Team:Penn State/Project
From 2010.igem.org
(→Genetic Circuit) |
(→Overall project) |
||
| (9 intermediate revisions not shown) | |||
| Line 48: | Line 48: | ||
| - | This system could also be implemented in other chassis, such as yeast (wine) or Lactobacillus (yogurt), so the food itself could be the sensor. Genetically Modified Organisms (GMOs) in food would obviously face severe regulatory constraints. To better understand public perception, we created and administered a survey to over 250 randomized students in cooperation with other iGEM teams, especially RMIT. For details, see our [[Human Practices|Human Practices]] page. | + | This system could also be implemented in other chassis, such as yeast (wine) or Lactobacillus (yogurt), so the food itself could be the sensor. Genetically Modified Organisms (GMOs) in food would obviously face severe regulatory constraints. To better understand public perception, we created and administered a survey to over 250 randomized students in cooperation with other iGEM teams, especially RMIT. For details, see our [[Team:Penn State/Human Practices|Human Practices]] page. |
| Line 212: | Line 212: | ||
| - | The Tet inverter behaves somewhat as a constitutive promoter when no promoter is in front of it. Following the Tet inverter is a cI coding sequence (which codes for the lambda repressor). The lambda repressor binds to the lux promoters in the front of the circuit, which are also lambda repressible. This, coupled with the fact that the conditions are anaerobic, provides a very strong " | + | The Tet inverter behaves somewhat as a constitutive promoter when no promoter is in front of it. Following the Tet inverter is a cI coding sequence (which codes for the lambda repressor). The lambda repressor binds to the lux promoters in the front of the circuit, which are also lambda repressible. This, coupled with the fact that the conditions are anaerobic, provides a very strong "OFF" switch that does not allow the expression of the Anaerobic Fluorescent Protein (AFP) to occur. |
===FNR breaks apart in oxygen=== | ===FNR breaks apart in oxygen=== | ||
| Line 248: | Line 248: | ||
=== Oxygen Sensor === | === Oxygen Sensor === | ||
| - | The design of the oxygen-sensing promoter was based on the dcuC gene of E. coli. The two binding sites for FNR surrounded the RNAP binding site and were systematically shifted upstream and downstream from this site in multiple constructs, causing overlap of the RNAP binding site in some cases. | + | The design of the oxygen-sensing promoter was based on the dcuC gene of E. coli. The two binding sites for FNR surrounded the RNAP binding site and were systematically shifted upstream and downstream from this site in multiple constructs, causing overlap of the RNAP binding site in some cases. Using this strategy, five constructs were designed with the constitutive promoter J23113 as the basis for the sequence following the downstream FNR binding site. The construct that worked best as an oxygen-sensing promoter (J6, [[http://partsregistry.org/Part:BBa_K376003 BBa_J376003]]) was fully characterized and is shown below. |
| Line 254: | Line 254: | ||
The oxygen promoters were characterized using a Tecan infinite M1000 microplate reader. Each sample was grown overnight on plates and transferred to a 96 deep-well plate for overnight growth. After 16 hours the samples were tested in the Tecan, where measurements would be taken for around 12 hours. Sometimes, measurement of growth of three successive plates (serially diluted from the preceding one) was needed before the appropriate steady state could be recorded. | The oxygen promoters were characterized using a Tecan infinite M1000 microplate reader. Each sample was grown overnight on plates and transferred to a 96 deep-well plate for overnight growth. After 16 hours the samples were tested in the Tecan, where measurements would be taken for around 12 hours. Sometimes, measurement of growth of three successive plates (serially diluted from the preceding one) was needed before the appropriate steady state could be recorded. | ||
| - | |||
| - | |||
== Results == | == Results == | ||
| Line 267: | Line 265: | ||
<center>[[Image:RawODoxygen.png|800px]]</center> | <center>[[Image:RawODoxygen.png|800px]]</center> | ||
| - | Shown below is the Fluorescence over OD of each sample. As the samples reach steady state, the graph should become a horizontal line. Notice how the graph comes closer and closer to reaching this steady state with each successive plate. Also, it is important to | + | Shown below is the Fluorescence over OD of each sample. As the samples reach steady state, the graph should become a horizontal line. Notice how the graph comes closer and closer to reaching this steady state with each successive plate. Also, it is important to remember that evaporation comes into play here when comparing anaerobic and anaerobic samples with each other. Since the anaerobic samples are covered in mineral oil, it is expected that the evaporation rates of the media underneath is less. This means that, for aerobic cells, the OD becomes artificially inflated as the same number of cells have become concentrated in less and less media. It is not known to what extent the evaporation effect has on these data. |
