Team:TU Delft/Project/alkane-degradation/results
From 2010.igem.org
(→Growth on alcohols as sole carbon source) |
(→ALDH) |
||
| Line 175: | Line 175: | ||
===ALDH=== | ===ALDH=== | ||
| - | + | ==== Results === | |
=====Growth on alcohols as sole carbon source===== | =====Growth on alcohols as sole carbon source===== | ||
| - | + | As we described before, we tried to grow our recombinant strains ''Escherichia coli'' 029A ([http://partsregistry.org/Part:BBa_K398029 BBa_K398029] on the plasmid pSB1A2) and ''Escherichia coli'' 030A ([http://partsregistry.org/Part:BBa_K398030 BBa_K398030] on the plasmid pSB1A2) on Octanal and Dodecanal. No growth was observed after 48 hours. We abandoned this experiment after this result. | |
=====Resting cell assays===== | =====Resting cell assays===== | ||
Revision as of 15:05, 26 October 2010

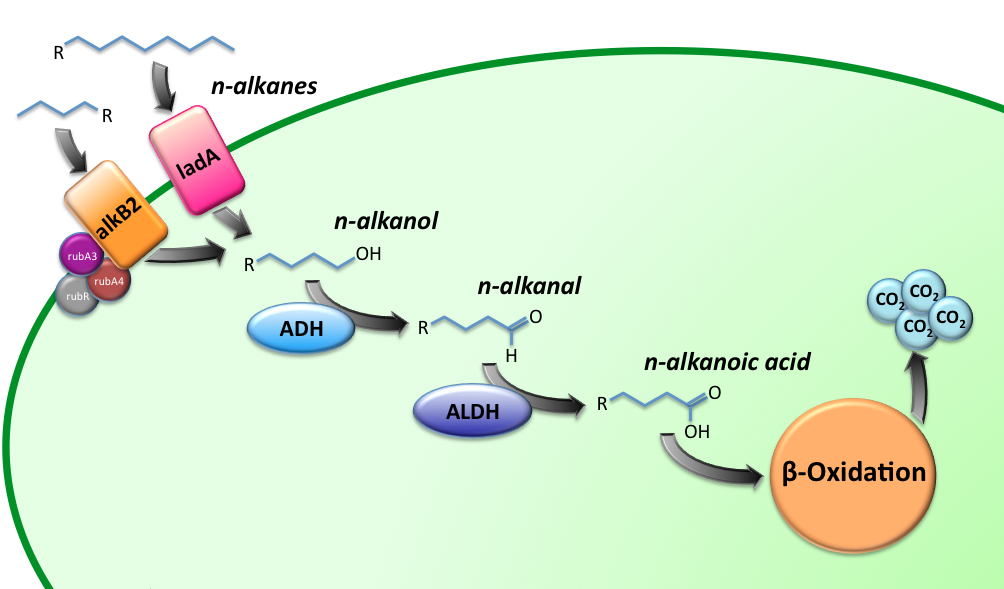
Alkane Degradation Results & Conclusions
Characterization of the alkane hydroxylase system
Growth analysis
We attempted to culture our recombinant AH-carrying E.coli K12 strains ([http://partsregistry.org/Part:BBa_K398014 BBa_K398014]) on 1% v/v octanol or 1% v/v dodecane. Negligible growth was observed.
Resting-cell assays
The resting cell assays were performed on our recombinant AH-carrying E.coli K12 cells ([http://partsregistry.org/Part:BBa_K398014 BBa_K398014]). 100 micromoles of octane was added to 6 mL of growth-stalled cells (1.5 mg cell dry weight total) and incubated at 37 degrees with shaking o/n. The organic phase was extracted using EtOAc and analysed by gas chromatography.
