Team:UNIPV-Pavia/Parts/Characterization/ExistingPartsRegistry
From 2010.igem.org
| (5 intermediate revisions not shown) | |||
| Line 9: | Line 9: | ||
{{UNIPV-Pavia/Style}} | {{UNIPV-Pavia/Style}} | ||
<table border="0" align="center" width="100%"><tr><td align="justify" valign="top" style="padding:20px"> | <table border="0" align="center" width="100%"><tr><td align="justify" valign="top" style="padding:20px"> | ||
| - | <html><p align="center"><font size="4"><b>EXISTING PARTS FROM THE REGISTRY</b></font></p></html><hr> | + | <html><p align="center"><font size="4"><b>EXISTING PARTS FROM THE REGISTRY<br>(all these reported informations are shared in the Registry of Standard Parts)</b></font></p></html><hr> |
<tr><td width="100%"> | <tr><td width="100%"> | ||
| Line 160: | Line 160: | ||
=<partinfo>BBa_K300009</partinfo>/<partinfo>BBa_I4102</partinfo> - PoPS->3OC6HSL sender device= | =<partinfo>BBa_K300009</partinfo>/<partinfo>BBa_I4102</partinfo> - PoPS->3OC6HSL sender device= | ||
| + | <partinfo>BBa_K300009</partinfo> and <partinfo>BBa_K300010</partinfo> was assembled downstream of the promoters the table reported in, thus obtaining the following parts: | ||
| + | |||
| + | {| border='1' align='center' | ||
| + | | '''BioBrick''' ||'''Description''' | ||
| + | |- | ||
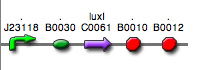
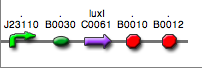
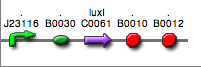
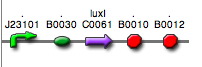
| + | | <partinfo>BBa_K300030</partinfo>|| [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23118 | ||
| + | |- | ||
| + | | <partinfo>BBa_K300028</partinfo>|| [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23110 | ||
| + | |- | ||
| + | | <partinfo>BBa_K300029</partinfo>|| [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23116 | ||
| + | |- | ||
| + | | <partinfo>BBa_K300025</partinfo>|| [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23101 | ||
| + | |- | ||
| + | | <partinfo>BBa_K300026</partinfo>|| [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23105 | ||
| + | |- | ||
| + | | <partinfo>BBa_K300027</partinfo>|| [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23106 | ||
| + | |} | ||
| + | |||
| + | For each part, a measurement system was built, exploiting the production of the reporter gene GFP (Green Fluorescent Protein) by <partinfo>BBa_F2620</partinfo> receiver device, to evaluate the "switch on" condition of every self-inducible promoter. Many different combinations were explored, in order to provide a library of promoters able to initiate transcription at the desired culture density. | ||
| + | |||
| + | ===<b>Quantification of the HSL produced by autoinducer generators</b>=== | ||
| + | |||
