) | 0,5 | 0,5 |
| BstEII (no.Lab:17) | 0,5 | 0,5 |
| H2O | 10,6 | 12,4 |
| Total volume | 20 | 20 |
Comments: Incubation at 25° after addition of SwaI for 1,5 hours. Then incubation at 60° after addition of BstEII for 1,5 hours.
Gel extraction:
Preparation of gel:
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time: 45 minutes
Expected sizes of constructs:
- pAAV_RC 6839 bp and 493 bp
- pMA_RC_Insert 3292 bp and 493 bp
| Marker | Sample P190, 20µl | Sample P 50, 20µl | |
|---|---|---|---|
| Lane | 3 | 5 | 7 |
Bands are really weak, tomorrow I`ll take more DNA in sample.
Next steps: Tomorrow repetition of experiment
Repetition: Biobrick production of Rep40, Rep68, Rep78 and AAP
Investigator: Stefan
Aim of the experiment:
We want to produce biobricks from Rep40, Rep68, Rep78 and AAP.
- Plasmids used as template:
Rep_68_ex (p119): c = 470,6 ng/µl
Rep_78_(p122): c = 566,8 ng/µl
pAAV-RC containing AAP ORF (p50): c = 378,5 ng/µl
- Primer used:
For Rep_68_ex: Praefix_68_78_ex & Suffix_40_68_ex
For Rep_78_ex: Praefix_68_78_ex & Suffix_52_78_ex
For AAP_ex: Praefix_AAP_ex & Suffix_AAP_ex
- PCR:
(was performed following the standard protocol)
| Ingredients | Volume / µl | Rep68 | Rep78 | AAP | Rep40 |
| 5X Phusion HF buffer | 10 | ||||
| 10 mM dNTP mix | 1 | ||||
| forward primer: | 2,5 | ||||
| reverse primer: | 2,5 | ||||
| DNA Template | *** | 2,5 µl | 2 µl | 3 µl | 1 µl |
| DMSO (2%) | - | - | 1 µl | - | |
| Phusion Polymerase | 0,5 | ||||
| H2O | *** | 31 µl | 31,5 µl | 29,5 µl | 31,5 µl |
| Total volume | 50 |
PCR program:
| PCR Program | temperature/ °C | Time | Rep68 | Rep78 | AAP | Rep40 |
| 1 | 98 | 1min | ||||
| 2 | 98 | 15s | ||||
| 3 | *** | 25s | 63°C | 62°C | 64°C | 63°C |
| 4 (step 2-4 8x) | 72 | *** | 24s | 27s | 10s | 15s |
| 5 | 98 | 15s | ||||
| 6 | *** | 25s | 68°C | 64°C | 68°C | 68°C |
| 7 (step 5-7 17x) | 72 | *** | 24s | 27s | 10s | 15s |
| 8 | 72 | 5min | ||||
| Hold | 4°C |
Used agarose gel: 0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time:45 minutes
Comments: I forgot to dilute Rep68. Therefore, a new PCR will have to be performed tonight.
Gel extraction
Gel extraction was perfomed according to protocol.
- Digestion of PCR products and vector:
| components | PCR products /µl | vector /µl |
| DNA | 29 | 11 |
| BSA (10x) | 4 | 2 |
| Buffer 4 (10x) | 4 | 2 |
| Enzyme XbaI | 1 | 1 |
| Enzyme SpeI | 1 | 1 |
| H2O | 1 | 3 |
| Total volume (e.g. 15,20,25,30 µl) | 40 | 20 |
Comments: Digestion was done with XbaI and SpeI, it has to be checked if the inserts are cloned into the vector in the right orientation.
- Purification of Rep40, 78 and AAP:
For the purification 200 µl of buffer PBI was used.
c(Rep40)= 6,5 ng/µl
c(Rep78)= 14,3 ng/µl
c(AAP)= 39,6 ng/µl
- Gelextraction of pSB1C3_RFC25_CFP:
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time:50 minutes
4µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas)

c(pSB1C3)= 19,5 ng/µl
- T4 ligation of PCR products and vector:
For the Ligation 1µl buffer (10x) and 1µl T4 ligase were used.
| vector /µl | insert /µl | |
| pSB1C3 + Rep40 | 1,57 | 6,43 |
| pSB1C32 + Rep78 | 1,71 | 6,29 |
| pSB1C3 + AAP | 5,56 | 2,44 |
- Transformation:
The transformation was done following the standard protocol using XL1b cells.
Design and Ordering of Oligos
Investigator: Hanna
SV40 terminator
Comment: The CMV promoter is often used in combination with a SV40 terminator. Because we want to use this promoter in the context of the VP2 fusion and VP1 insertion approaches, oligos were designed in order to produce a SV40 terminator BioBrick (from the pEGFP-C1 plasmid).
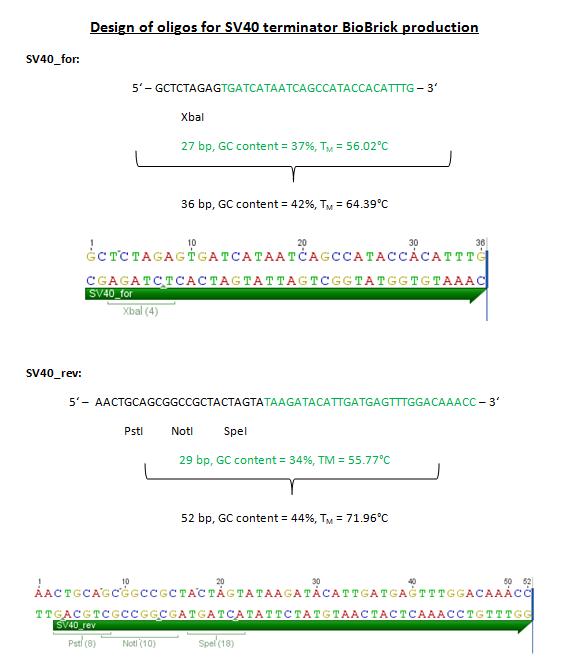
pEGFP_backbone
Comment: In order to perform the VP2 N-terminal fusion and VP1 insertion approaches, we decided to use pEGFP-C1 as expression plasmid. This plasmid contains an EGFP under the control of the CMV promoter and the SV40 terminator. In order to exchange the EGFP with our targeting constructs a PCR of the CMV_backbone_SV40Terminator should be performed. The referring primers were designed with EcoRI and PstI restriction sites. One should keep in mind that this expression plasmid mustn't digested with NgoMIV and it contains Kanamycin-resistance!
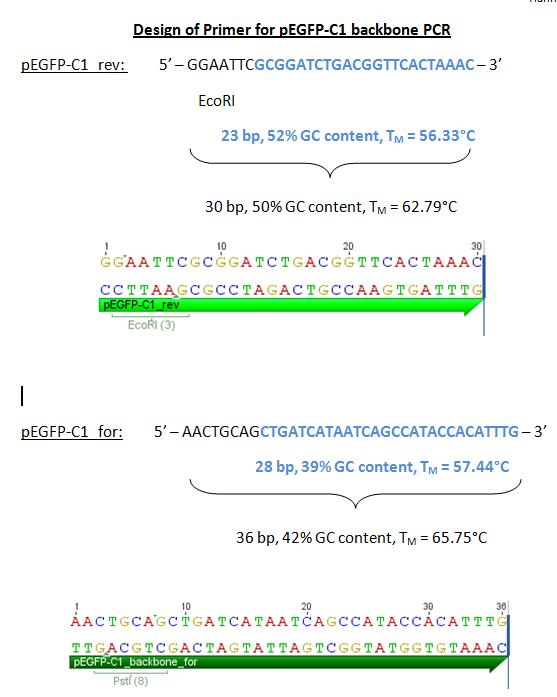
92. labday 17.08.2010
Overview of the state-of the art plasmid preparation
Investigator: Bea
Comments: In order to obtain a overview (more or less plausible ;-)) I sorted out how the final plasmid need to be constructed. In the picture you can see the most important plasmids which we will need for the retargeting approaches starting tomorrow.
[http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/index.php/Laborjournal top of page]
Further pSB1C3_SDM_SspI_Bla14FM sequencing
Investigators: Bea, Patrick
Sequencing results: there is no BLA in clone 4. Clone 5 showed a flawless BLA so it will be used for the next site directed mutagenesis.
pSB1C3_SDM_SspI_Bla14FM clone 5 SDM (P222)
Investigator: Patrick
Intention: remove PvuII restriction site
PCR program:
- 95°C 2 min (1x)
- 95°C 20s, 60°C 10 s, 68°C 30 s (30s/kb) (18x)
- 68°C 5 minutes
| Ingredients | Volume |
| 10x reaction buffer | 2,5 |
| DNA template ( about 10 ng) | 1 µl diluted P222 |
| forward primer: | 0,58 µl O108 (pSB1C3 PvuII rev) |
| reverse primer: | 0,59 µl O109 (pSB1C3 PvuII for) |
| DMSO (2%) | 0,5 |
| dNTP Mix from the kit | 0,5 µl |
| QuickSolution Reagent | 0,75 µl |
| Quickchange Lightning Enzyme (1.25U) | 0,5 µl |
| H2O | 18,08 µl |
| Total volume | 25 µl |
Further procedure including the transformation was performed according to the standard protocol.