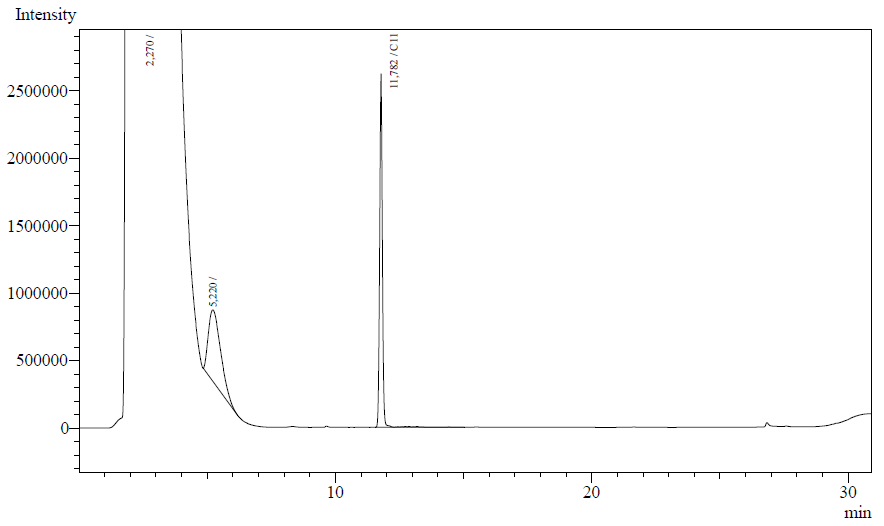
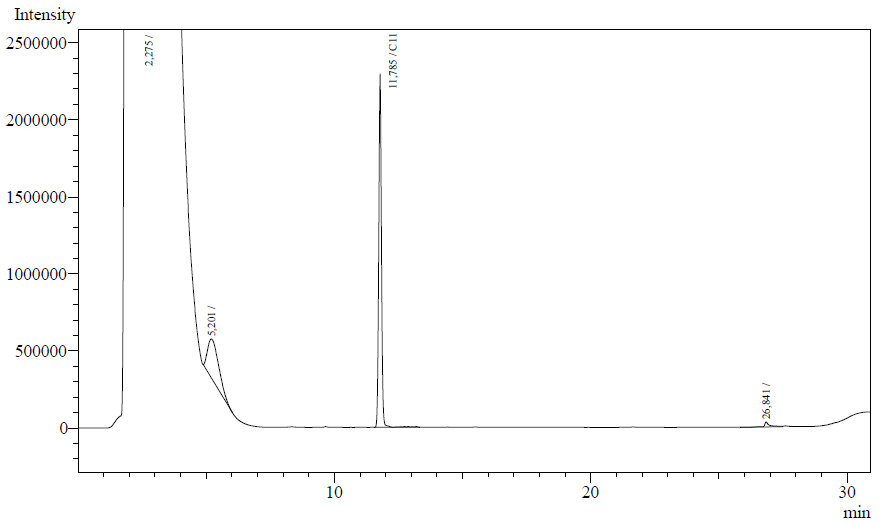
The following are examples of typical chromatographs obtained:
The peaks shown in the chromatographs belong to the following compounds:
| Retention time [min] | Compound |
| ~2.2 | Ethyl acetate (solvent) |
| ~5.2 | Octane (substrate) |
| ~11.8 | Undecane (internal standard, 0.1% v/v of solvent) |
The surface areas of the peaks correspond to the amount of molecules present in the sample. If we assume that a negligible amount hydrocarbon will remain present in the aqueous phase, we can state that 2.5 uL (
LadA
Alcohol DeHydrogenase (ADH) system
= Results
Growth on alcohols as sole carbon source
- As we described before, we tried to grow our strain Escherichia coli 018A ([http://partsregistry.org/Part:BBa_K398018 BBa_K398018] on the plasmid pSB1A2) on the alkanols Octanol-1 and Dodecanol-1 (1% V/V on M9 medium without any other carbon source). No growth was observed after 48 hours. We abandoned this experiment after this result.
Resting cell assays
- We grew Escherichia coli 018A and Escherichia coli negative control (Biobrick BBa_J13002 on plasmid pSB1A2 ) in 50 mL of M9 medium with glucose and CAS aminoacids.