<center>[[Image:FLUORoverODoxygen.png|800px]]</center> | <center>[[Image:FLUORoverODoxygen.png|800px]]</center> | ||
| Line 286: | Line 284: | ||
===Characterization of Old BioBrick parts=== | ===Characterization of Old BioBrick parts=== | ||
| - | When designing our project, we were thinking about using the violacein pigment, BBa_K274002, in our genetic circuit. This part is supposed to be just the coding sequence | + | When designing our project, we were thinking about using the violacein pigment, BBa_K274002, in our genetic circuit. This part is supposed to be just the coding sequence. Because the tube in the picture below contains only this part with no promoter added, there should be no expression of violacein. However, as can be seen below, the violacein is present. This could be caused by either an undocumented promoter within the part, or a non-insulated BioBrick vector with inefficient flanking transcriptional terminators. |
<center>[[Image:Violacein.jpg|400px]]</center> | <center>[[Image:Violacein.jpg|400px]]</center> | ||
| Line 293: | Line 291: | ||
Two Lux-inducible promoters that were already on the registry (K091107 and K091146) were characterized. Because these promoters require AHL and LuxR in order to activate, a construct was created that consisted of a Lux promoter followed by RFP, and a constitutive promoter followed by the LuxR gene. LuxR was expected to be present in excess so that the limiting factor would be AHL. The coding sequences for LuxR and RFP were preceded by the standard RBS B0034. | Two Lux-inducible promoters that were already on the registry (K091107 and K091146) were characterized. Because these promoters require AHL and LuxR in order to activate, a construct was created that consisted of a Lux promoter followed by RFP, and a constitutive promoter followed by the LuxR gene. LuxR was expected to be present in excess so that the limiting factor would be AHL. The coding sequences for LuxR and RFP were preceded by the standard RBS B0034. | ||
| - | The AHL used was | + | The AHL used was OC6-homoserine lactone. It was added to samples in a 96-well plate in concentrations of 0, .1, 1, 10, 100 and 1000nM. |
| + | |||
| + | The results below show no trend in protein expression based on concentration of OC6-homoserine lactone. More research is warranted to provide information about what are the best conditions for chemically inducing the Lux promoters. | ||
<center>[[Image:1107_108.png|800px]]</center> | <center>[[Image:1107_108.png|800px]]</center> | ||
| Line 303: | Line 303: | ||
<center>[[Image:146_116_tetR.png|800px]]</center> | <center>[[Image:146_116_tetR.png|800px]]</center> | ||
| - | + | ||
| + | ====Inconsistent Sequencing Results==== | ||
| + | |||
| + | Certain parts on the registry have reportedly had inconsistent sequencing results, or not enough information is available to say for certain that the actual part is what its creators think it is. Below is a list of parts for which the Penn State iGEM team repeatedly received the same but inaccurate sequencing results. These findings have been posted on each part's registry page in order to hopefully start a discussion with other teams who may have experienced the same results. | ||
| + | |||
| + | constitutive promoter [http://partsregistry.org/Part:BBa_J23108:Experience#User_Reviews J23108] | ||
| + | |||
| + | |||
| + | RBS+LuxR [http://partsregistry.org/Part:BBa_J37033:Experience J37033] | ||
| + | |||
| + | |||
| + | Lux promoter [http://partsregistry.org/Part:BBa_K091107:Experience#User_Reviews K091107] | ||
Latest revision as of 03:00, 28 October 2010
| Home | Team | Official Team Profile | Project | Parts Submitted to the Registry | Notebook | Human Practices | Safety | Sponsors |
|---|
Contents |
Overall project
The Penn State Team combined quorum sensing and oxygen sensing promoters to construct a bacterial fireworks genetic circuit. We characterized the transcriptional regulation of the FNR oxygen sensing protein, which derepresses protein expression in the presence of oxygen. We then combined this oxygen sensing promoter with AHL sensing promoters in a positive and negative feedback loop system. While in the "ON" state, the circuit expresses an anaerobic fluorescent protein (AFP), which can emit blue light even when oxygen is not present. When a single cell senses oxygen, it produces the quorum sensing molecule AHL, which then diffuses to activate the circuit in nearby non-oxygenated cells, amplifying the output of the system. The fireworks genetic circuit has memory; once exposed to oxygen it will stay on even in the absence of oxygen.
Here is a simple example of a problem that our research with oxygen promoters and quorum sensing could solve:
Imagine a sealed glass vessel. Using our engineered microorganism as an oxygen sensor and light emitting reporter, we want to detect the presence of oxygen, which indicates the potential for spoilage of food, wine, or medicine. Jacketing the preserved contents of the vessel is a thin layer of porous media enmeshed with E. coli. The microorganism contains our plasmid which includes an oxygen-sensing promoter and quorum sensing genes.