| + | '''Experimental implementation:''' The autoinducer generators <partinfo>BBa_K300030</partinfo>, <partinfo>BBa_K300028</partinfo>, <partinfo>BBa_K300029</partinfo>, <partinfo>BBa_K300025</partinfo>, <partinfo>BBa_K300026</partinfo> and <partinfo>BBa_K300027</partinfo> were, thus, characterized by measuring the concentration of HSL released in the medium of cultures grown for 6 hours. All the details are available in [[Team:UNIPV-Pavia/Parts/Characterization#Microplate reader experiments for 3OC6-HSL quantification by means of BBa_T9002 biosensor - Protocol #3|this section]]. | ||
| + | |||
| + | <partinfo>BBa_T9002</partinfo> contained in <partinfo>pSB1A3</partinfo> in ''E. coli'' TOP10 was used as a HSL->GFP biosensor. In every experiment, a HSL-GFP calibration curve with known concentration of HSL was produced. | ||
| + | |||
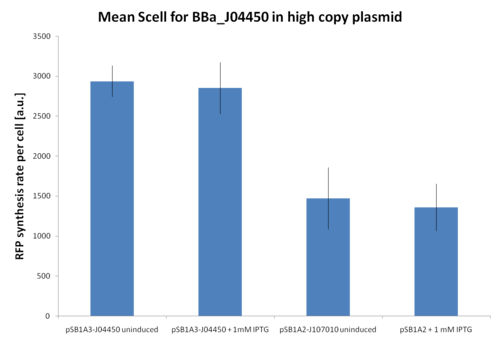
| + | '''Results:''' The amount of 3OC6-HSL produced after a 6 hours growth by ''E. coli'' DH5alpha bearing the parts in high copy plasmid <partinfo>pSB1A2</partinfo> is reported in Fig.8 and in the table below: | ||
| + | |||
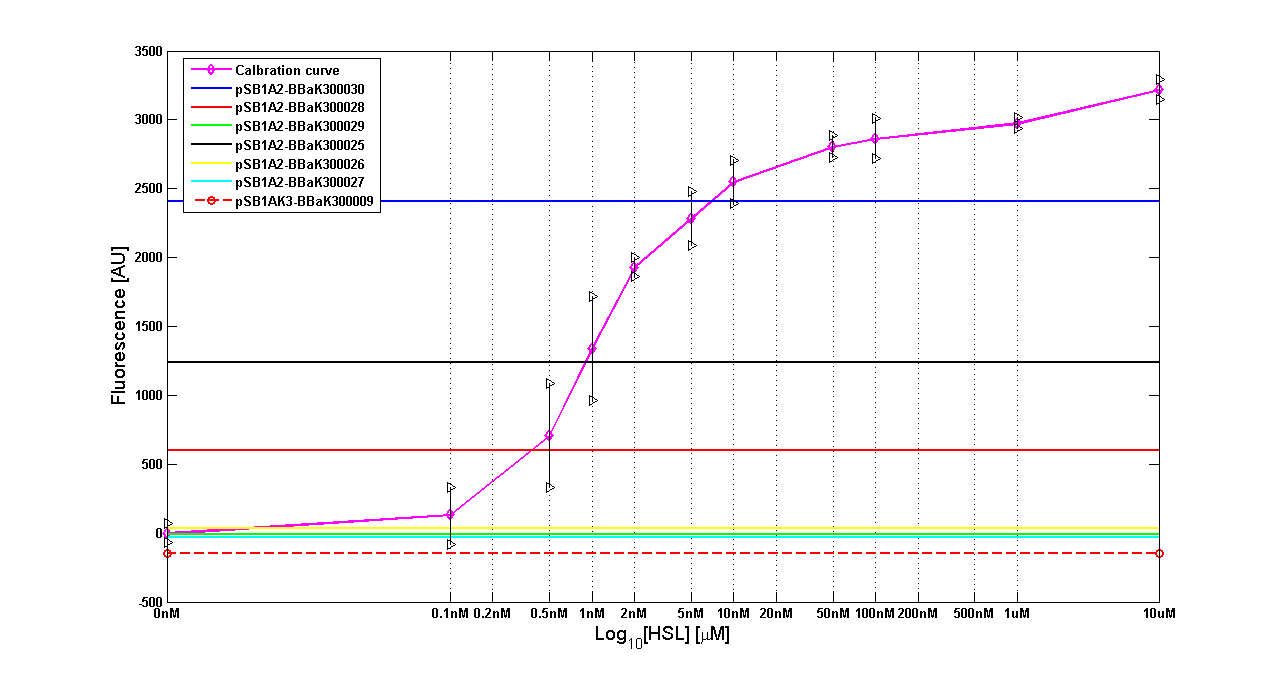
| + | {| align='center' | ||
| + | |[[Image:pv_HCT9002sensor.png|500px|thumb|center|Figure 8 - <partinfo>BBa_T9002</partinfo> calibration curve for detection of [HSL] produced by autoinducer generators in high copy vector.]] | ||
| + | |} | ||
| + | |||
| + | {| border='1' align='center' | ||
| + | | ''BioBrick'' || ''Wiki name''|| E. coli ''strain'' || [HSL] | ||
| + | |- | ||
| + | | <partinfo>BBa_K300030</partinfo> || I14|| DH5alpha || 0.7 uM | ||
| + | |- | ||
| + | | <partinfo>BBa_K300028</partinfo> || I15|| DH5alpha || 0.04 uM | ||
| + | |- | ||
| + | | <partinfo>BBa_K300029</partinfo> || I16|| DH5alpha || not detected | ||
| + | |- | ||
| + | | <partinfo>BBa_K300025</partinfo> || I17|| DH5alpha || 0.09 uM | ||
| + | |- | ||
| + | | <partinfo>BBa_K300026</partinfo> || I18|| DH5alpha || not detected | ||
| + | |- | ||
| + | | <partinfo>BBa_K300027</partinfo> || I19|| DH5alpha || 0.002 uM | ||
| + | |} | ||
| + | |||
| + | The amount of 3OC6-HSL produced by the parts contained in low copy plasmid <partinfo>pSB4C5</partinfo> after a 6 hour cell growth is reported in figure and in the table below: | ||
| + | |||
| + | {| align='center' | ||
| + | |[[Image:pv_LCT9002sensor.png|500px|thumb|center|Figure 9 - <partinfo>BBa_T9002</partinfo> calibration curve for detection of [HSL] produced by the autoinducer generators in low copy vector.]] | ||
| + | |} | ||
| + | |||
| + | |||
| + | {| border='1' align='center' | ||
| + | | ''BioBrick'' || ''Wiki name''|| E. coli ''strain'' || [HSL] | ||
| + | |- | ||
| + | | <partinfo>BBa_K300030</partinfo> || I14|| DH5alpha || 0.005 uM | ||
| + | |- | ||
| + | | <partinfo>BBa_K300028</partinfo> || I15|| DH5alpha || 0.002 uM | ||
| + | |- | ||
| + | | <partinfo>BBa_K300029</partinfo> || I16|| DH5alpha || not detected | ||
| + | |- | ||
| + | | <partinfo>BBa_K300025</partinfo> || I17|| DH5alpha || 0.003 uM | ||
| + | |- | ||
| + | | <partinfo>BBa_K300026</partinfo> || I18|| DH5alpha || not detected | ||