- The cells were harevested when the O.D. at 600nm was around 0.3; they were spun down at 4000 rpm, for 10 min at 4ºC. And the resting cell assays were prepared according to the standard protocol
| Strain | O.D. |
| J13002 | 0.327 |
| 018A | 0.338 |
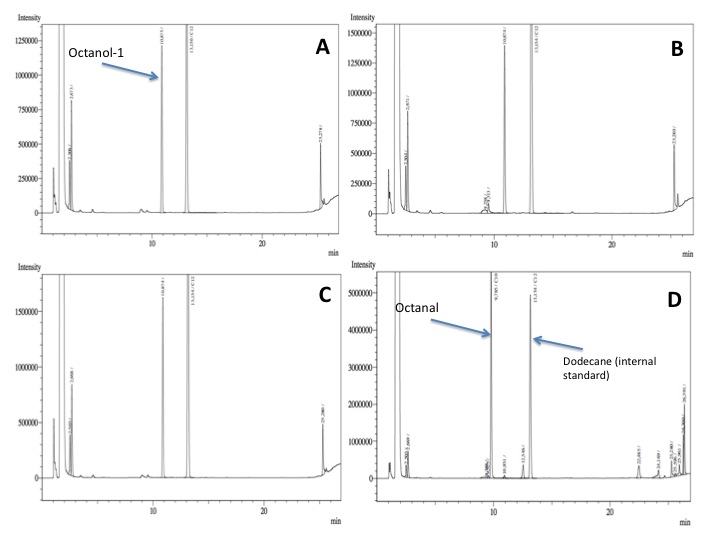
- After an overnight incubation at 37ºC, the organic phase was extracted using 3 mL of ethyl acetate (dodecane was used as internal standard). We tried to determine production of alkanals or alkanoic acids by Gas Chromatography measurements. The chromatograms are shown below:
- According to our results there was no degradation of octanol-1 ='( , after this experiment the protocol was abandoned. We think that the cells lack of transporters for long-chain alcohols, we preferred to check the cell extract in order to know if there was any biological activity in the cytoplasma.
NADH production in cell extracts
- EXPERIMENT 1
Cells were cultured in 50mL of LB medium and harvested when the O.D. 600nm of the culture was around 0.6. Two different cultures of the recominant strain and the negative control were prepared.
| Strain | O.D. 600nm |
| J13002 (1) | 0.654 |
| J13002 (2) | 0.621 |
| 018A (1) | 0.615 |
| 018A (2) | 0.580 |
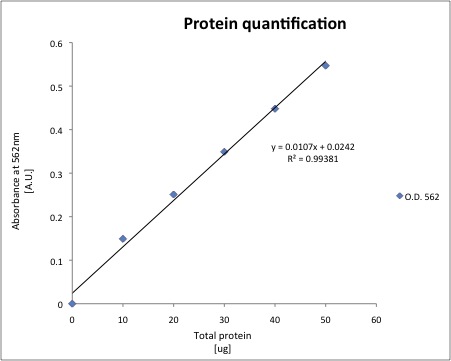
Cytoplasmic proteins were extracted using our standard protocol. And a standard curve for protein quantification was prepared.
The total protein of each sample was quantified using 20uL of cell extract, the results are shown below.
| Sample | O.D. 562 nm | Total ug of protein | ug/uL |
| J13002(1A) | 0.135 | 10.393 | 1.0393 |
| J13002 (1B) | 0.135 | 10.393 | 1.0393 |
| J13002 (2A) | 0.128 | 9.7363 | 0.9736 |
| J13002 (2B) | 0.129 | 9.8302 | 0.9830 |
| 018A (1A) | 0.097 | 6.8275 | 0.6827 |
| 018A (1B) | 0.091 | 6.2645 | 0.6264 |
| 018A (2A) | 0.092 | 6.3583 | 0.6358 |
| 018A (2B) | 0.092 | 6.3583 | 0.6358 |
The alcohol dehydrogenase activity was measured using the standard protocol and Dodecanol-1 as substrate. You can download our raw data by clicking on the link: File:TUDelft ADH raw.xls
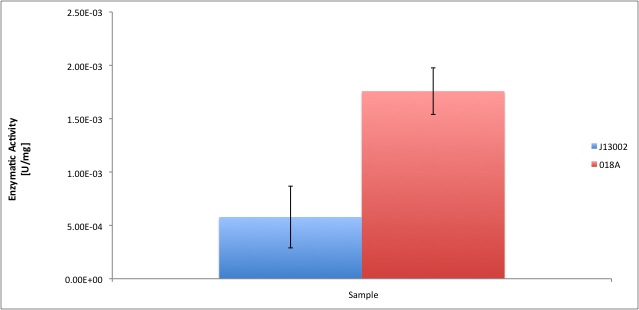
After the data treatment, the results that we obtained were the following:
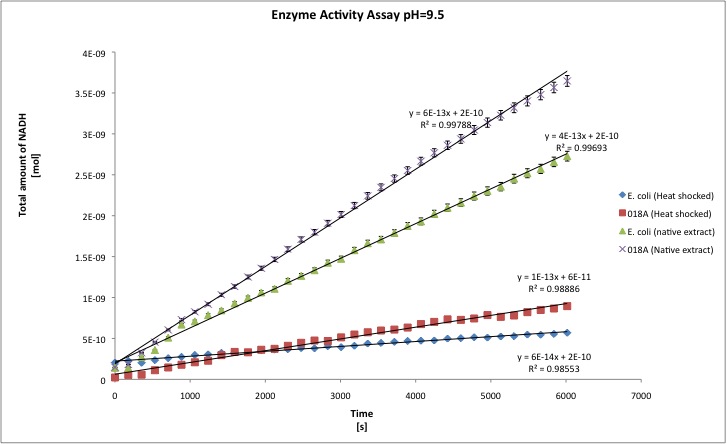
 All the enzyme activities were normalized to the TOTAL amount of protein in the cell extract. The results are shown in katal per mg of protein in the cell extract and Enzymatic Units per mg of protein in the cell extract.