Sent for sequencing
- pMA_RC (P222) sent for sequencing with O41 (Cap 4000 for) to check if the EaglII restiction site was successfully changed to PvuII.
- pAAV_RC (P158) sent for sequencing with GATC_std_SK, Rep_1250 for (O35), GATC_std_UP-2 to check if all deleted restriction sites are present (PstI, BamHI, SalI).
Investigators: Bea, Patrick
Cloning of pMA_RC-Insertparts in pAAV_RC 1.2 SDM SalI
Investigator: Jessica
P211 will be cloned in the P158 vector that contains 4 mutation (BamHI, SalI, PstI 320 + 4073)
Digestion:
| components | pMA_RC_Insertparts | pAAV_RC 1.2 SDM SalI |
| DNA | 5 | 4 |
| BSA (10x) | 1 | 1 |
| Buffer 3 (10x) | 1 | 1 |
| Enzyme1 BstEII | 0,5 | 0,5 |
| Enzyme2 SwaI | 0,5 | 0,5 |
| H2O | 2 | 3 |
| Total volume | 10 | 10 |
- 1. digestion with SwaI at 25°C for 1,5h
- 2. digestion with BstEII at 60°C for 1,5h
Preparation of gel:
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at Volt, running time: minutes
Expected sizes of constructs:
- pMA_RC-Insertparts: 493bp, 3287bp
- pAAV_RC 1.2 SDM SalI: 6839bp, 478bp
The corresponding bands were cut out and Gel-Extraction was performed according to protocol.

concentrations measured via NanoDrop:
- pMA_RC-Insertparts: 5,9 ng/µl
- pAAV_RC 1.2 SDM SalI:8,6 ng/µl
Ligation
Volume insert: 1,95 µl
Volume vector: 6,08 µl
Trafo was prepared with BL21
Repetition: Biobrick production of Rep 68
Investigator: Christian L. , Stefan
Aim of the experiment:
We want to produce biobricks from Rep 68.
- Plasmid used as template:
Rep_68_ex (p224): c = 350,29 ng/µl
- Primer used:
For Rep_68_ex: Praefix_68_78_ex & Suffix_40_68_ex
- PCR:
(was performed following the standard protocol)
| Ingredients | Volume Rep68 / µl |
| 5x Phusion HF buffer | 10 |
| 10 mM dNTP mix | 1 |
| forward primer: | 2,5 |
| reverse primer: | 2,5 |
| DNA Template | 3 |
| DMSO (2%) | - |
| Phusion Polymerase | 0,5 |
| H2O | 30,5 |
| Total volume | 50 |
PCR program:
| PCR Program | temperature/ °C | Time |
| 1 | 98 | 1min |
| 2 | 98 | 15s |
| 3 | 63 | 25s |
| 4 (step 2-4 8x) | 72 | 24s |
| 5 | 98 | 15s |
| 6 | 68 | 25s |
| 7 (step 5-7 17x) | 72 | 24s |
| 8 | 72 | 5min |
| Hold | 4°C |
Used agarose gel: 0,5 g Agarose,50 ml TBE (1%), 3 µl GELRED , at 115 Volt, running time:45 minutes
Gel extraction was perfomed according to protocol.
comment: Elution was put in 4°C room. Work will be continued tomorrow.
Transduction of HT1080 cells
Transduction plan
Investigator: Adrian, Kerstin
| A | control no cells | 300 µl virus (40 µg YFP) | 300 µl virus (20 µg YFP) |
|---|---|---|---|
| B | control no virus | 500µl P38 (pAAV2_mVenus) | 1000µl P38 (pAAV2_mVenus) |
| A | control no cells | 300 µl virus (40 µg YFP) | 300 µl virus (20 µg YFP) |
|---|---|---|---|
| B | control no virus | 500µl P38 (pAAV2_mVenus) | 1000µl P38 (pAAV2_mVenus) |
| A | control no cells | 300 µl virus (40 µg YFP) | 300 µl virus (20 µg YFP) |
|---|---|---|---|
| B | control no virus | 500µl P38 (pAAV2_mVenus) | 1000µl P38 (pAAV2_mVenus) |
| A | control no cells | 300 µl virus (40 µg YFP) | 300 µl virus (20 µg YFP) |
|---|---|---|---|
| B | control no virus | 500µl P38 (pAAV2_mVenus) | 1000µl P38 (pAAV2_mVenus) |
| A | control no cells | 150 µl virus (TKGMK clone 1) | 180 µl virus (TKGMK clone 1) |
|---|---|---|---|
| B | control no virus | 200 µl (TKGMK clone2) | 200µl (no GOI) |
Midi-Prep of pAAV_igEM_mVenus_YFP and dsAAV_CMV_EGFP
Investigators: Chris, Anna
Comment: Midi-Preps of dsAAV_CMV_EGFP (p31, B24) and pAAV_iGEM_mVenus_YFP (p39, B34)
The Midi-Preps were performed according to the standard protocol yielding the following concentrations:
| plasmid-no. | p31 | p39 |
| concentration (ng/µl) | 539,05 | 162,37 |
BioBrick assembly for pSB1C3_lITR_CMV_beta-globin_YFP_rITR
investigator:Achim
- This construct without the hGH polyadenylation sequence will be used to determine wether/how much hGH affects translation of the virus gene.
Vector:
- pSB1C3_lITR_CMV_ß_YFP (P209): c=274,86 ng/µl
Insert:
- pSB1C3_rITR (P172/P191): c=73,3 ng/µl
| components | Vector | Insert |
| DNA | 3,6 | 20,5 |
| BSA (10x) | 3 | 3 |
| Buffer 4 (10x) | 3 | 3 |
| Enzyme 1 | 1(SpeI) | 1(XbaI) |
| Enzyme 2 | 1(PstI) | 1 (PstI) |
| H2O | 18,4 | 1,5 |
| Total volume | 30 | 30 |
- 1% Agarose gel
- 3 µl Gelred
- 7 µl DNA-Ladder-Mix
- 115 Volt, running time: 30 minutes
The Insert containing the right ITR was cut out after 30 minutes, the Vector was cut out after 45 minutes
| Sample/µl | Expected size/bp |
| Vector | 4100 |
| Insert | 150 |
- weight of insert gel extract: 170 mg
- weight of vector gel extract: 200 mg
Nanodrop
- insert : 11,4 ng/µl
- vector : 26,7 ng/µl
Ligation
- vector: 6,36 µl
- insert: 1,64 µl
Transformation
BioBrick assembly for pSB1C3_lITR_pTERT
Investigator:Anissa
- Construct with the pTERT tumor specific promoter
Vector:
- pSB1C3_lITR (P196): 174 ng/µl
Insert:
- pSB1C3_phTERT (P177): c=312 ng/µl
| components | Vector | Insert |
| DNA | 5,75 | 6,41 |
| BSA (10x) | 1,5 | 1,5 |
| Buffer 4 (10x) | 1,5 | 1,5 |
| Enzyme 1 | 1(SpeI) | 1(XbaI) |
| Enzyme 2 | 1(PstI) | 1 (PstI) |
| H2O | 4,25 | 3,59 |
| Total volume | 15 | 15 |
- 1% Agarose gel
- 3 µl Gelred
- 7 µl DNA-Ladder-Mix
- 115 Volt, running time: 45 minutes
| Sample/µl | Expected size/bp |
| Vector | 2200 |
| Insert | 500 |
- weight of insert gel extract: 350 mg
- weight of vector gel extract: 250 mg
Nanodrop
- insert : 17,7 ng/µl
- vector : 27,6 ng/µl
Ligation
- vector: 5,57 µl
- insert: 2,43 µl
Transformation
Picking clones of pSB1C3_lITR_CMV_(betaglobin)_mVenus_hGH_rITR, EGFP_C1 and pMA_T_affibody
Investigator: Anna
| Plasmid | Resistance | Bacterial strain |
| pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR° | Chloramphenicol | XL1 blue |
| pSB1C3_lITR_CMV_mVEnus_hGH_rITR° | Chloramphenicol | XL1 blue |
| EGFP_C1 | Kanamycin | |
| pMA_T_affibody | Ampicillin |
°: 3 clones each
Clones were inoculated in 10 mL DYT medium containing 10 µL of antibiotics and were put in 37°C room on shaker.