In the event that oxygen leaks into the jar, the top layer of bacteria react by expressing AFP. However, this is a weak indicator. If only the top layer of cells is fluorescing, it will be hard to detect a change at all! Quorum sensing solves this problem by sending a signal to all the surrounding bacteria that will cause them to begin producing AFP.

In this way, only when oxygen-sensing and quorum-sensing are used together will the AFP indicator be a strong enough signal to measure.

This system could also be implemented in other chassis, such as yeast (wine) or Lactobacillus (yogurt), so the food itself could be the sensor. Genetically Modified Organisms (GMOs) in food would obviously face severe regulatory constraints. To better understand public perception, we created and administered a survey to over 250 randomized students in cooperation with other iGEM teams, especially RMIT. For details, see our Human Practices page.
Project Contributors
Penn State iGEM would like to give a big thank you to those who helped us through the process of our project. We really appreciate the help the following people have given us:
- Mike Kang
- Mike designed the oxygen promoters that deal with FNR in Dr. Salis' lab. With his help we were able to take his design and assemble the promoters.
- Trip Garland
- Trip designed the oxygen promoters that deal with arcA in Dr. Salis' lab. Although, we did not get to assemble his designed promoters, they were a potential use for our project.
- Collectively, Mike and Trip designed and constructed the anaerobic fluorescent protein expression vector.
- Dr. Leland Glenna
- Dr. Glenna helped guide us in the process of designing our Human Practices survey and also helped us with our statistical analysis of the results.
- Mike Speer
- Mike was the driving force behind our iGEM team to help us accomplish our goals while we worked in Dr. Richard's lab. Under his guidance, and tested patience, we overcame many obstacles to complete our tasks. Specifically, we would like to thank Mike for his help with the assembly of the Tet inverter to the cI repressor.
- RMIT-Australia
- RMIT collaborated with us by helping with our Human Practice's project. They distributed our survey to random students at their school, so we could better understand some of the different perspectives on Genetically Modified Organisms across the globe.
Project Details
Genetic Circuit
The Penn State iGEM team began work and research in mid May. After learning lab techniques from the grad students, we began researching possible topics. We decided to create an oxygen sensing promoter(s) to be coupled with a quorum sensing device.
Initially, we transformed Lux Promoters, Constitutive Promoters, LuxI + RBS, LuxR + RBS, Terminators, LacI, LuxR, pLac, cI, RBS, Pigments, FP, LacI + RBS, and cI + RBS. There were issues with several of these parts, which are documented in the [Characterization of Old BioBrick Parts] section.
For a better understanding of the parts used throughout our project, please refer to the tables below.
| Lux Promoters | Lux Promoters | Constitutive Promoters | LuxI + RBS | LuxR + RBS | Terminator | AFP | Tet Promoters |
|---|---|---|---|---|---|---|---|
| [http://partsregistry.org/Part:BBa_K091107 K091107] | [http://partsregistry.org/Part:BBa_R1062 R1062] | [http://partsregistry.org/Part:BBa_J23108 J23108] | [http://partsregistry.org/Part:BBa_C0061 C0061] | [http://partsregistry.org/Part:BBa_K376010 K376010] | [http://partsregistry.org/Part:BBa_B0010 B0010] | [http://partsregistry.org/Part:BBa_K376004 K376004] | [http://partsregistry.org/Part:BBa_R0040 R0040] |
| [http://partsregistry.org/Part:BBa_K091146 K091146] | [http://partsregistry.org/Part:BBa_R0063 R0063] | [http://partsregistry.org/Part:BBa_J23116 J23116] | [http://partsregistry.org/Part:BBa_B0012 B0012] | ||||
| [http://partsregistry.org/Part:BBa_J23100 J23100] | [http://partsregistry.org/Part:BBa_B1006 B1006] |
| LuxR | cI | RBS | RFP + RBS | LacI + RBS | cI + RBS | Tet Repressor | Super RBS | Anaerobic Promoters |
|---|---|---|---|---|---|---|---|---|
| [http://partsregistry.org/Part:BBa_C0062 C0062] | [http://partsregistry.org/Part:BBa_C0051 C0051] | [http://partsregistry.org/Part:BBa_B0033 B0033] | [http://partsregistry.org/Part:BBa_E1010 E1010] | [http://partsregistry.org/Part:BBa_J24679 J24679] | [http://partsregistry.org/Part:BBa_K081007 K081007] | [http://partsregistry.org/Part:BBa_C0040 C0040] | [http://partsregistry.org/Part:BBa_K376001 K376001] | [http://partsregistry.org/Part:BBa_K376002 K376002] |
| [http://partsregistry.org/Part:BBa_B0034 B0034] | [http://partsregistry.org/Part:BBa_P0451 P0451] |
| Oxygen Sensitive Promoters | AFP+RBS | AFP Generators | RFP Generators | Constitutive LuxR Generators | Tet Inverted Lambda Repressor |
|---|---|---|---|---|---|
| [http://partsregistry.org/Part:BBa_K376003 K376003] | [http://partsregistry.org/Part:BBa_K376004 K376004] | [http://partsregistry.org/Part:BBa_K376005 K376005] | [http://partsregistry.org/Part:BBa_K376008 K376008] | [http://partsregistry.org/Part:BBa_K376011 K376011] | [http://partsregistry.org/Part:BBa_K376019 K376019] |
| [http://partsregistry.org/Part:BBa_K376006 K376006] | [http://partsregistry.org/Part:BBa_K376009 K376009] | [http://partsregistry.org/Part:BBa_K376012 K376012] | [http://partsregistry.org/Part:BBa_K376025 K376025] | ||
| [http://partsregistry.org/Part:BBa_K376013 K376013] |
Here is an overview of our genetic circuit. Below, we will take a look at the function of each individual part in detail.