All the enzyme activities were normalized to the TOTAL amount of protein in the cell extract. The results are shown in katal per mg of protein in the cell extract and Enzymatic Units per mg of protein in the cell extract.
| pH=9.5 | pH=8 | |||||||
| Heat | Native | Heat | Native | |||||
| J13002 | 018A | J13002 | 018A | J13002 | 018A | J13002 | 018A | |
| kat/mg | 6,156E-12 | 1,941E-11 | 1,053E-11 | 2,806E-11 | 3,211E-12 | 4,496E-11 | 1,302E-11 | 2,638E-11 |
| U/mg | 3,694E-04 | 1,091E-03 | 6,317E-04 | 1,694E-03 | 1,927E-04 | 2,139E-03 | 7,811E-04 | 1,608E-03 |
| stdev | 1,157E-12 | 4,579E-12 | 1,343E-12 | 9,961E-13 | 6,395E-12 | 2,331E-11 | 6,511E-13 | 9,961E-13 |
| Improvement | - | 215,34% | - | 166,51% | - | 1299,87% | - | 166,51% |
| ttest | - | 0,9963 | - | 1,0000 | - | 0,9988 | - | 1,000 |
You can download our results file here: File:TUDelft ADH results.xls Maybe you are not familiar with the term "katal", officially the katal is the STANDARD UNIT FOR CATALYTIC ACTIVITY in the International System of Units; the katal is defined as the amount of enzyme that converts 1 mol of substrate each second. [http://www.clinchem.org/cgi/content/full/48/3/586 Click here] to know more about it, and because people are not that familiar with the term, we report the activities in U/mg. One enzyme activity unit is defined as the amount of protein that converts 1 umol of substrate each minute. T-tests were performed in order to show that there is a difference between E. coli and our recombinant strain; a result for t-test of 1 means that both samples are statistically different; whereas a t-test of nearly 0 means that samples belong to the same group (no difference between both). Our level of significance is 0.025, in two tailed test. That means that if our t-test result is 0.95 we conclude that both samples are statistically different, whereas 0.949 or lower means that both samples are not statistically different.
- EXPERIMENT 2
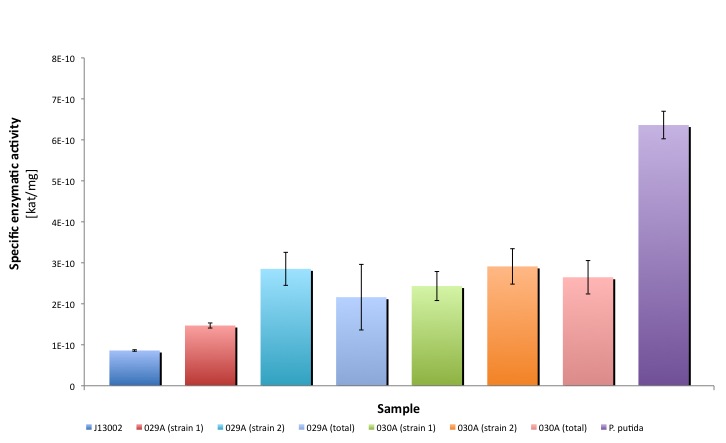
We ran a second experiment at pH 9.5, with two other strains that seemed to be positive in the colony PCR test. We followed the same procedure as in the experiment 1, the results obtained were the following:
| kat/mg | U/mg | STDEV | Improvement | T-test | Relative to putida | ||
|---|---|---|---|---|---|---|---|
| E. Coli | 8.75E-12 | 5.25E-04 | 8.18987E-12 | - | - | 0.90% | |
| 018A-2 | 9.30E-12 | 5.58E-04 | 1.84877E-12 | 6.26% | 0.4708 | 0.96% | |
| 018A-3 | 3.05E-11 | 1.83E-03 | 6.27802E-12 | 249.14% | 0.9808 | 3.15% | |
| 018A-4 | 8.26E-12 | 4.96E-04 | 1.8256E-12 | -5.59% | 0.4053 | 0.85% | |
| P. Putida | 9.69E-10 | 5.82E-02 | 2.09109E-10 | - | 0.9883 | 100.00% |
According to these results, the other two strains (018A-2 and 018A-4) do not functionally express the part BBa_K398018. However, these results confirm our findings from the first experiment. You can download our result in this link: File:TUDelft ADH results2.xls You can also find a summary in this file:File:TUDelft ADH summary.xls
Conclusions
According to our results, a normal E. coli cell extract has a dodecanol-1 dehydrogenase activity of 9.64e-12 kat/mg (0.58 mU/mg); whereas our recombinant strain 018A has an activity of 2.93e-11 kat/mg (1.76 mU/mg). According to the statistical analysis, the enzymatic activities of both strains are statistically different at confidence level of 0.95 which means that the part BBa_K398018 increases 2 times the alcohol dehydrogenase activity in the cell extract. We can conclude from our data that the part has a biological activity in vitro.