Trafo evaluation of SDM PstI in CD and inoculation
Investigator: Kira
The plate contained lots of colonies and 2 of them were picked and inoculated into DYT+Amp
93. labday 18.08.2010
Continuation: pSB1C3_SDM_SspI_Bla14FM clone 5 SDM (P222)
There is a lawn on the plate although chloramphenicol was added. Some clones were picked and a new agar plate inoculated to hopefully receive some 'pickable' clones.
Additionally a new SDM was performed: PCR program:
- 95°C 2 min (1x)
- 95°C 20s, 60°C 10 s, 68°C 30 s (30s/kb) (18x)
- 68°C 5 minutes
| Ingredients | Volume |
| 10x reaction buffer | 2,5 |
| DNA template ( about 10 ng) | 1 µl diluted P222 |
| forward primer: | 0,58 µl O108 (pSB1C3 PvuII rev) |
| reverse primer: | 0,59 µl O109 (pSB1C3 PvuII for) |
| DMSO (2%) | 0,5 |
| dNTP Mix from the kit | 0,5 µl |
| QuickSolution Reagent | 0,75 µl |
| Quickchange Lightning Enzyme (1.25U) | 0,5 µl |
| H2O | 18,08 µl |
| Total volume | 25 µl |
All further steps including the transformation were performed according to the standard protocol. This time 2 agar plates were inoculated. One of them with the resuspended pellet and the other one with a 1:100 dilution of the resuspended pellet.
Investigator: Patrick
Mini Preps pEGFP_C1,pSB1C3_Rep40,pSB1C3_Rep78,pMA_T_Affibody,pSB1C3_SspIdel_BLA clone 2+3,pSB1C3_AAP, pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR,pSB1C3_lITR_CMV_mVenus_hGH_rITR,pSK_CD_SDMPstI
Investigator: Jessica
Preps and test digestions were made:
pSK_CD_SDMPstI
- Given Plasmid-Number: P248; DNA concentration: 302,7 ng/µL ;
- Given Plasmid-Number: P249; DNA concentration: 267,0 ng/µL ;
- P248, P249 and P223 as control digested with PstI --> no satisfying result,control was not cut but samples two times... sent for sequencing with primer: GATC_std_T7minus1
pSB1C3_Rep40
- Given Plasmid-Number: P231 (clone3); DNA concentration: 234,00 ng/µL
- Given Plasmid-Number: P232 (clone2); DNA concentration: 234,96 ng/µL
- Given Plasmid-Number: P233 (clone1); DNA concentration: 237,12 ng/µL
pSB1C3_Rep78
- Given Plasmid-Number: P234 (clone2); DNA concentration: 277,56 ng/µL
- Given Plasmid-Number: P235 (clone1); DNA concentration: 294,19 ng/µL
- Given Plasmid-Number: P236 (clone3); DNA concentration: 378,87 ng/µL
pSB1C3_AAP
- Given Plasmid-Number: P239 (clone1); DNA concentration: 219,82 ng/µL
- Given Plasmid-Number: P240 (clone2); DNA concentration: 220,45 ng/µL
- Given Plasmid-Number: P241 (clone3); DNA concentration: 209,57 ng/µL
@Jessica: What are these plasmid numbers?? (Bea)
- Given Plasmid-Number: P230; DNA concentration: 367,36 ng/µL ;
- Given Plasmid-Number: P242; DNA concentration: 202,09 ng/µL ;
- Given Plasmid-Number: P243; DNA concentration: 369,34 ng/µL ;
- Given Plasmid-Number: P245; DNA concentration: 207,12 ng/µL ;
- Given Plasmid-Number: P246; DNA concentration: 198,35 ng/µL ;
- Given Plasmid-Number: P247; DNA concentration: 212,22 ng/µL ;
Test digestion
- buffer used: 4 ; Restriction-enzymes used: Enzyme XbaI ; Enzyme SpeI
- Plasmid
Components P231-233 Volume/µL P234 Volume/µL P235 Volume/µL 236 Volume/µL P239-242 Volume/µL DNA 4 3.5 3 2.5 4 BSA (10x) 2 2 2 2 2 Buffer 4 (10x) 2 2 2 2 2 SpeI 0.5 0.5 0.5 0.5 0.5 XbaI 0.5 0.5 0.5 0.5 0.5 H2O 11 11,5 12 12.5 11 Total volume 20 20 20 20 20
Comment: Test digestion of P231-233 and P239-242 contained equal volumes of ingredients. Therefore the charts were simplyfied.
Comment: P231 and P239 will be sent for sequenzing tomorrow. We picked for additional clones of Rep78 for Mini-Prep and test digestion tomorrow.
Continuation of Repetition: Biobrick production of Rep 68 (17.08.2010)
Investigator: Christian L. , Stefan
Aim of the experiment:
We want to produce biobricks from Rep 68.
- Vector: name: pSB1C3_CFP P51.1
- Insert: name: Rep68_SDM_PstI clone 1 P224
- buffer used: 4
- Restriction-enzymes used:
- XbaI
- SpeI
- DNA concentration (P51.1): 132,1 ng/µl
- DNA concentration (P224): 350.29 ng/µl
Digestion of plasmid backbone and PCR product:
| components | volume of pSB1C3_CFP /µl | volume of Rep68_SDM_PstI clone 1 /µl |
| DNA | 10 | 29 |
| BSA (10x) | 2 | 4 |
| Buffer 4 (10x) | 2 | 4 |
| SpeI | 1 | 1 |
| XbaI | 1 | 1 |
| H2O | 4 | 1 |
| Total volume | 20 | 40 |
Incubation at 37°C, 2 h
Dephoshorylation of Vector pSB1C3_CFP after digestion:
- add 2 µl of 10x Antarctic Phopshatase Reaction Buffer to DNA cut with XbaI and SpeI
- add 1 µl of Antarctic Phosphatase and mix
- incubate for 60 minutes at 37°C
Gelextration of pSB1C3
Gelextration of pSB1C3 was performed according to standard protocol
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time:45
PCR Purification
Purification of Rep68 was performed according to the standard protocol
Concentration measured: 16,4 ng/µl
Ligation
For the ligation 1µl T4 buffer (10x) and 1µl T4 ligase were used.
| vector /µl | insert /µl | |
| pSB1C3 + Rep68 | 3,9 | 4,1 |
Incubation at room temperature for 45 minutes.
Transformation:
The transformation was done following the standard protocol using XL1b cells.
Transfection of AAV293 cells
Investigator: Adrian, Kerstin
- 2x106 cells were seeded (instead of 3x106) --> hopefully better virus-production
- used plasmids:
- P50 c= 429 ng/µl (RC)
- P64 c= 500 ng/µl (pHelper)
- P229 c= 162 ng/µl (GOI=YFP)
- P228 c= 540 ng/µl (GOI=eGFP)
- P158a c= 1800ng/µl (RC mutant)
- P158a c= 1770ng/µl (RC mutant)
- 1. plate: Lipo-Transfection
- 2. plate: Lipo-Transfection
- 3. plate: 3,3 µg (RC), 3,3 µg (pHelper), 3,3 µg (YFP)
- 4. plate: 10 µg (RC), 10 µg (pHelper), 3,3 µg (YFP)
- 5. plate: 10 µg (RC mutant P158a), 10 µg (pHelper), 3,3 µg (YFP)
- 6. plate: 10 µg (RC mutant P158b), 10 µg (pHelper), 3,3 µg (YFP)
- 7. plate: 3,3 µg (RC mutant P158a), 3,3 µg (pHelper), 3,3 µg (YFP)
- 8. plate: 3,3 µg (RC mutant P158b), 3,3 µg (pHelper), 3,3 µg (YFP)
- 9. plate: 10 µg (RC), 10 µg (pHelper), 3,3 µg (eGFP)
- 10. plate: 10 µg (RC), 10 µg (pHelper), 10 µg (eGFP)
- 11. plate: 10 µg (RC), 10 µg (pHelper), 20 µg (eGFP)
- 12. plate: 10 µg (RC), 10 µg (pHelper), 40 µg (eGFP)
Seeding Ht1080
Investigators: Adrian, Kerstin
Cloning of pMA_T_affibody and pSB1C3_CFP
Comment: pMA_T_affibody (p237) from Retrafo and pSB1C3_CFP (p51.5) were digested with EcoRI and PstI.