| + | |- | ||
| + | | <partinfo>BBa_K300027</partinfo> || I19|| DH5alpha || not detected | ||
| + | |} | ||
| + | |||
| + | '''Discussion''' These experiments provided extremely useful informations about the capability of the signal generators to produce the 3OC6-HSL signal molecule. Data are quantitative, but incomplete because for weak promoters or medium-strength promoters contained in a low copy number plasmid the amount of 3OC6-HSL was not detectable using this system. However, this simple experiment shows that there is a strong correlation between the strength of promoter and the amount of signal molecule produced. These results confirm that the production of the autoinducer can be engineered in ''E. coli'' and different expression systems reach different amounts of 3OC6-HSL in the growth media as a function of the promoter strength. Thus, these results demonstrate that self-inducible circuits can be rationally designed from a set of well characterized standard parts. | ||
| + | |||
| + | ===<b>Modulation of plasmid copy number</b>=== | ||
| + | |||
| + | The signal generator was assembled on a low copy number plasmid (<partinfo>pSB4C5</partinfo>) and the receiver device on high copy number plasmid (<partinfo>pSB1A2</partinfo>). | ||
| + | The circuits we obtained and tested are summarized in tables. | ||
| + | |||
| + | |||
| + | <div align='center>Sender/Receiver devices assembled as a unique BioBrick part on the same vector</div> | ||
| + | |||
| + | <div align='center>Sender devices assembled on low copy number vector and Receiver device on high copy number vector</div> | ||
| + | |||
| + | {| border='1' align='center' | ||
| + | | '''BioBrick'''<br> '''Sender''' ||'''Description ''' || '''Sender Vector''' || '''<partinfo>BBa_F2620</partinfo><br> Receiver vector''' | ||
| + | |- | ||
| + | | <partinfo>BBa_K300030</partinfo> | ||
| + | | [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23118 | ||
| + | |<partinfo>pSB4C5</partinfo><br>LC | ||
| + | | <partinfo>pSB1A2</partinfo><br>HC | ||
| + | |- | ||
| + | | <partinfo>BBa_K300028</partinfo> | ||
| + | | [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23110 | ||
| + | |<partinfo>pSB4C5</partinfo><br>LC | ||
| + | | <partinfo>pSB1A2</partinfo><br>HC | ||
| + | |- | ||
| + | | <partinfo>BBa_K300029</partinfo> | ||
| + | | [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23116 | ||
| + | |<partinfo>pSB4C5</partinfo><br>LC | ||
| + | | <partinfo>pSB1A2</partinfo><br>HC | ||
| + | |- | ||
| + | | <partinfo>BBa_K300025</partinfo> | ||
| + | | [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23101 | ||
| + | |<partinfo>pSB4C5</partinfo><br>LC | ||
| + | | <partinfo>pSB1A2</partinfo><br>HC | ||
| + | |- | ||
| + | | <partinfo>BBa_K300026</partinfo> | ||
| + | | [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23105 | ||
| + | |<partinfo>pSB4C5</partinfo><br>LC | ||
