Team:ETHZ Basel/Modeling/Light Switch
From 2010.igem.org
(→Modeling of the light switch) |
(→Modeling of the light switch) |
||
| Line 12: | Line 12: | ||
Inspired by the modular approach used for the [[Team:ETHZ_Basel/Biology/Cloning|Cloning Strategy]] in the wet-lab, we decided to decrease the combinatorial explosion by also applying a novel modular approach. Not only did this approach reduce the amount of models by a factor of four, it also allowed to widely separate the differential equations of the chemotaxis pathway from those of the light-induced localization system by simultaneously decreasing redundancies and thus decreasing the danger of slip of the pens. The underlying mathematical model for the light-induced relocation was completely developed by us (for a short discussion of the recently by Sorokina et al. published light-induced relocation system and why we did not use it, see [[Team:ETHZ_Basel/Modeling/Sorokina|here]]) as well as - to our knowledge - the approach to couple this model to models of the chemotaxis pathway. | Inspired by the modular approach used for the [[Team:ETHZ_Basel/Biology/Cloning|Cloning Strategy]] in the wet-lab, we decided to decrease the combinatorial explosion by also applying a novel modular approach. Not only did this approach reduce the amount of models by a factor of four, it also allowed to widely separate the differential equations of the chemotaxis pathway from those of the light-induced localization system by simultaneously decreasing redundancies and thus decreasing the danger of slip of the pens. The underlying mathematical model for the light-induced relocation was completely developed by us (for a short discussion of the recently by Sorokina et al. published light-induced relocation system and why we did not use it, see [[Team:ETHZ_Basel/Modeling/Sorokina|here]]) as well as - to our knowledge - the approach to couple this model to models of the chemotaxis pathway. | ||
| - | [[Image:ETHZ_Basel_Flagellum.png|thumb|center|550px|'''Flagellum of a gram negative bacterium. Free cytosolic CheYp binds to the flagellar motor complex and induces tumbling]] | + | [[Image:ETHZ_Basel_Flagellum.png|thumb|center|550px|'''Flagellum of a gram negative bacterium. Free cytosolic CheYp binds to the flagellar motor complex and induces tumbling.]] |
Revision as of 13:08, 27 October 2010
Modeling of the light switch
Background
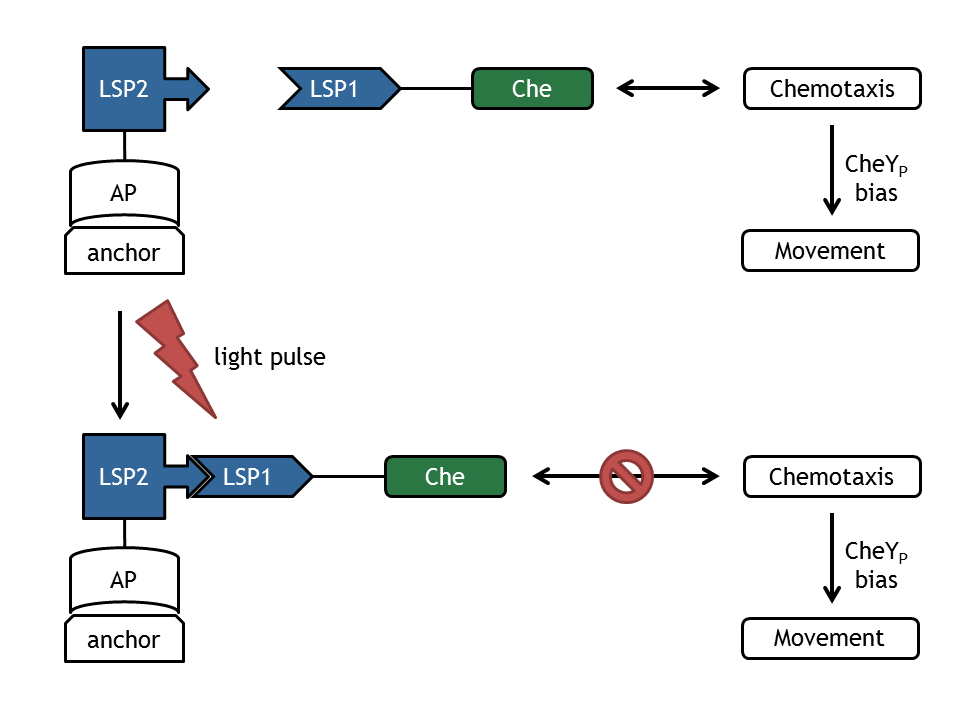
In our biological setup, the relocation of one of the chemotaxis pathway proteins (either CheR, CheB, CheY or CheZ) is achieved by fusing them to a light-sensitive protein LSP1 (either to PhyB or to PIF3), which dimerizes by red light induction with the corresponding LSP2 (PIF3 or PhyB), fused to a spatially dislocated anchor. Since we decided to implement two different models of the chemotaxis pathway (see section Chemotaxis Pathway), modeling all setups implemented in the wet-lab would have resulted in 16 different models:
|{CheR, CheB, CheY, CheZ} × {PhyB, PIF3} × {Model 1, Model 2}|=16.
Inspired by the modular approach used for the Cloning Strategy in the wet-lab, we decided to decrease the combinatorial explosion by also applying a novel modular approach. Not only did this approach reduce the amount of models by a factor of four, it also allowed to widely separate the differential equations of the chemotaxis pathway from those of the light-induced localization system by simultaneously decreasing redundancies and thus decreasing the danger of slip of the pens. The underlying mathematical model for the light-induced relocation was completely developed by us (for a short discussion of the recently by Sorokina et al. published light-induced relocation system and why we did not use it, see here) as well as - to our knowledge - the approach to couple this model to models of the chemotaxis pathway.
Facing the Combinatorial Explosion
The main problem in separating in silico the chemotaxis pathway from the light-induced relocation system is the non-existence of a hierarchical relationship between the two sub-models: The properties of the chemotaxis pathway are clearly influenced if the active concentration of one of its key species drops, such that giving the amount of localized proteins, obtained by the localization model, as an input to the chemotaxis model is a natural conclusion. However, also the concentration of the localized species of the localization model change depending on the reactions in the chemotaxis model, since several of the Che proteins are modeled as two or more molecular species. The CheY protein for example is modeled as two species, one representing the phosphorylated and one the non-phosphorylated molecular concentration. Thus, the two models cannot be represented as e.g. a hierarchical block structure, which is making a modular approach significant more complicated.
Our approach to modularize the model is based on an observation of the reaction directions which can take place in the overall model. For example, Figure 2 shows all eight species which have to be implemented to adequately describe the localization and phosphorylation states of CheY when fused to PhyB.
Download
The light switch model is included within the Matlab Toolbox and can be downloaded there.
 "
"