Investigator: Anna
Vector:
- pSB1C3_CFP (p51.1): c= 132,1 ng/µl
Insert:
- pMA_T_affibody (p237): c= 233,96 ng/µl
| components | Vector p51.1 | Insert p237 |
| DNA | 9 | 8 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| Enzyme EcoRI | 1 | 1 |
| Enzyme PstI | 1 | 1 |
| H2O | 5 | 6 |
| Total volume | 20 | 20 |
- Gelextraction of pSB1C3_RFC25_CFP:
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 120 Volt, running time:55
4µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas)
- c(pSB1C3)= 5,4 ng/µl
- c(affibody)= 5,6 ng/µl
- Ligation of PCR products and vector:
For the Ligation 1µl T4 buffer (2x) and 1µl T4 ligase were used. Incubation time: 30 min
| vector /µl | insert /µl | |
| pSB1C3 + affibody | 5,91 | 2,09 |
- Transformation:
The transformation was done following the standard protocol using XL1 blue cells.
Result: Clones of pSB1C3_affibody will be picked tomorrow.
94. labday 19.08.2010
Continuation: pSB1C3_SDM_SspI_Bla14FM clone 5 SDM (P222)
Again there was a bacterial lawn on the agar plate. Therefore some clones from the overgrown agar plated were picked and diluted. The 1:10^4, 1:10^5, 1:10^6 dilution each were put onto an agar plate (37°C, over night). Additionally a transformation with XL1B cells from the remaining SDM DNA sample was performed and diluted, too. The 1:10^4 and the 1:10^6 dilution were plated. Hopefully there will be some 'pickable' clones tomorrow.
Investigator: Patrick
Cloning of pSB1C3_CFP and middlelinker
Investigator: Jessica
- buffer used: 4; Restriction-enzymes used: Enzyme 1: AgeI ; Enzyme 2: PstI ; Enzyme3: NgoMIV
- Plasmids:
- pSB1C3_CFP P51.2
- pGA14_middle linker P65
| Components | Mastermix | P51.2/µL | P65/µL |
| DNA | - | 9,9 | 10,8 |
| BSA (10x) | - | 2 | 2 |
| Buffer 4 (10x) | - | 2 | 2 |
| Enzyme 1 AgeI | - | 1 | - |
| Enzyme 2 PstI | - | 1 | 1 |
| Enzyme 3 NgoMIV | - | - | 1 |
| H2O | - | 4,1 | 3,2 |
| Total volume | - | 20 | 20 |
- Incubation: 90 minutes
Agarosegel
0.5 g Agarose, 50 ml TAE (1 %), 3 µL GELRED, at 115 Volt, running time: 45 minutes
concentrations measured via NanoDrop:
- SB1C3_CFP: 5,0 ng/µl
- middle linker:1,7 ng/µl
T4 Ligation
Volume insert: 1,24 µl
Volume vector: 6,76 µl
Trafo was prepared with XL1blue
Test digestion of pSB1C3_Rep78
Investigator: Chris L., Stefan
Preps and test digestions were made:
pSB1C3_Rep78
- Given Plasmid-Number: P252 (clone4); DNA concentration: 352,47 ng/µL
- Given Plasmid-Number: P253 (clone5); DNA concentration: 408,23 ng/µL
- Given Plasmid-Number: P254 (clone6); DNA concentration: 349,18 ng/µL
- Given Plasmid-Number: P255 (clone7); DNA concentration: 320,25 ng/µL
Test digestion
- buffer used: 4 ; Restriction-enzymes used: Enzyme XbaI ; Enzyme SpeI
- Plasmid
Components P252 Volume/µL P253 Volume/µL P254 Volume/µL 255 Volume/µL DNA 2.5 2 2.5 3 BSA (10x) 2 2 2 2 Buffer 4 (10x) 2 2 2 2 SpeI 0.5 0.5 0.5 0.5 XbaI 0.5 0.5 0.5 0.5 H2O 12.5 13 12.5 12 Total volume 20 20 20 20
Incubation time: 1 h; Incubation temperature: 37°
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , running time:5 minutes at 90 Volt and 40 minutes at 115 Volt
4µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas)
Sent for sequencing
Investigator: Chris L., Stefan
pSB1C3_Rep52 clone3 (P202) sent for sequencing with O162 (VF2). pSB1C3_Rep40 clone3 (P231), pSB1C3_AAP clone1 (P239), pSB1C3_Rep78 clone4 (P252) and pSB1C3_Rep78 clone7 (255) sent for sequencing with O162 (VF2) and O51 (VR2) pAAV_RC_ins-rep clone1 (P250) and pAAV_RC_ins-rep clone2 (P251) sent for sequencing with O19 (VP1 primer for pKEX reverse)
Test Digestion of P242, P243, P245, P246, P247
Investigator: Achim
P242: pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR clone1
P243: pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR clone2
P245: pSB1C3_lITR_CMV_mVenus_hGH_rITR clone1
P246: pSB1C3_lITR_CMV_mVenus_hGH_rITR clone2
P247: pSB1C3_lITR_CMV_mVenus_hGH_rITR clone3
(Left picture: Test digestion from 18.8., right picture: Repetition of test digestion from 19.8.)
Fragments sizes:
- P242/243: 1400/3400
- P245/P246/P247: 1400/2900
Bands are diffuse and size estimation is difficult because of strange marker behaviour, but at least the second digestion shows two distinct bands which look like they have the correct sizes. P242 and P245 were sent for sequencing.
Test digestion of pAAV_RC-ins-rep
Investigator: Chris L., Stefan
Preps and test digestions were made:
pAAV_RC-ins-rep
- Given Plasmid-Number: P250 (clone1); DNA concentration: 467,5 /µL
- Given Plasmid-Number: P251 (clone2); DNA concentration: 108,4 ng/µL
Test digestion
- buffer used: 3 ; Restriction-enzymes used: Enzyme SwaI ; Enzyme BstEII
- Plasmid
Components P250 Volume/µL P251 Volume/µL DNA 1,7 7.3 BSA (10x) 1 1 Buffer 3 (10x) 1 1 SpeI 0.5 0.5 XbaI 0.5 0.5 H2O 5.3 - Total volume 10 10.3
Incubation time with SwaI: 60 minutes; Incubation temperature: 25° Incubation time with BstEII: 75 minutes; Incubation temperature: 60°
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , running time: 45 minutes at 115 Volt
4µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas)
Midiprep of dsAAV_CMV_EGFP and pAAV_iGEM_mVenus_YF
Investigators: Chris W.
Comment: Midi-Preps of dsAAV_CMV_EGFP (p31, B24) and pAAV_iGEM_mVenus_YFP (p39, B34)
The Midi-Preps were performed according to the standard protocol yielding the following concentrations:
| plasmid-no. | p31 | p39 |
| concentration (ng/µl) | 2420,46 & 2586,64 | 1148,07 & 978,99 |
Harvest viral particles and Transduction of HT1080 cells
Investigators: Adrian, Kerstin
| Plate | amount of cells |
| 1 | 100 000 |
| 2 | 200 000 |
| 3 | 400 000 |
| 4 | 500 000 |
| 5 | 800 000 |
| 6 | 1 000 000 |
| 7 | 1 200 000 |
| 8 | 1 500 000 |
| 9 | 1 750 000 |
| 10 | 1 750 000 not sure |
The cells were transfected with 40µg YFP, 3,3µg Rep/Cap and 3,3µg pHelper.
Comments: We want to find out, if the efficiency of virus production can be influenced by the amount of transfected cells.
Plate 1 YFP: 200.000 cells per well
| 1 | 2 | 3 | |
|---|---|---|---|
| A | control, no cells | 165µl virus (1) | 165µl virus (2) |
| B | control, no virus | 165µl virus (1) | 165µl virus (2) |
Plate 2 YFP: 200.000 cells per well
| 1 | 2 | 3 | |
|---|---|---|---|
| A | control, no cells | 165µl virus (3) | 165µl virus (4) |
| B | control, no virus | 165µl virus (3) | 165µl virus (4) |
Plate 3 YFP: 200.000 cells per well
| 1 | 2 | 3 | |
|---|---|---|---|
| A | control, no cells | 165µl virus (5) | 165µl virus (6) |
| B | control, no virus | 165µl virus (5) | 165µl virus (6) |
Plate 4 YFP: 200.000 cells per well
| 1 | 2 | 3 | |
|---|---|---|---|
| A | control, no cells | 165µl virus (7) | 165µl virus (8) |
| B | control, no virus | 165µl virus (7) | 165µl virus (8) |
Plate 5 YFP: 200.000 cells per well
| 1 | 2 | 3 | |
|---|---|---|---|
| A | control, no cells | 165µl virus (9) | 165µl virus (10) |
| B | control, no virus | 165µl virus (9) | 165µl virus (10) |
New ladder mix and testing
Investigator: Jessica
new Generuler ladder mix from Fermentas arrived
one tube (containing 100µl solution) was prepared with 100µl loading dye and 400 µl miliporewater
two test-gels were prepared:
- 1% TAE agarose gel with samples of
- 0,5 µl
- 1,0 µl
- 2,0 µl
- 3,0 µl
- 4,0 µl
- 5,0 µl
- 6,0 µl
- 7,0 µl
- 8,0 µl
- 9,0 µl
- 10,0 µl
- sample P250
- 1% TAE agarose gel containing 20% H2O with samples of ( 20% H2O was an idea I have read in a journal but it doesn't seem like a good idea
- 0,5 µl
- 1,0 µl
- 2,0 µl
- 3,0 µl
- 4,0 µl
- 5,0 µl
- 6,0 µl
- 7,0 µl
- 8,0 µl
- 9,0 µl
- 10,0 µl
- sample P251
Comments: an example of these laddervolumes is patches at the bench (right up)
SDM of CD NgoMIV
Investigator:Kira
Comment: Lightning kit was used for quickchange reaction
| component | Volume/µl |
| DNA: (1:100) | 3,3 |
| 10x reaction buffer: | 2,5 |
| primer forward NgoMIV (1:10) | 0,55 |
| primer reverse NgoMIV (1:10) | 0,55 |
| dNTP | 0,5 |
| DMSO | 0 |
| QuickSolution Reagent | 0,75 |
| H2O | 16,35 |
| Lightning Polymerase | 0,5 |
PCR
| segment | cycles | temperature | time |
| 1 | 1 | 95 | 2min |
| 2 | 18 | 95 | 20sec |
| 60 | 10 sec | ||
| 68 | 2 min15 sec | ||
| 3 | 68 | 5 min |
sample was digested with 1 ul DpnI for 10 min at 37°C
Transformation was performed according to the standard protocol.