| + | | <partinfo>pSB1A2</partinfo><br>HC | ||
| + | |- | ||
| + | | <partinfo>BBa_K300027</partinfo> | ||
| + | | [[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23106 | ||
| + | |<partinfo>pSB4C5</partinfo><br>LC | ||
| + | | <partinfo>pSB1A2</partinfo><br>HC | ||
| + | |} | ||
| + | |||
| + | ===<b>Results</b>=== | ||
| + | |||
| + | The following measurement systems were realized assembling GFP downstream of the receiver circuit. The parts characterized are reported in this table: | ||
| + | |||
| + | {| border='1' align='center' width='80%' | ||
| + | | '''Sender device''' | ||
| + | | '''Sensor systems with GFP''' | ||
| + | |'''Measurement Device''' | ||
| + | |- | ||
| + | |<partinfo>BBa_K300030</partinfo> in <partinfo>pSB4C5</partinfo><br>[[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23118 | ||
| + | |<partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo><br>[[Image:pv_T9002.png|300px]] | ||
| + | |Sender and Receiver are contained <br>in two different plasmids, <br>cotransformed in the same cell | ||
| + | |- | ||
| + | |<partinfo>BBa_K300028</partinfo> in <partinfo>pSB4C5</partinfo><br>[[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23110 | ||
| + | |<partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo><br>[[Image:pv_T9002.png|300px]] | ||
| + | |Sender and Receiver are contained <br>in two different plasmids, <br>cotransformed in the same cell | ||
| + | |- | ||
| + | |<partinfo>BBa_K300029</partinfo> in <partinfo>pSB4C5</partinfo><br>[[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23116 | ||
| + | |<partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo><br>[[Image:pv_T9002.png|300px]] | ||
| + | |Sender and Receiver are contained<br> in two different plasmids, <br>cotransformed in the same cell | ||
| + | |- | ||
| + | |<partinfo>BBa_K300026</partinfo> in <partinfo>pSB4C5</partinfo><br>[[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23105 | ||
| + | |<partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo><br>[[Image:pv_T9002.png|300px]] | ||
| + | |Sender and Receiver are contained<br> in two different plasmids, <br>cotransformed in the same cell | ||
| + | |- | ||
| + | |<partinfo>BBa_K300025</partinfo> in <partinfo>pSB4C5</partinfo><br>[[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23101 | ||
| + | |<partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo><br>[[Image:pv_T9002.png|300px]] | ||
| + | |Sender and Receiver are contained<br> in two different plasmids, <br>cotransformed in the same cell | ||
| + | |- | ||
| + | |<partinfo>BBa_K300027</partinfo> in <partinfo>pSB4C5</partinfo><br>[[Image:pv_SignalGeneratorDevice.png|150px]]<br>J23106 | ||
| + | |<partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo><br>[[Image:pv_T9002.png|300px]] | ||
| + | |Sender and Receiver are contained<br> in two different plasmids, <br>cotransformed in the same cell | ||
| + | |} | ||
| + | |||
