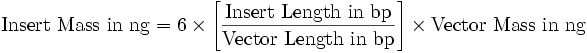Team:UNIPV-Pavia/Material Methods/Protocols
From 2010.igem.org
(New page: <!-- originale della home disponibile a https://2010.igem.org/Team:UNIPV-Pavia/homebackup --> <table width="100%" border="0"> <tr> <td colspan="2"> {{UNIPV-Pavia/header}} </td> ...) |
m |
||
| Line 69: | Line 69: | ||
==M9 supplemented with glycerol (M9gly)== | ==M9 supplemented with glycerol (M9gly)== | ||
| - | |||
For 1L of medium, add: | For 1L of medium, add: | ||
* 716 ml of autoclaved (and cooled to Tamb) ddH2O | * 716 ml of autoclaved (and cooled to Tamb) ddH2O | ||
| Line 93: | Line 92: | ||
<div align="right"><small>[[#indice|^top]]</small></div> | <div align="right"><small>[[#indice|^top]]</small></div> | ||
| - | <br> | + | <br/><br/> |
==Antibiotics== | ==Antibiotics== | ||
| Line 107: | Line 106: | ||
* Chloramphenicol 12.5 ug/ml | * Chloramphenicol 12.5 ug/ml | ||
<div align="right"><small>[[#indice|^top]]</small></div> | <div align="right"><small>[[#indice|^top]]</small></div> | ||
| - | |||
<br><br> | <br><br> | ||
| Line 129: | Line 127: | ||
* take 650ul of supernatant and resuspend the pellet in the remaining LB (~150ul) | * take 650ul of supernatant and resuspend the pellet in the remaining LB (~150ul) | ||
* plate the entire culture and incubate the plate at 37°C overnight | * plate the entire culture and incubate the plate at 37°C overnight | ||
| - | |||
Variants: | Variants: | ||
| Line 136: | Line 133: | ||
* if you use commercial Invitrogen TOP10 the protocol changes and it is reported below. | * if you use commercial Invitrogen TOP10 the protocol changes and it is reported below. | ||
<div align="right"><small>[[#indice|^top]]</small></div> | <div align="right"><small>[[#indice|^top]]</small></div> | ||
| - | + | <br/><br/> | |
| - | <br> | + | |
==Transforming commercial competent cells== | ==Transforming commercial competent cells== | ||
| Line 157: | Line 153: | ||
* the remaining 150ul can be stored at +4°C | * the remaining 150ul can be stored at +4°C | ||
<div align="right"><small>[[#indice|^top]]</small></div> | <div align="right"><small>[[#indice|^top]]</small></div> | ||
| - | |||
<br><br> | <br><br> | ||
| Line 191: | Line 186: | ||
* DB3.1 (5*10^4 with 100ul of cells); | * DB3.1 (5*10^4 with 100ul of cells); | ||
<div align="right"><small>[[#indice|^top]]</small></div> | <div align="right"><small>[[#indice|^top]]</small></div> | ||
| - | |||
<br><br> | <br><br> | ||
=E. coli strains (all in -80°C freezer)= | =E. coli strains (all in -80°C freezer)= | ||
| - | |||
==TOP10== | ==TOP10== | ||
| Line 290: | Line 283: | ||
=Ethanol precipitation with sodium acetate= | =Ethanol precipitation with sodium acetate= | ||
| - | |||
* Add 1/10 DNA solution volume of sodium acetate 3 M, pH 5.2 | * Add 1/10 DNA solution volume of sodium acetate 3 M, pH 5.2 | ||
* Add 2.5 DNA solution volume of absolute ethanol | * Add 2.5 DNA solution volume of absolute ethanol | ||
| Line 302: | Line 294: | ||
* Elute with 5-10 µl of ddH2O | * Elute with 5-10 µl of ddH2O | ||
<div align="right"><small>[[#indice|^top]]</small></div> | <div align="right"><small>[[#indice|^top]]</small></div> | ||
| - | |||
<br><br> | <br><br> | ||
=Ligation= | =Ligation= | ||
| - | |||
After the purification of two digested DNA fragments: | After the purification of two digested DNA fragments: | ||
* add a volume containing 20-50 ng of vector | * add a volume containing 20-50 ng of vector | ||
| Line 331: | Line 321: | ||
=DNA resuspension from iGEM plates= | =DNA resuspension from iGEM plates= | ||
| - | |||
*Find the right position of the DNA of interest in iGEM plates | *Find the right position of the DNA of interest in iGEM plates | ||
*Resuspend with 15 ul ddH2O and transfer it in 0,5 ml sterile Eppendorf tubes | *Resuspend with 15 ul ddH2O and transfer it in 0,5 ml sterile Eppendorf tubes | ||
| Line 381: | Line 370: | ||
</td> | </td> | ||
</tr></table> | </tr></table> | ||