Picking clones of pSB1C3_affibody and pSB1C3_Rep68
Investigator: Anna
| Plasmid | Resistance | Bacterial strain |
| pSB1C3_affibody° | Chloramphenicol | XL1 blue |
| pSB1C3_Rep68°° | Chloramphenicol | XL1 blue |
°: 2 clones
°°: 3 clones each
Clones were inoculated in 10 mL DYT medium containing 10 µL of antibiotics and were put in 37°C room on shaker.
95. labday 20.08.2010
Sequence analysis of sequencing reads prepared yesterday (19.08.2010)
Investigator: Bea
GENERAL COMMENT: pSB1C3 {Rep 52 and Rep 78{} were sent for sequencing BUT they cannot be used for subcloning the ordered rep protein because in the produced BioBricks the restriction enzyme needed for subcloning (SwaI) is NOT present in the BioBricks pSB1C3_Rep52 and rep78.
Comments: Sequence analysis of pAAV_RC containing the subcloned "rep" (ordered)
- Primer used: VP1 primer fro pKex
- Plasmid sequenced: P250
- Sequence sample: CL-9
- Stored in folder:
Next steps: xy
Comments: Sequence analysis of pAAV_RC containing the subcloned "rep" (ordered)
- Primer used: VP1 primer fro pKex
- Plasmid sequenced: P251
- Sequence sample: CL-10
- Stored in folder:
Next steps: xy
Comments: Sequence analysis of pSB1C3_Rep40
- Primer used: VR-2
- Plasmid sequenced: P231
- Sequence sample: CL-2
- Stored in folder: DKFZ Rep40ex
Sequence read is good (prefix is ok) but there was found a mutation in Rep52 protein: G2886A (nucleotides) --> V337I.
Next steps: xy
Comments: Sequence analysis of pSB1C3_AAP
- Primer used: VR-2
- Plasmid sequenced: P239
- Sequence sample: CL-3
- Stored in folder: DKFZ AAP
xy
Next steps: xy
[http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/index.php/Laborjournal top of page]
Continuation: pSB1C3_SDM_SspI_Bla14FM clone 5 SDM (P222)
Investigator: Patrick
The XL1B transformation didnt work or was too much diluted because there were no clones on the plates. The dilution steps of the transformed XL10-Gold cells worked quite well. Now there are 'pickable' clones.

[http://www.molbiotech.uni-freiburg.de/iGEM/wiki2010/index.php/Laborjournal top of page]
Subcloning cap from pMA_RepCap into pAAV_RC which contains four mutations and the synthesized Rep part
Investigator: Patrick
pAAV_RC_ins-rep clone 1 (P250) and pMA_RepCap Vector_SDM_InsPvuII clone 1 (P211) were digested with PspLI and BspMI according to the standard protocol (Subcloning_cap_into_pAAV_RC).
Expected size of the bands:
- Insert: 920 bp (P211)
- Vector: 6414 bp (P250)
The gelextraction yielded the following DNA concentrations:
- P 211 : 2,45 ng/µl
- P 250 : 11,15 ng/µl
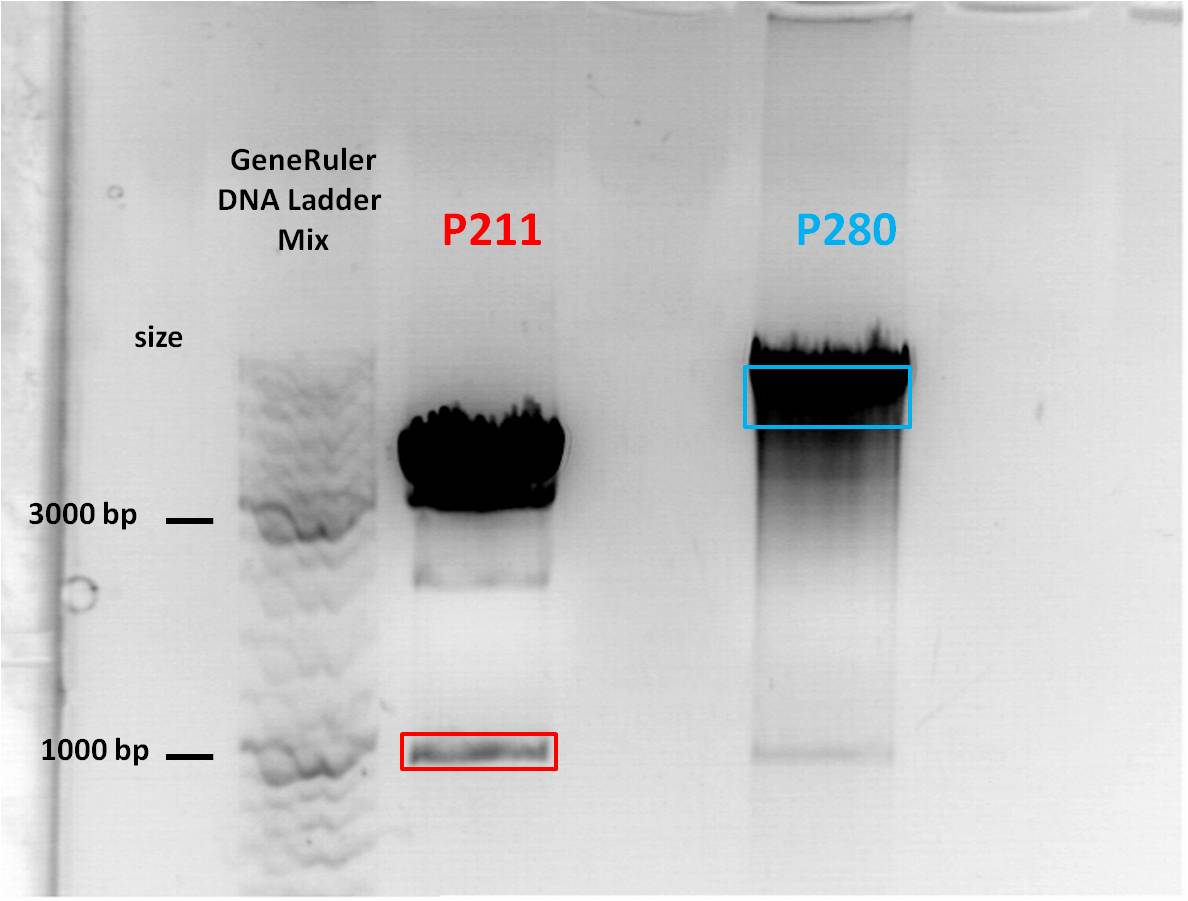
The marked bands were cut out and the DNA was extracted. The ligation was performed accodring to the standard prtocol with a total volume of 10 µl: 2,67 µl vector backbone(P250), 5,33 µl Insert following a transformation according to the standard protocol with XL1B cells.