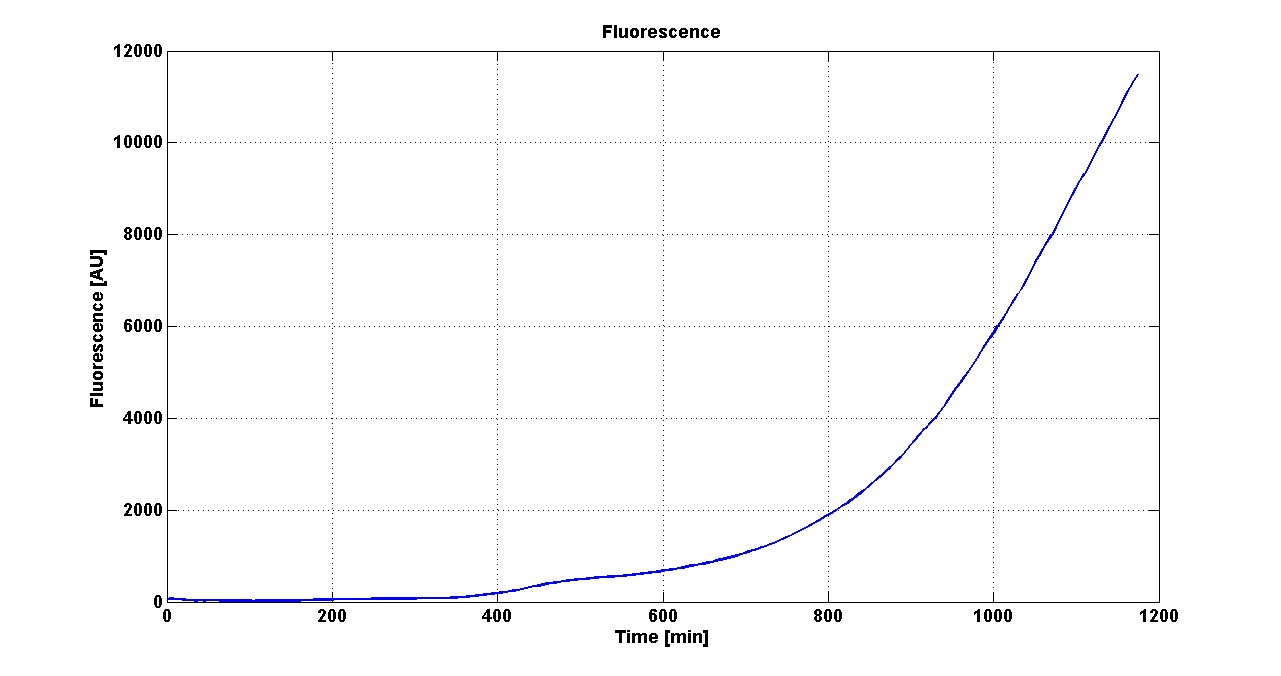
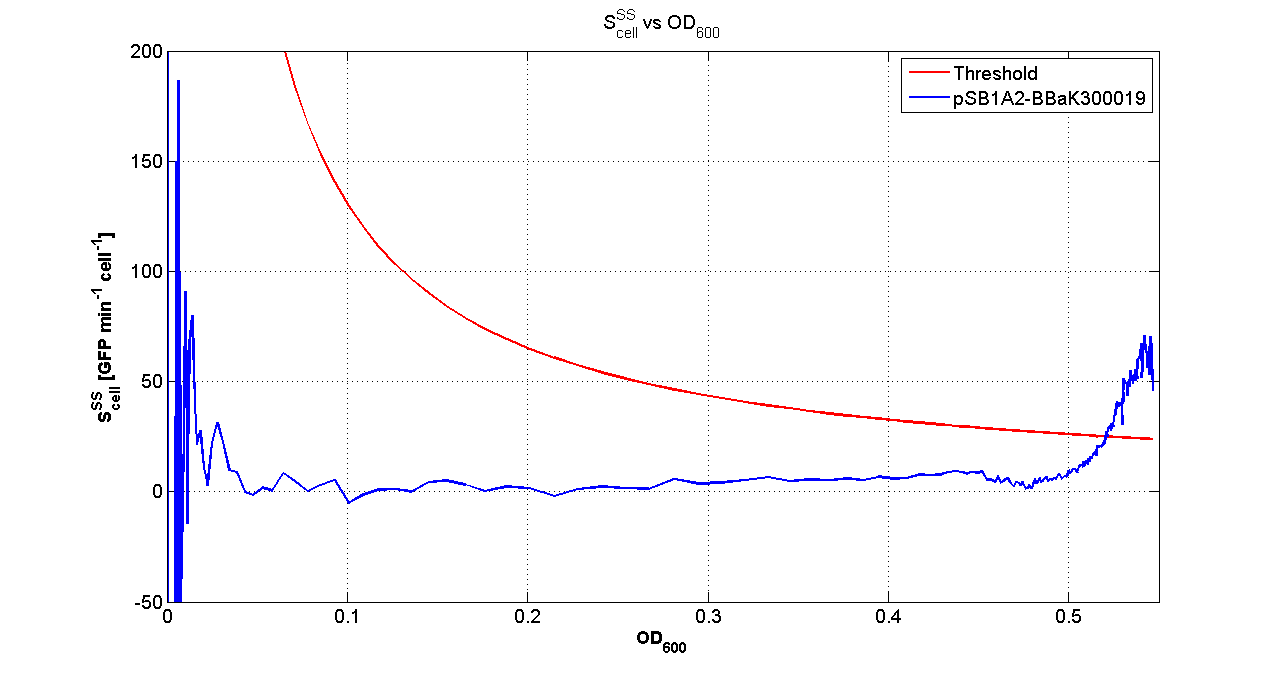
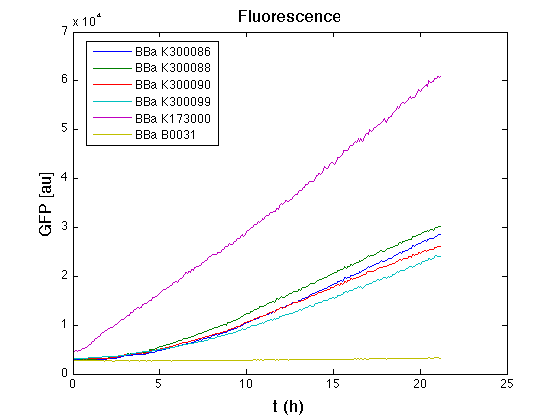
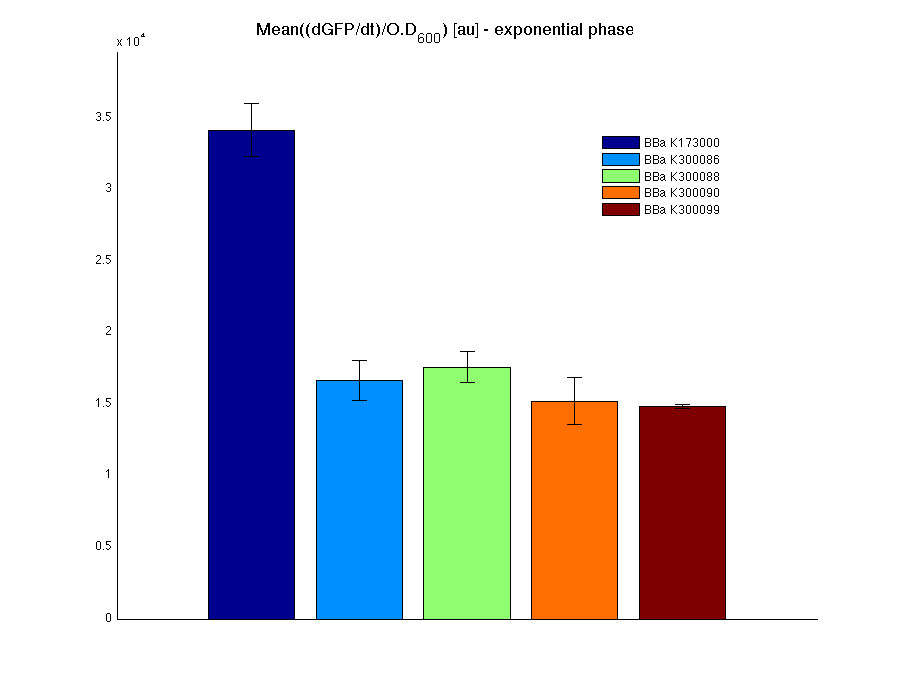
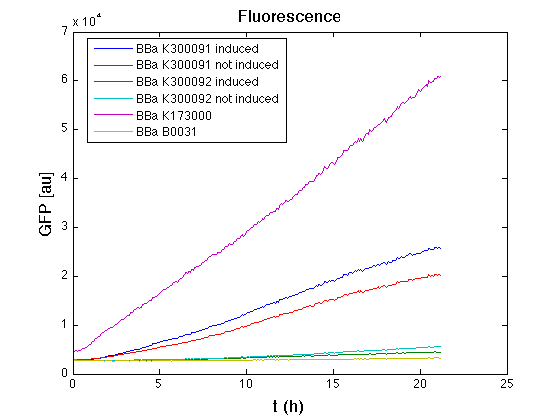
| + | Cultures of ''E. coli'' TOP10 bearing the plasmids containing the self-inducible devices expressing GFP were grown according to [[Team:UNIPV-Pavia/Parts/Characterization#Microplate reader experiments for self-inducible promoters - Protocol #1|this protocol]] and all data collected were analyzed as explained in [[Team:UNIPV-Pavia/Parts/Characterization#Data analysis for self-inducible promoters (initiation-treshold determination)|this section]]. An example of O.D.600 and fluorescence signals for a self-inducible device expressing GFP (<partinfo>BBa_K300026</partinfo>), as well as its Scell signal and the estimated threshold value, is reported below. | ||
| + | |||
| + | {| | ||
| + | |[[Image:pv_GrowthCurveSelf.png|300px|thumb|center|Growth curve of <partinfo>BBa_K300019</partinfo> (O.D.600)]] | ||
| + | |[[Image:pv_FluoCurveSelf.png|300px|thumb|center|Fluorescence curve of <partinfo>BBa_K300019</partinfo> (G.F.P.)]] | ||
| + | |- | ||
| + | |[[Image:pv_FLUvsASB.png|300px|thumb|center|Fluorescence VS Optical density curve of <partinfo>BBa_K300019</partinfo>]] | ||
| + | |[[Image:pv_Scell_Threshold.png|300px|thumb|center|Scell=(dGFP/dt)/O.D.600 and threshold]] | ||
| + | |} | ||
| + | |||
| + | For every self-inducible device, several parameters were evaluated: | ||
| + | *O.D.start is the O.D.600 corresponding to the transcription initiation of the gene of interest; it was evaluated as reported [[Team:UNIPV-Pavia/Parts/Characterization#Data analysis for self-inducible promoters (initiation-treshold determination)|in this section]]; | ||
| + | *K_HSL is the HSL synthesis rate per cell; it was estimated with the algorithm described [[Team:UNIPV-Pavia/Parts/Characterization#Data analysis to estimate the HSL synthesis rate per cell|here]] | ||
| + | *Doubling time is the period of time required for a cell population to double; it was evaluated as described in [[Team:UNIPV-Pavia/Parts/Characterization#Doubling time evaluation|Doubling time evaluation section]] | ||
| + | *Scell_ratio was evaluated as (Scell_max_Phi)/(Scell_max_J101). Phi is the self-inducible device of ineterst, J101 is the reference standard <partinfo>BBa_J23101</partinfo> contained in the same vector of the receiver device. Scell_max_phi was evaluated for times subsequent to the transcription initiation. | ||
| + | |||
| + | Results are summarized in the following tables: | ||
| + | |||
| + | '''Tab. 1 - Sender on low copy plasmid <partinfo>pSB4C5</partinfo> and Receiver on high copy plasmid <partinfo>pSB1A3</partinfo>''' | ||
| + | |||
| + | {| border='1' width='80%' | ||
| + | |rowspan='2' align='center'|'''Self-inducible device''' <br><font color='#50C878'>working</font><br><font color='red'>not working</font> | ||