Our genetic circuit is designed to initially be in an anoxic state. However, the cells may be grown aerobically and then switched off using a Tet analog such ATC and/or doxycycline. The goal here was to create a circuit with a very strong "OFF" during anaerobic conditions, and a very strong "ON" during aerobic conditions. Anaerobic fluorescent protein (AFP) is used as an indicator.
cI mechanism

The Tet inverter behaves somewhat as a constitutive promoter when no promoter is in front of it. Following the Tet inverter is a cI coding sequence (which codes for the lambda repressor). The lambda repressor binds to the lux promoters in the front of the circuit, which are also lambda repressible. This, coupled with the fact that the conditions are anaerobic, provides a very strong "OFF" switch that does not allow the expression of the Anaerobic Fluorescent Protein (AFP) to occur.
FNR breaks apart in oxygen

FNR can act as a repressor or inducer depending on its position relative to the RBS. In nature, an activated FNR dimer represses aerobic functions while inducing anaerobic functions.
In the event that conditions change to aerobic in our circuit, oxygen molecules break up the dimer formed by FNR, causing the FNR dimer to detach from the oxygen promoter. Assuming that the oxygen promoter used in this assembly is a repressor, the end result will be that the coding sequences following the promoter will begin to be expressed.
Ptet is repressed, and the positive feedback loop is switched on

The oxygen promoter is followed by 3 protein coding sequences. Two of them are genes that code for the Tet repressor and LuxI (which produces [N-]acyl-homoserine lactone, or AHL). These two parts serve to strengthen the genetic switch once it has been turned on. The Tet repressor binds to the Tet inverter, causing the expression of the lambda repressor to be switched off. Meanwhile, AHL production resulting from the LuxI gene couples with the constitutively-expressed LuxR in order to activate the Lux promoter at the beginning of the circuit.
In addition, the Anaerobic Fluorescent Protein begins to be expressed at a high rate.
AFP expression ramps up

Because both the oxygen promoter and the Lux promoter are turned on as part of the positive feedback loop, AFP is now expressing at a very high rate. This is also bolstered by the fact that that the Super RBS is in front of this protein.
The Experiments
Oxygen Sensor
The design of the oxygen-sensing promoter was based on the dcuC gene of E. coli. The two binding sites for FNR surrounded the RNAP binding site and were systematically shifted upstream and downstream from this site in multiple constructs, causing overlap of the RNAP binding site in some cases. Using this strategy, five constructs were designed with the constitutive promoter J23113 as the basis for the sequence following the downstream FNR binding site. The construct that worked best as an oxygen-sensing promoter (J6, http://partsregistry.org/Part:BBa_K376003 BBa_J376003) was fully characterized and is shown below.