Mini-Preps of pSB1C3_Rep68_ex and pSB1C3_ZEGFR:1907
Investigator: Chris L.,Stefan
-
Glycerol stocks were prepared:
- B218 = pSB1C3_Rep68_ex clone1
- B219 = pSB1C3_Rep68_ex clone2
- B220 = pSB1C3_Rep68_ex clone3
- B221 = pSB1C3_ZEGFR:1907 clone 1
- B222 = pSB1C3_ZEGFR:1907 clone 2
- B218 = pSB1C3_Rep68_ex clone1 c=286.72 ng/µl
- B219 = pSB1C3_Rep68_ex clone2 c=277.06 ng/µl
- B220 = pSB1C3_Rep68_ex clone3 c=298.18 ng/µl
- B221 = pSB1C3_ZEGFR:1907 clone 1 c=174.20 ng/µl
- B222 = pSB1C3_ZEGFR:1907 clone 2 c=162.48 ng/µl
Mini-Prep following the standard protocol
Test digestion
- buffer used: 4 ; Restriction-enzymes used: XbaI ; SpeI ; EcoRI ; PstI
| Components | P264 Volume/µL | P265 Volume/µL | P266 Volume/µL | P267 Volume/µL | P268 Volume/µL |
| DNA (800 ng) | 2.8 | 2.9 | 2.7 | 4.6 | 4.9 |
| BSA (10x) | 1 | 1 | 1 | 1 | 1 |
| Buffer 4 (10x) | 1 | 1 | 1 | 1 | 1 |
| SpeI | 0.5 | 0.5 | 0.5 | - | - |
| XbaI | 0.5 | 0.5 | 0.5 | - | - |
| PstI | - | - | - | 0.5 | 0.5 |
| EcoRI | - | - | - | 0.5 | 0.5 |
| H2O | 4.2 | 4.1 | 4.3 | 2.4 | 2.1 |
| Total volume | 10 | 10 | 10 | 10 | 10 |
Incubation time: 1 h; Incubation temperature: 37°
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , running time:5 minutes at 90 Volt and 40 minutes at 115 Volt
4µl loading dye (6x) for the sample, Marker: GeneRuler ladder mix (Fermentas)

Comment:pSB1C3_Rep68_ex clone 3 (P266) sent for sequencing with O162 (VF2) and O51 (VR2). pSB1C3_ZEGFR:1907 clone 1 (P267) sent for sequencing with O51 (VR2).
Hybridisation of the oligos for NLS and cloning NLS into pSB1C3
Investigator: Anissa
- Hybridisation: program: Origami 1
Hybridisation was peformed according the standard protocol - Digestion of pSB1C3_CFP
components Vector /µl DNA 6,62 BSA (10X) 1,5 Buffer 4 1,5 Enzyme EcoRI 1 Enzyme PstI 1 H2O 3,38 Endvolume 15 - Gel:After digestion, the sample was loaded on a 1% agarose-gel.
- Gelextraction: After the gel run, the vector backbone pSB1C3 was cut and purified according to the standard protocol
- Ligation: the original concentration of the hybridisation was 209,7 ng/µl, for the ligation a 1:10 dilution was made
components Volume /µl Concentration/ ng/µl Insert (NLS) 6,79 2,097 Vector (pSB1C3_cut) 1,21 8,3 - Transformation: Trafo into XL1 blue was performed according the standard protocol
Seeding HT1080 cells
Investigator: Kerstin, Adrian
- Seeding HT1080 cells for transduction with viral stocks from 18.08.
- We harvest four T75 flask (two of them were used three times (3 passages)) and seed them into eight 6er dishes (200.000 cells per well)
PCR and ligation for biobrick-production of pCerulean
Investigator: Christian L. , Stefan
DNA template: pEGFP_C1 (P230)
C=367,4 ng/µl
PCR:
(was performed following the standard protocol)
| Ingredients | Volume / µl |
| 5X Phusion HF buffer | 10 |
| 10 mM dNTP mix | 1 |
| forward primer: O163 (pEGFP-C1_rev) | 2.5 |
| reverse primer: O164 (pEGFP-C1_for) | 2.5 |
| DNA Template(1:1000) | 3 µl |
| DMSO (2%) | 1 |
| Phusion Polymerase | 0,5 |
| H2O | 29.5 |
| Total volume | 50 |
PCR program:
| PCR Program | temperature/ °C | time/ sec. |
| 1 | 98 | 60 |
| 2 | 98 | 15 |
| 8x | 59 | 25 |
| 3 | 72 | 60 |
| 4 | 98 | 15 |
| 17x | 66 | 25 |
| 5 | 72 | 60 |
| 1x | 72 | 5 |
| Hold | 4 | - |
Comments: Errors in table, please correct!!! (TB)
Gelextraction of PCR product
Gelextraction was performed according to the standard protocol of the PCR product.
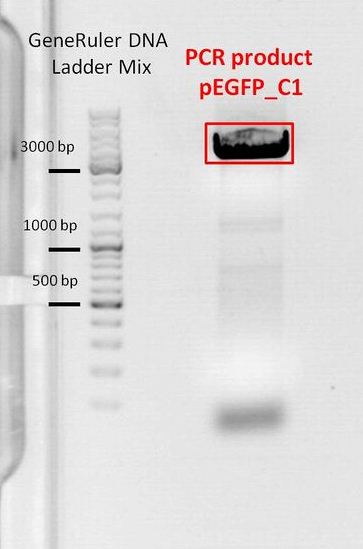
Comment: 0,4 g Agarose,50 ml TAE (0,8%), 3 µl GELRED , at 115 Volt, running time: 45 minutes
Digestion of PCR product and vector:
| components | PCR product /µl | vector /µl |
| DNA | 10 | 29 |
| BSA (10x) | 2 | 2 |
| Buffer 4 (10x) | 2 | 2 |
| Enzyme EcoRI | 1 | 1 |
| Enzyme PstI | 1 | 1 |
| H2O | 4 | 5 |
| Total volume (e.g. 15,20,25,30 µl) | 20 | 40 |
Purification of the PCR product with PCR purification was made according to the standard protocol.
Gelextraction of CFP
Gelextraction was performed according to the standard protocol.
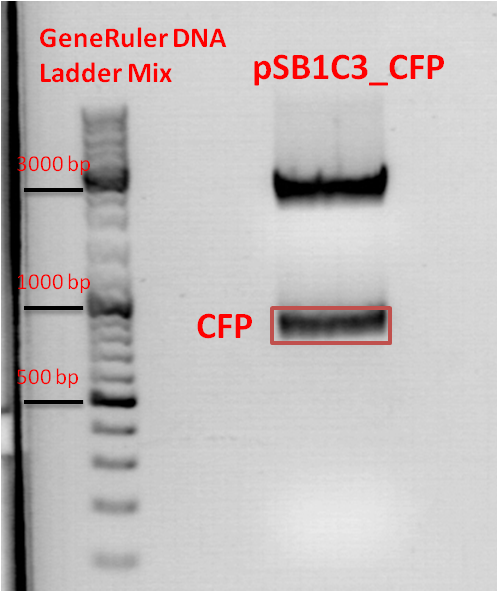 >
>
Comment: 0.5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 115 Volt, running time: 45 minutes
Ligation
The Ligation was performed as following:
- Vector Volume: 4,1 µl
- Insert Volume: 3,9 µl
- 1µl T4-Ligase buffer (10x)
- 8µl (Vector + Insert) mix
- 1µl T4-Ligase
Incubation for 45 minutes at room temperature
Transformation:
performed according to the standard protocol (XL1B). The cells were plated on a agar plate with Kanamycin.
Biobrick production of VP1_up
Investigator: Achim
DNA template: pKex_VP1 (P27)
C=99,3 ng/µl
PCR:
(was performed following the standard protocol)
| Ingredients | Volume / µl |
| 5X Phusion HF buffer | 10 |
| 10 mM dNTP mix | 1 |
| forward primer: O89 (Präfix_VP1_ex_RFC25_for) | 2.5 |
| reverse primer: O167 (pEGFP-C1_for) | 2.5 |
| DNA Template(1:100) | 1 µl |
| DMSO (2%) | - |
| Phusion Polymerase | 0,5 |
| H2O | |
| Total volume | 50 |
PCR program:
| PCR Program | temperature/ °C | time/ sec. |
| 1 | 98 | 60 |
| 2 | 98 | 15 |
| 8x | 60 | 25 |
| 3 | 72 | 15 |
| 4 | 98 | 15 |
| 17x | 66 | 25 |
| 1x | 72 | 15 |
| Hold | 4 | - |
Digestion of PCR product and vector:
| components | PCR product /µl | vector /µl |
| DNA | 20 | 5,2 |
| BSA (10x) | 3 | 2 |
| Buffer 4 (10x) | 3 | 2 |
| Enzyme NgOMIV | 1 | 1 |
| Enzyme SpeI | 1 | 1 |
| H2O | 2 | 8,8 |
| Total volume (e.g. 15,20,25,30 µl) | 30 | 20 |
Ligation with T4-Ligase and transformation with XL1B were carried out. Clones have to be picked tomorrow.
BioBrick assembly for pSB1C3_Affibody_Linker and pSB1C3_YFP_His-Tag
Investigators: Volker
The aim of this experiment is to assemble the affibodies with the the five linkers and the YFP with the His-Tag. All cloning is done according to RFC 25 and will be used to test the affibody if it really binds the EGF-receptor, for the N-terminal fusions and to supply it to other groups in the registry.