| + | |colspan='4' style="background: yellow" align='center' |LB | ||
| + | |colspan='4' style="background: cyan" align="center" |M9 | ||
| + | |- | ||
| + | |align='center'| '''O.D.start''' | ||
| + | |align='center'| '''K_HSL'''<br> [nmol/min] | ||
| + | |align='center'| '''Doubling time'''<br>[min] | ||
| + | |align='center'| '''Scell ratio''' | ||
| + | |align='center'|'''O.D.start''' | ||
| + | |align='center'| '''K_HSL'''<br> [nmol/min] | ||
| + | |align='center'| '''Doubling time'''<br>[min] | ||
| + | |align='center'|'''Scell ratio''' | ||
| + | |- | ||
| + | |<table><tr><td><font color='#50C878'><partinfo>BBa_K300030</partinfo><br> (wiki name: I14) <br>in <partinfo>pSB4C5</partinfo> plasmid</font> | ||
| + | <br></td><td><partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo></td></tr> | ||
| + | <tr><td>[[Image:pv_BBa_K300030.png|170px]]<br>LC</td> | ||
| + | <td>[[Image:pv_BBa_F2620.png|170px]]<br>HC</td></tr></table> | ||
| + | |Constitutive | ||
| + | | 2.92 10^-16 <br> ± <br> 8.16 10^-18 | ||
| + | | 33.20 <br> ± <br> 1.46 | ||
| + | | 1.20 <br> ± <br> 0.20 | ||
| + | | 0.05 <br> ± <br> 0.005 | ||
| + | | 8.06 10^-16 <br> ± <br> 1.28 10^-16 | ||
| + | | 58.27 <br> ± <br> 5.66 | ||
| + | | 0.91 <br> ± <br> 0.15 | ||
| + | |- | ||
| + | |<table><tr><td><font color='#50C878'><partinfo>BBa_K300028</partinfo><br> (wiki name: I15)<br> in <partinfo>pSB4C5</partinfo> plasmid</font><br></td><td><partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo></td></tr> | ||
| + | <tr><td>[[Image:pv_BBa_K300028.png|170px]]<br>LC</td> | ||
| + | <td>[[Image:pv_BBa_F2620.png|170px]]<br>HC</td></tr></table> | ||
| + | | 0.32 <br> ± <br> 0.04 ** | ||
| + | | 8.31 10^-17 <br> ± <br> 3.92 10^-17 ** | ||
| + | | 33.24 <br> ± <br> 1.27 | ||
| + | | 0.61 <br> ± <br> 0.04 ** | ||
| + | | 0.15 <br> ± <br> 0.007 | ||
| + | | 1.72 10^-16 <br> ± <br> 6.65 10^-18 | ||
| + | | 65.57 <br> ± <br> 5.99 | ||
| + | | 0.26 <br> ± <br> 0.03 | ||
| + | |- | ||
| + | |<table><tr><td><font color='#50C878'><partinfo>BBa_K300029</partinfo> (wiki name: I16) in <partinfo>pSB4C5</partinfo> plasmid</font> | ||
| + | </td><td><partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo></td></tr> | ||
| + | <tr><td>[[Image:pv_BBa_K300029.png|170px]]<br>LC</td> | ||
| + | <td>[[Image:pv_BBa_F2620.png|170px]]<br>HC</td></tr></table> | ||
| + | | 0.31 <br> * | ||
| + | | 1.17 10^-16 <br> * | ||
| + | | 35.46 <br> ± <br> 2.85 | ||
| + | | 0.57 <br> * | ||
| + | | <font color='red' size='+2'>X</font> | ||
| + | | <font color='red' size='+2'>X</font> | ||
| + | | <font color='red' size='+2'>X</font> | ||
| + | | <font color='red' size='+2'>X</font> | ||
| + | |- | ||
| + | |<table><tr><td><font color='#50C878'><partinfo>BBa_K300025</partinfo><br> (wiki name: I17) <br>in <partinfo>pSB4C5</partinfo> plasmid</font> | ||
| + | </td><td><partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo></td></tr> | ||