The oxygen promoters were characterized using a Tecan infinite M1000 microplate reader. Each sample was grown overnight on plates and transferred to a 96 deep-well plate for overnight growth. After 16 hours the samples were tested in the Tecan, where measurements would be taken for around 12 hours. Sometimes, measurement of growth of three successive plates (serially diluted from the preceding one) was needed before the appropriate steady state could be recorded.
Results
All measurements were taken using the Tecan Infinite M 1000 spectrophotometer. The following is a graph of the raw fluorescence values recorded by the Tecan for two oxygen promoters, D5 and J6. Each construct was duplicated in the 96-well plate. One was allowed to grow normally under aerobic conditions, while the other sample was covered in 100uL of mineral oil to create an anaerobic environment. Serial dilution of two plates resulted in 3 sets of data for this experiment, as seen below.

The next graph shown is the Raw data for OD for the same constructs shown above. The OD is proportional to the number of cells growing in the media, so dividing the fluorescence by these values is approximately proportional to the fluorescence per cell.

Shown below is the Fluorescence over OD of each sample. As the samples reach steady state, the graph should become a horizontal line. Notice how the graph comes closer and closer to reaching this steady state with each successive plate. Also, it is important to remember that evaporation comes into play here when comparing anaerobic and anaerobic samples with each other. Since the anaerobic samples are covered in mineral oil, it is expected that the evaporation rates of the media underneath is less. This means that, for aerobic cells, the OD becomes artificially inflated as the same number of cells have become concentrated in less and less media. It is not known to what extent the evaporation effect has on these data.

Next, the fold-change was calculated for each sample. That is, the fluorescence/OD for anaerobic growth of the sample was divided bye fluorescence/OD for the aerobic sample. The same was done for a constitutive promoter expressing AFP. That way, a comparison could be made between constant expression of AFP in an anoxic environment and induced expression.
Oxygen Promoter J6

Oxygen Promoter D5

The results are that both promoters indeeed were inducible under anaerobic conditions. The fold-change expression when switching from aerobic to anaerobic is significantly higher than the fold-change expression of the constitutive promoter.

Characterization of Old BioBrick parts
When designing our project, we were thinking about using the violacein pigment, BBa_K274002, in our genetic circuit. This part is supposed to be just the coding sequence. Because the tube in the picture below contains only this part with no promoter added, there should be no expression of violacein. However, as can be seen below, the violacein is present. This could be caused by either an undocumented promoter within the part, or a non-insulated BioBrick vector with inefficient flanking transcriptional terminators.