This experiment went wrong on Saturday 21.08. and was repeated on Sunday 22.08. and gave the following results. Results from the first experiment are not shown but can be senn in the written Labjournal.
Digestion
The following digestions were performed:
| components | Affibody | Affibody | Affibody | Short Linker | Middle Linker | Long Linker | SEG Linker | GSAT Linker | YFP | His-Tag |
| DNA | 8.6 µl | 8.6 µl | 8.6 µl | 7.7 µl | 8.2 µl | 8.0 µl | 8.1 µl | 8.4 µl | 5.4 µl | 14 µl |
| BSA (100x) | 2µl | 2µl | 2µl | 0µl | 0µl | 0µl | 0µl | 0µl | 2µl | 0µl |
| Buffer (10x) | 2µl | 2µl | 2µl | 2µl | 2µl | 2µl | 2µl | 2µl | 2µl | 2µl |
| Enzyme 1 (1µl each) | PstI | PstI | PstI | PstI | PstI | PstI | PstI | PstI | EcoRI | EcoRI |
| Enzyme 2 (1µl each) | AgeI | AgeI | AgeI | NgoMIV | NgoMIV | NgoMIV | NgoMIV | NgoMIV | AgeI | NgoMIV |
| H2O | 5.4 µl | 5.4 µl | 5.4 µl | 8.3 µl | 7.8 µl | 8.0 µl | 7.9 µl | 7.6 µl | 8.6 µl | 2.0 µl |
| Total volume | 20µl | 20µl | 20µl | 20µl | 20µl | 20µl | 20µl | 20µl | 20µl | 20µl |
Incubation: 2 h
Preparative Agarose-Gel
1 g Agarose, 100 ml TAE (1%), 6 GELRED (3-6µl), at 120Volt, running time: 130 minutes
| Lane | Sample | Sample/µl] | Loading dye (5x/6x)/µl | Expected size 1 (Geneious) | Expected size 2 (Geneious) |
|---|---|---|---|---|---|
| 1 | Affibody | 20 µl | 4 µl | bp | bp |
| 2 | Affibody | 20 µl | 4 µl | bp | bp |
| 3 | Affibody | 20 µl | 4 µl | bp | bp |
| 10 | His-Tag | 20 µl | 4 µl | bp | bp |
2 g Agarose, 100 ml TAE (2%), 6 GELRED (3-6µl), at 120 Volt, running time: 130 minutes
| Lane | Sample | Sample/µl] | Loading dye (5x/6x)/µl |
|---|---|---|---|
| 4 | Short Linker | 20 µl | 4 µl |
| 5 | Medium Linker | 20 µl | 4 µl |
| 6 | Long Linker | 20 µl | 4 µl |
| 7 | SEG Linker | 20 µl | 4 µl |
| 8 | GSAT Linker | 20 µl | 4 µl |
| 9 | YFP | 20 µl | 4 µl |
Gel extraction
| Sample | Weight | Volume | Concentration |
| Affibody | 0.4g | 1200µl | 29.78 |
| Affibody | 0.44g | 1320µl | 24.86 |
| Short-Linker | 0.27 g | 810µl | 0.22 |
| Middle-Linker | 0.38 g | 1140µl | 3.67 |
| Long-Linker | 0.35 g | 1050µl | 1.31 |
| SEG-Linker | 0.34 g | 1020µl | 2.13 |
| GSAT-Linker | 0.22 g | 660µl | 3.58 |
Ligation
| Sample | Insert | V(Insert) | Vector | V(Vector) |
| pSB1C3_Affibody_Short-Linker | Affibody | 1.06µl | Short-Linker | 6.94µl |
| pSB1C3_Affibody_Middle-Linker | Affibody | 5.25µl | Middle-Linker | 2.75µl |
| pSB1C3_Affibody_Long-Linker | Affibody | 2.93µl | Long-Linker | 5.07µl |
| pSB1C3_Affibody_SEG-Linker | Affibody | 2.46µl | SEG-Linker | 5.54µl |
| pSB1C3_Affibody_GSAT-Linker | Affibody | 3.41µl | GSAT-Linker | 4.59µl |
96. labday 21.08.2010
Transfection of AAV293, Harvest viral particles, Transduction of HT1080 and Fluorescent microscopy
Investigators: Adrian, Kerstin
The Motivation
Measure the Transduction efficiency from 19.8
The Motivation
Transfection of AAV293 cells from 19.8 (ten 10cm2 dishes with 2.000.000 cells each) with original standard protocoll!!! We want to try different 2xHBS Buffers (pH) and their influence on transfection efficiency!
The Plan
Requiered goods: 0,3 M CaCl2 , 2x HBS-Buffer , autoclaved millipore water, falcons, eppis
Working steps:
- Remove the three plasmids to be co-transfected (the recombinant pAAV expression plasmid or control plasmid, pAAV-RC, and pHelper) from storage at –20°C.
- Pipet 10 µg of each of the three plasmid DNA solutions into an eppi.
- Add 1ml of 0.3 M CaCl2 and mix gently
- Pipet 1ml of 2 × HBS into a 15-ml falcon.
- pipet the DNA/CaCl2 solution dropwise into the falcon (use the Pipetus).
- Apply the DNA/CaCl2/HBS suspension to the plate of cells in a dropwise fashion, swirling gently to distribute the DNA suspension evenly in the medium.
- Return the tissue culture plate to the 37°C room for 6 hours.
- At the end of the incubation period, remove the medium from the plate and wash cells with warm PBS, replace it with 10 ml of fresh DMEM growth medium.
- Return the plate to the 37°C incubator for an additional 72 hours.
We try 5 different 2xHBS buffers:
- pH: 7.06
- pH: 7.08
- pH: 7.10
- pH: 7.12
- pH: 7.14
- 1. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.06
- 2. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.08
- 3. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.10
- 4. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.12
- 5. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.14
- 6. Plate: 10µg of Rep/Cap, pHelper and GOI=TKGMK; pH(2xHBS)= 7.08
- 7. Plate: 10µg of Rep/Cap, pHelper and GOI=TKGMK; pH(2xHBS)= 7.10
- 8. Plate: 10µg of Rep/Cap, pHelper and GOI=mVenus; pH(2xHBS)= 7.14
The transfected cells were seeded at 19.8 (2.000.000 cells per 10 cm2)
Used Plasmids: Standart protocol: 10µg of each plasmid
- GOI: mVENUS => P262, conc:1148ng/µl; used amount: 8,71µl
- GOI: TKGMK => P82b, conc:1500ng/µl; used amount: 6,67µl
- pHELPER => P64, conc:503ng/µl; used amount: 20µl
- RepCap => 10.8: 1348ng/µl; used amount: 7,41µl
- Harvest viral paricles from 18.08.2010 and transduction of HT1080 cells following the standard protocol
- Analysis via Fluorescent microscopy (Transduction from 19.08.2010)
- Results:
Midi-Prep of pSB1C3_lITR_CMV_betaglobin_mVenus_hGH_rITR clone1
Investigators: Chris W.
Comment: Midi-Preps of P 242, B200
The Midi-Preps were performed according to the standard protocol yielding the following concentrations:
| plasmid-no. | P269 | p270 |
| concentration (ng/µl) | 1035,10 | 1376,95 |
Re-Trafo of pSB1C3_SDM_SspI_Bla14FM
Investigator: Stefan
Comment: Mini-Prep of pSB1C3_SDM_SspI_Bla14FM showed extremely low plasmid concentrations. It was proposed that this could correspond to using not the standard lab cells but rather XL-10 Gold cells for trafo. Therefore a re-trafo was performed using BL21 cells. 2 µl of each clone was added to BL21 cells and trafo was performed. Clones can be picked tonight (Volker)!