| + | <tr><td>[[Image:pv_BBa_K300025.png|170px]]<br>LC</td> | ||
| + | <td>[[Image:pv_BBa_F2620.png|170px]]<br>HC</td></tr></table> | ||
| + | | Constitutive | ||
| + | | 2.85 10^-16 <br> ± <br> 1.90 10^-17 | ||
| + | | 33.27 <br> ± <br> 2.85 | ||
| + | | 1.09 <br> ± <br> 0.28 | ||
| + | | 0.1 <br> ± <br> 0.01 | ||
| + | | 3.81 10^-16 <br> ± <br> 4.68 10^-17 | ||
| + | | 59.02 <br> ± <br> 8.28 | ||
| + | | 0.45 <br> ± <br> 0.08 | ||
| + | |- | ||
| + | |<table><tr><td><font color='#50C878'><partinfo>BBa_K300026</partinfo><br> (wiki name: I18) <br>in <partinfo>pSB4C5</partinfo> plasmid</font> | ||
| + | </td><td><partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo></td></tr> | ||
| + | <tr><td>[[Image:pv_BBa_K300026.png|170px]]<br>LC</td> | ||
| + | <td>[[Image:pv_BBa_F2620.png|170px]]<br>HC</td></tr></table> | ||
| + | | <font color='red' size='+2'>X</font> | ||
| + | | <font color='red' size='+2'>X</font> | ||
| + | | 34.63 <br> ± <br> 1.31 | ||
| + | | <font color='red' size='+2'>X</font> | ||
| + | | 0.53 <br> ± <br> 0.03 ** | ||
| + | | 1.78 10^-17 <br> ± <br> 1.36 10^-18 ** | ||
| + | | 61.68 <br> ± <br> 7.08 | ||
| + | | 0.03 <br> ± <br> 0.002 ** | ||
| + | |- | ||
| + | |<table><tr><td><font color='#50C878'><partinfo>BBa_K300027</partinfo><br> (wiki name: I19)<br> in <partinfo>pSB4C5</partinfo> plasmid</font> | ||
| + | </td><td><partinfo>BBa_T9002</partinfo> in <partinfo>pSB1A3</partinfo></td></tr> | ||
| + | <tr><td>[[Image:pv_BBa_K300027.png|170px]]<br>LC</td> | ||
| + | <td>[[Image:pv_BBa_F2620.png|170px]]<br>HC</td></tr></table> | ||
| + | | <font color='red' size='+2'>X</font> | ||
| + | | <font color='red' size='+2'>X</font> | ||
| + | | 33.80 <br> ± <br> 1.78 | ||
| + | | <font color='red' size='+2'>X</font> | ||
| + | | 0.33 <br> ± <br> 0.05 | ||
| + | | 5.46 10^-17 <br> ± <br> 7.86 10^-18 | ||
| + | | 52.84 <br> ± <br> 4.29 | ||
| + | | 0.06 <br> ± <br> 0.002 | ||
| + | |} | ||
| + | |||
| + | <div id="box" style="width: 70%; margin-left: 137px; padding: 5px; border: 1px solid #000; background-color: #f6f6f6;"> | ||
| + | <div id="template" style="text-align: center; font-weight: bold; font-size: large; color: black; padding: 5px;"> | ||
| + | LEGEND OF TABLES: | ||
| + | </div> | ||
| + | <div id="legend" style="font-weight: normal; font-size: small; color: black; padding: 5px;"> | ||
| + | ''Constitutive'': the induction point, in term of O.D.600, is under the minimum detectable value calculated by the algorithm. This minimum value was estimated by running the algorithm on data acquired from a culture that constitutively produces GFP. For this reason, the devices labelled as ''constitutive'' can be considered as constitutive GFP producers. | ||
| + | |||
| + | <nowiki>*</nowiki>: in two of three experiments the self-induction failed, thus having a non-induced culture for all the cell densities. The standard errors were not computed for these cultures. | ||
