Team:Tsinghua/Notebook/25 September 2010
From 2010.igem.org
(Difference between revisions)
(New page: == Module Ⅱ, group 4 == After getting and then diluting the primer, do Two-step PCR. PCR with Pyrobest DNA polymerase. PCR system(50ul): 10×Pyrobest buffer Ⅱ 5ul ...) |
(→Module Ⅱ, group 4) |
||
| Line 16: | Line 16: | ||
Total 50μl | Total 50μl | ||
| - | + | PCR program: | |
| - | + | Step Condition Time | |
| - | + | 1 95℃ 5min | |
| - | + | 2 95℃ 30s | |
| - | + | 3 58℃ 30s | |
| - | + | 4 72℃ 1min | |
| - | + | 5 return to step2 15cycles | |
| + | 6 72℃ 5min | ||
| + | 7 add amplification primers | ||
| + | 8 95℃ 5min | ||
| + | 9 95℃ 30s | ||
| + | 10 56℃ 30s | ||
| + | 11 72℃ 1min | ||
| + | 12 return to step2 30cycles | ||
| + | 13 72℃ 1min | ||
| + | 14 4℃ HOLD | ||
| - | Run a gel | + | |
| + | Run a gel. | ||
== result == | == result == | ||
Revision as of 03:54, 28 October 2010
Module Ⅱ, group 4
After getting and then diluting the primer, do Two-step PCR.
PCR with Pyrobest DNA polymerase.
PCR system(50ul):
10×Pyrobest buffer Ⅱ 5ul Pyrobest 0.3ul dNTPmix 1ul Primer1 1μl Primer2 1ul Template DNA 0.5ul Mgcl2 0.5ul ddH2O 40.5ul Total 50μl
PCR program:
Step Condition Time 1 95℃ 5min 2 95℃ 30s 3 58℃ 30s 4 72℃ 1min 5 return to step2 15cycles 6 72℃ 5min 7 add amplification primers 8 95℃ 5min 9 95℃ 30s 10 56℃ 30s 11 72℃ 1min 12 return to step2 30cycles 13 72℃ 1min 14 4℃ HOLD
Run a gel.
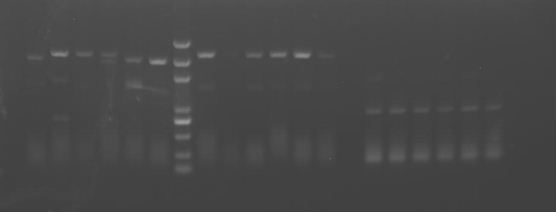
result
P+E+M② is positive. It seems that PKC 1-6 are positive. But it still remains unclear whether the PCR bands are so weak and why the digestion result seemed negative.
 "
"