97. labday 22.08.2010
Test-digestion of pCerulean (P273,4,5), pSB1C3_VP1up (P279, 280), pSB1C3_NLS (P281,282), pAAV_RC_final (P283)
Investigator: Patrick
Digestions:
- 0,5 µl EcoRI, 0,5 µl Enzyme PstI, 1 µl BSA (10x), 1 µl Buffer 4 (10x), 1 µl pCerulean (P73), 6 µl H2O; total volume = 10 µl
- 0,5 µl EcoRI, 0,5 µl Enzyme PstI, 1 µl BSA (10x), 1 µl Buffer 4 (10x), 1 µl pCerulean (P74), 6 µl H2O; total volume = 10 µl
- 0,5 µl EcoRI, 0,5 µl Enzyme PstI, 1 µl BSA (10x), 1 µl Buffer 4 (10x), 1 µl pCerulean (P75), 6 µl H2O; total volume = 10 µl
- 1 µl NotI, 1 µl BSA (10x), 1 µl Buffer 4 (10x), 1 µl pSB1C3_VP1up (P279), 6 µl H2O; total volume = 10 µl
- 1 µl NotI, 1 µl BSA (10x), 1 µl Buffer 4 (10x), 1 µl pSB1C3_VP1up (P280), 6 µl H2O; total volume = 10 µl
- 0,5 µl EcoRI, 0,5 µl Enzyme PstI, 1 µl BSA (10x), 1 µl Bufer 4 (10x), 1 µl pSB1C3_NLS (P281) , 6 µl H2O; total volume = 10 µl
- 0,5 µl EcoRI, 0,5 µl Enzyme PstI, 1 µl BSA (10x), 1 µl Bufer 4 (10x), 1 µl pSB1C3_NLS (P282) , 6 µl H2O; total volume = 10 µl
- 0,5 µl Acc65I, 0,5 µl BspMI, 1 µl BSA (10x), 1 µl Buffer 3 (10x) , 1 µl pAAV_RC_final , 6 µl H2O; total volume = 10 µl
Incubation time: 37°C, 1h
Expected size of the fragments:
- pCerulean clone 1 : 771 & 3915 bp
- pCerulean clone 2 : 771 & 3915 bp
- pCerulean clone 3 : 771 & 3915 bp
- pSB1C3_VP1up clone 1 : 468 & 2030 bp
- pSB1C3_VP1up clone 2 : 468 & 2030 bp
- pSB1C3_NLS clone 1 : 78 & 2031 bp
- pSB1C3_NLS clone 2 : 78 & 2031 bp
- pAAV_RC_final : 615 & 6720 bp
pCerulean clone 1 (P273), pSB1C3_VP1up clone 2 (P280), pSB1C3_NLS clone 2 (P282) and pAAV_RC_final (P283) were sent for sequencing although the results don't look very promising.
Labelling:
- pCerulean clone 1 (P273) : iGEM2-CMV-F
- pSB1C3_VP1up clone 2 (P280) : iGEM1-VF2
- pSB1C3_NLS clone 2 (P282) : iGEM276-O51_VR2 and iGEM3-O51_VR2 (was ordered twice due to an ordering mistake)
- pAAV_RC_final (P283) : iGEM4-SK
- pAAV_RC_final (P283) : iGEM4-VP1 pKEX rev
- pAAV_RC_final (P283) : iGEM4-Cap2800for
- pAAV_RC_final (P283) : iGEM4-Cap2800rev
- pAAV_RC_final (P283) : iGEM4-Cap3500for
Mini-Prep of pCerulean
Investigator: Stefan
-
Glycerol stocks were prepared:
- B225 = pCerulean clone1
- B226 = pCerulean clone2
- B227 = pCerulean clone3
-
Mini-Prep following the standard protocol
- P273 = pCerulean clone1 c= 419,52 ng/µl
- P274 = pCerulean clone2 c= 383,37 ng/µl
- P275 = pCerulean clone3 c= 382,72 ng/µl
Test digestion will be performed tomorrow.
98. labday 23.08.2010
Primer design: Sequencing primer for BioBrick Assembly of vectorplasmid and phTERT qPCR primer
Investigator: Bea
Comment: Two goals will be accomplished with this primers.
- The first primer pair (left picture) serve as sequencing primers for the pSB1C3 vector plasmid. Problem was that the designed VR-2/VF-2 primers cannot be used for sequencing because the ITRs flank several assembled parts, mostly CMV, beta globin, GOI and hGH. These primers were designed to bind in the flanked region, directly after/before left and right ITR repsectivel and can therfore be used to sequence most of the flanked sequence inbetween the ITRs.
- The second primer pair (right picture) will be used for performing the qPCR with th econtruct containing the phTERT promoter. The qPCR primers used for the other qRT-PCR cannot be used because these two primers anneal in the CMV region. The new designed primers bind in the phTERT region and can therefore bee used for determinining the infectious titers.
Mini-Prep of pSB1C3_CFP_middlelinker and pSB1C3_ZEGFR:1907 / testdigestion of pSB1C3_CFP_middlelinker and pSB1C3_lITR_pHTERT
Investigator: Jessica
Mini-Prep following the standard protocoll
- P276 PSB1C3_CFP_middlelinker clone1 c=207,7 ng/µl
- P277 PSB1C3_CFP_middlelinker clone2 c=203,0 ng/µl
- P278 PSB1C3_CFP_middlelinker clone3 c=227,9 ng/µl
- P279 pSB1C3_VP1up clone1 c=183,0 ng/µl
- P280 pSB1C3_VP1up clone2 c=178,1 ng/µl
- P281 pSB1C3_NLS clone1 c=155,7 ng/µl
- P282 pSB1C3_NLS clone2 c=142,6 ng/µl
- P283 pAAV_RC_final c=147,7 ng/µl
- P284 pSB1C3_ZEGFR:1907 clone 1 c=146,5 ng/µl
- P285 pSB1C3_ZEGFR:1907 clone 1 c=161,9 ng/µl
-
Test digestion:
- Perfomed with 10 µL of total volume
| Mastermix/µL | P256/µL | P257/µL | P276/µL | P277/µL | P278/µL | |
| DNA | - | 5 | 5,2 | 4,3 | 4,4 | 4,0 |
| BSA (10x) | 6 | 1 | 1 | 1 | 1 | 1 |
| Buffer 4 (10x) | 6 | |||||
| Enzyme XbaI | 3 | |||||
| Enzyme PstI | 3 | |||||
| H2O | - | 2 | 1,8 | 2,7 | 2,6 | 3,0 |
| Total volume | 18 | 10 | 10 | 10 | 10 | 10 |
- incubation time: 50'
- incubation temparatur: 37°C
- 5µl new ladder mix
Sequencing:
- Plasmid used: P276: pSB1C3_CFP_middlelinker
- Primer used: VR-2
- Tube name: JG 276
Continuation of BioBrick assembly for pSB1C3_Affibody_Linker
Investigator: Anna
Comment: Transformation was done of pSB1C3_Affibody with Short-Linker, Middle-Linker, Long-Linker, SEG-Linker and GSAT-Linker. Unfortunately, the digestion of pSB1C3_YFP_His-Tag didn't work (see 20.08).
Transformation was done following the standard protocol using XL1 blue cells.
To do: Repetition of BioBrick assembly for pSB1C3_YFP_His-Tag.
Cloning of pSB1C3_ß-globin_YFP and pSB1C3_hGH_rITR
Investigator: Anna
Comment: pSB1C3_ß-globin_mVenus (p178) was digested with SpeI and PstI and pSB1C3_hGH_rITR (p186) was digested with XbaI and PstI .
Vector:
- pSB1C3_ß-globin_YFP (p178): c= 237,2 ng/µl
Insert:
- pSB1C3_hGH_rITR (p186): c= 105,2 ng/µl
| components | Vector (p178) | Insert (p186) |
| DNA | 4,2 | 14 |
| BSA (10x) | 1,5 | 2,5 |
| Buffer 4 (10x) | 1,5 | 2,5 |
| Enzyme EcoRI | 1 | 1,5 |
| Enzyme PstI | 1 | 1,5 |
| H2O | 5,8 | 3 |
| Total volume | 15 | 25 |
- Gelextraction of pSB1C3_RFC25_CFP:
0,5 g Agarose,50 ml TAE (1%), 3 µl GELRED , at 120 Volt, running time:55
Marker: GeneRuler ladder mix (Fermentas)
- c(pSB1C3_ß-globin_YFP)= 16.9 ng/µl; size: 3293 bp
- c(hGH_rITR)= 7,5 ng/µl; size: 657 bp
- Ligation of PCR products and vector:
For the Ligation 1µl T4 buffer (2x) and 1µl T4 ligase were used. Incubation time: 30 min
| vector /µl | insert /µl | |
| pSB1C3_ß-globin_mVenus + hGH_rITR | 3,41 | 4,59 |
- Transformation:
The transformation was done following the standard protocol using XL1 blue cells.
To do: Picking clones of pSB1C3_ß-globin_mVenus_hGH_rITR.
Repetition: Quickchange site-directed mutagenesis of pSB1C3_SDM_SspI_Bla14FM
Investigator: Stefan
Comment: To delete a PvuII restriction site in the CAT marker, a SDM will be performed.
- PCR reaction:
| Ingredients | v / µl |
| 10x Pfu Ultra II buffer | 2,5 |
| dNTP | 0,5 |
| forward primer: O109 (pSB1C3 PvuII for) | 0,59 |
| reverse primer: O108 (pSB1C3 PvuII rev) | 0,58 |
| DNA Template 10ng (1:100 dilution) | 4 |
| DMSO (1%) | 0,5 |
| PfuUltra II fusion | 0,5 |
| H2O | 15,83 |
| Total volume | 25 |
- PCR program:
 "
"