| + | <div align="right"><small>[[#indice|^top]]</small></div> | ||
| + | <br/><br/> | ||
| + | |||
| + | =Glycerol stocks= | ||
| + | *Mix 750 ul from the 5 ml sample of the incubated bacteria with 250 ul of 80% glycerol | ||
| + | *Leave at -20°C for one day | ||
| + | *Move to -80°C the day after | ||
| + | <div align="right"><small>[[#indice|^top]]</small></div> | ||
| + | <br/><br/> | ||
| + | =X-Gal staining protocol for beta galactosidase (blue/white screening)= | ||
| + | *The principle is that X-Gal (5-bromo-4-chloro-3-indolyl-b-D-galactopyranoside) turns blue when reacts with beta-galactosidase. | ||
| + | *Mix 20 ul X-Gal 20 mg/ml and 80 ul SOC and spread on LB+amp plates (X-Gal and DMF are toxic, use face-mask for your safety!!! X-Gal is light-sensitive, remember to keep it in the dark, when possible) | ||
| + | *Incubate with 200 ul bacteria | ||
| + | *Result: blue colonies express LacZ, while white colonies don't | ||
<div align="right"><small>[[#indice|^top]]</small></div> | <div align="right"><small>[[#indice|^top]]</small></div> | ||
<br/><br/> | <br/><br/> | ||
Revision as of 14:33, 24 June 2010
|
|
|||||||||||||
|
Media & AntibioticsLB
to a sterile pyrex bottle
LB Agar
to a sterile 1L flask
SOB
to a sterile pyrex bottle
SOC
M9 supplemented with glycerol (M9gly)For 1L of medium, add:
NOTE:
can be stored at +4°C
can be stored at room temperature or +4°C
AntibioticsStocks at -20°C freezer:
These stocks are 1000x for high copy number plasmids. For low copy number plasmids, you should use these final concentrations in media:
E. coli transformationTransforming home-made competent cells
Variants:
Transforming commercial competent cells(according to manufacturer’s protocol)
E. coli competent cells preparationH. Inoue et al. (1990), High efficiency transformation of Escherichia coli with plasmids, Gene 96 23-28.
ALWAYS TEST THE EFFICIENCY IN [CFU/ug] UNITS This protocol has shown to work with:
E. coli strains (all in -80°C freezer)TOP10F- mcrA Δ(mrr-hsdRMS-mcrBC) φ80lacZΔM15 ΔlacX74 nupG recA1 araD139 Δ(ara-leu)7697 galE15 galK16 rpsL(StrR) endA1 λ-
NOTE: they have
DH5alphaF- endA1 glnV44 thi-1 recA1 relA1 gyrA96 deoR nupG Φ80dlacZΔM15 Δ(lacZYA-argF)U169, hsdR17(rK- mK+), λ–
BW20767F-, RP4-2(Km::Tn7,Tc::Mu-1), leu-163::IS10, ΔuidA3::pir+, recA1, endA1, thi-1, hsdR17, creC510
NOTE: they have
XL1-BlueendA1 gyrA96(nalR) thi-1 recA1 relA1 lac glnV44 F'[ ::Tn10 proAB+ lacIq Δ(lacZ)M15] hsdR17(rK- mK+)
NOTE: they have lacIQ DB3.1F- gyrA462 endA1 glnV44 Δ(sr1-recA) mcrB mrr hsdS20(rB-, mB-) ara14 galK2 lacY1 proA2 rpsL20(Smr) xyl5 Δleu mtl1
NOTE: they have a working lacZ, but a deleted lacY, they become slightly blue on IPTG/X-Gal plates STBL3F- glnV44 recA13 mcrB mrr hsdS20(rB-, mB-) ara-14 galK2 lacY1 proA2 rpsL20 xyl-5 leu mtl-1
NOTE: they cannot be used for blue/white screening CW2553 + pJat8Genotype: Khlebnikov A et al. (2000), Regulatable Arabinose-Inducible Gene Expression System with Consistent Control in All Cells of a Culture, Journal of Bacteriology, Vol. 182, No. 24, p.7029-7034.
NOTE:
Long term bacterial glycerol stocks
Plasmid digestion for BioBrick Standard Assembly
NOTE: if you are performing a digestion for screening, 1 hour of incubation is sufficient.
Ethanol precipitation with sodium acetate
LigationAfter the purification of two digested DNA fragments:
(“6” can be lowered to “2”)
NOTE: When the purified DNA of the insert also contains its native vector, you can perform the ligation anyway, but its antibiotic resistance must be different from the acceptor vector’s resistance in order to select correct transformants on agar plates. When doing this, you should modify the ligation protocol:
DNA resuspension from iGEM plates
Now you can store them at -20°C or transform in your favorite strain.
PCR
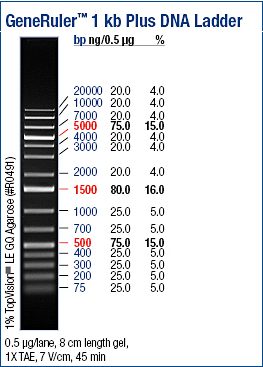
Electrophoresis
Glycerol stocks
X-Gal staining protocol for beta galactosidase (blue/white screening)
| ||||||||||||
 "
"