We still don't know about the in vivo activity of this part, maybe there is a lack of long-chain alcohol transporter proteins or it is necessary to over-express this part in order to see/measure any biological activity.
ALDH
= Results
Growth on alcohols as sole carbon source
As we described before, we tried to grow our recombinant strains Escherichia coli 029A ([http://partsregistry.org/Part:BBa_K398029 BBa_K398029] on the plasmid pSB1A2) and Escherichia coli 030A ([http://partsregistry.org/Part:BBa_K398030 BBa_K398030] on the plasmid pSB1A2) on Octanal and Dodecanal. No growth was observed after 48 hours. We abandoned this experiment after this result.
Resting cell assays
- We grew Escherichia coli 018A and Escherichia coli negative control (Biobrick BBa_J13002 on plasmid pSB1A2 ) in 50 mL of M9 medium with glucose and CAS aminoacids.
- The cells were harevested when the O.D. at 600nm was around 0.3; they were spun down at 4000 rpm, for 10 min at 4ºC. And the resting cell assays were prepared according to the standard protocol
| Strain | O.D. |
| J13002 | 0.327 |
| 018A | 0.338 |
- After an overnight incubation at 37ºC, the organic phase was extracted using 3 mL of ethyl acetate (dodecane was used as internal standard). We tried to determine production of alkanals or alkanoic acids by Gas Chromatography measurements. The chromatograms are shown below:
- According to our results there was no degradation of octanol-1 ='( , after this experiment the protocol was abandoned. We think that the cells lack of transporters for long-chain alcohols, we preferred to check the cell extract in order to know if there was any biological activity in the cytoplasma.
NADH production in cell extracts
We called this assay: THE LAST RESOURCE =s
Basically, we grew cells until an O.D. (600nm) between 0.5-1.0. We needed chubby healthy happy exponential-growth cells because most of our constructions are meant for protein production in this growth phase.
Once, we got a lot of cells... we disrupted them by sonication and we analyze the NADH production by their guts using a simple spectrophotometrical analysis. If our parts are working, we should see a higher NADH production when the substrate (dodecanol-1 or dodecanal) to the buffer with microbial guts and NAD buffer. We can use this method because NAD is the natural co-factor of our proteins. Fortunately for us, NAD reduction could be easily quantified by absorbance measurements at 340 nm.
We called this protocol "the last resource" because the others didn't work. Which means that alcohols and aldehydes do not cross the cell membrane, thus in order to grow cell on these compounds as sole carbon sources... WE NEEDED TO ADD TRANSPORTERS =(
BBa_K398030 and BBa_K398029
Both Biobricks are the protein generators of Bt-ALDH. We compared the activities of bot parts, parental strain with an empty plasmid (J13002) and a natural oil-degrader (Pseudomonas putida).
View [http://partsregistry.org/wiki/index.php?title=Part:BBa_K398029 BBa_K398029 in the parts registry]
Specified Components <partinfo>K398029 SpecifiedComponents</partinfo>
Designer: <partinfo>K398029 Designer</partinfo>
Status: <partinfo>K398029 Status</partinfo>
View [http://partsregistry.org/wiki/index.php?title=Part:BBa_K398030 BBa_K398030 in the parts registry]
Specified Components <partinfo>K398206 SpecifiedComponents</partinfo>
Designer: <partinfo>K398030 Designer</partinfo>
Status: <partinfo>K398030 Status</partinfo>

 "
"