Lux Promoter Characterization
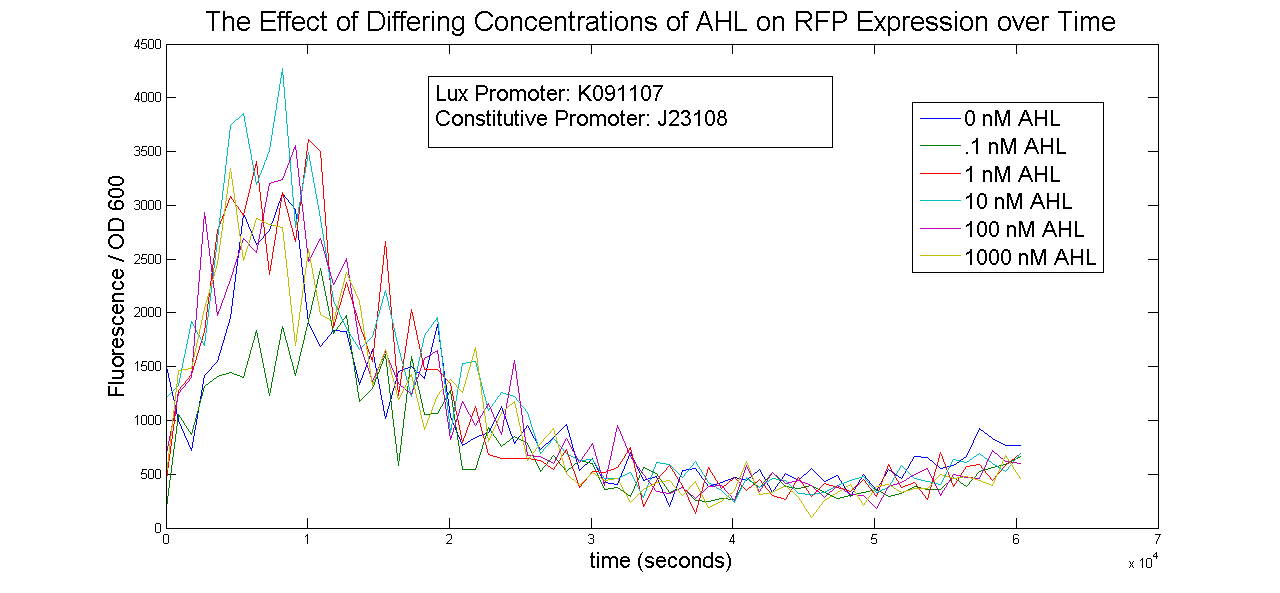
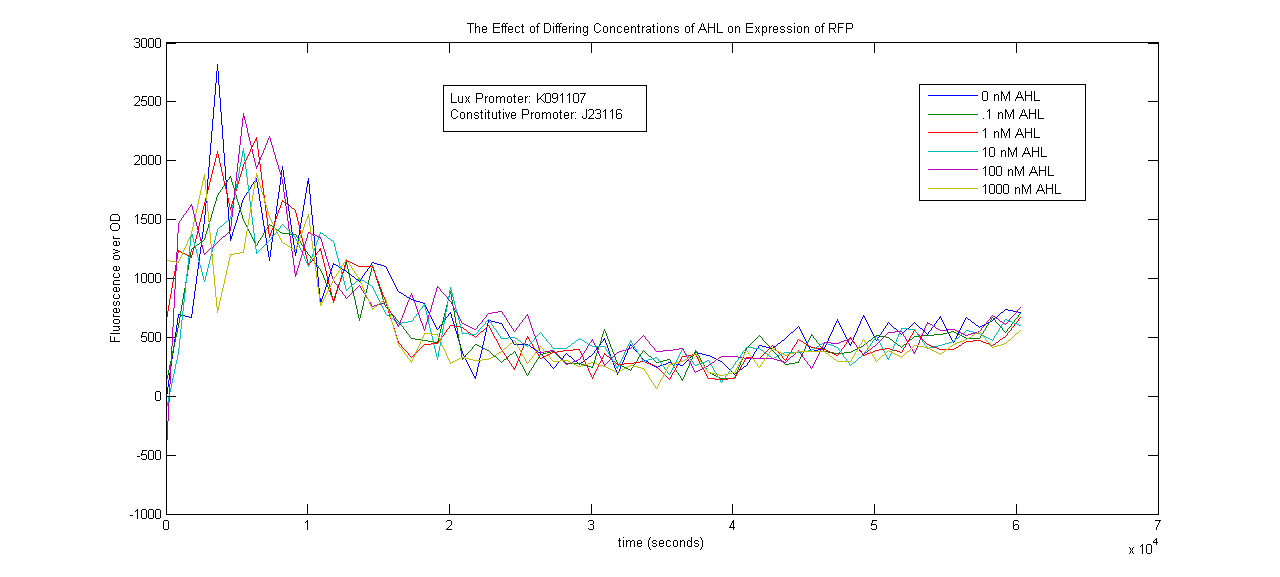
Two Lux-inducible promoters that were already on the registry (K091107 and K091146) were characterized. Because these promoters require AHL and LuxR in order to activate, a construct was created that consisted of a Lux promoter followed by RFP, and a constitutive promoter followed by the LuxR gene. LuxR was expected to be present in excess so that the limiting factor would be AHL. The coding sequences for LuxR and RFP were preceded by the standard RBS B0034.
The AHL used was OC6-homoserine lactone. It was added to samples in a 96-well plate in concentrations of 0, .1, 1, 10, 100 and 1000nM.
The results below show no trend in protein expression based on concentration of OC6-homoserine lactone. More research is warranted to provide information about what are the best conditions for chemically inducing the Lux promoters.


Inconsistent Sequencing Results
Certain parts on the registry have reportedly had inconsistent sequencing results, or not enough information is available to say for certain that the actual part is what its creators think it is. Below is a list of parts for which the Penn State iGEM team repeatedly received the same but inaccurate sequencing results. These findings have been posted on each part's registry page in order to hopefully start a discussion with other teams who may have experienced the same results.
constitutive promoter [http://partsregistry.org/Part:BBa_J23108:Experience#User_Reviews J23108]
RBS+LuxR [http://partsregistry.org/Part:BBa_J37033:Experience J37033]
Lux promoter [http://partsregistry.org/Part:BBa_K091107:Experience#User_Reviews K091107]
 "
"