| + | |||
| + | <nowiki>**</nowiki>: in one of three experiments the self-induction failed, thus having a non-induced culture for all the cell densities. The standard errors were computed on two independent experiments. | ||
| + | |||
| + | <partinfo>BBa_K300016</partinfo> is labelled with <nowiki>*</nowiki>, but probably induction failed in two of the three experiments because the culture didn't reach the O.D.start point (the experiment was stopped before the culture reached the O.D.600 critical value). | ||
| + | </div> | ||
| + | </div> | ||
| + | |||
| + | '''Discussion''': two modular PoPS-based devices (<partinfo>BBa_K300010</partinfo> and <partinfo>BBa_K300009</partinfo>/<partinfo>BBa_F2620</partinfo>) were designed and used to realize a library of self-inducible devices, able to start the production of the heterologous protein at a defined culture density. They were characterized in many different experimental conditions: | ||
| + | *varying the strength of the promoter controlling the production of the signal molecule (Sender Modulation) | ||
| + | *varying the copy number of vectors bearing both Sender and Receiver circuits | ||
| + | *varying the growth medium (LB or M9) | ||
| + | |||
| + | A graphical summary is reported in the figures below: | ||
| + | {| | ||
| + | |[[Image:pv_SwitchPointLB.png|700px|thumb|center| Typical ''E. coli'' growth curve with O.D.start evaluated by Threshold algorithm in LB]] | ||
| + | |- | ||
| + | |[[Image:pv_SwitchPointM9.png|700px|thumb|center| Typical ''E. coli'' growth curve with O.D.start evaluated by Threshold algorithm in M9]] | ||
| + | |} | ||
| + | |||
| + | A model-based approach was proposed to estimate many interesting parameters, such as the HSL synthesis rate per cell, and an algorithm was proposed in order to evaluate the O.D.start for every self-inducible device. | ||
| + | |||
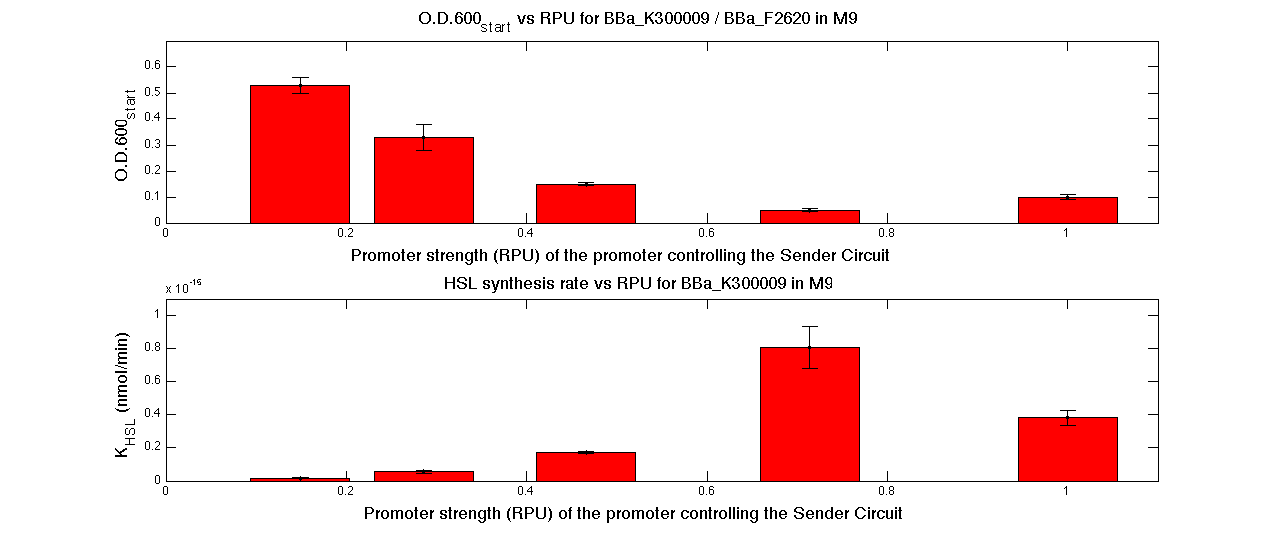
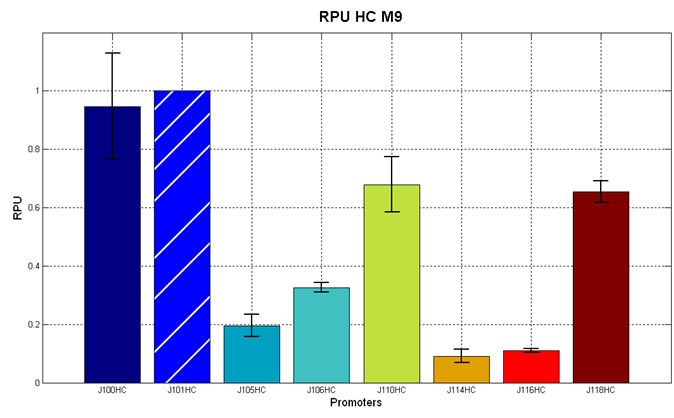
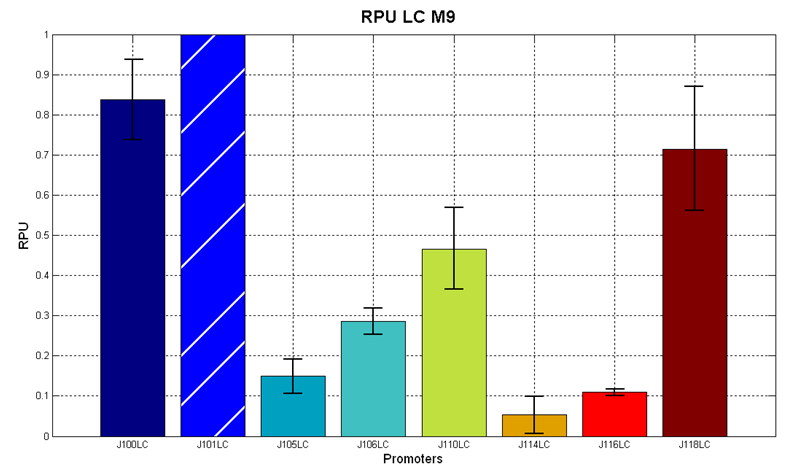
| + | In the figures below, the O.D.start and the HSL synthesis rate as a function of the strength of the promoter (RPU) controlling the signal molecule production are reported for <partinfo>BBa_K300010</partinfo> (Sender&Receiver device in HC plasmid) and for <partinfo>BBa_K300009</partinfo>/<partinfo>BBa_F2620</partinfo> (Sender device in LC plasmid in combination with Receiver device in HC plasmid). | ||
| + | |||
| + | {| | ||
| + | |[[Image:pv_RPUvsOD_HCHC_M9.png|700px|thumb|center| O.D.start and K_HSL as a function of RPUs of the promoters controlling the signal molecule production for <partinfo>BBa_K300010</partinfo> in high copy number plasmid in M9 medium]] | ||
| + | |- | ||
| + | |[[Image:pv_RPUvsOD_HCLC_M9.png|700px|thumb|center| O.D.start and K_HSL as a function of RPUs of the promoters controlling the autoinducer production for <partinfo>BBa_K300010</partinfo> in low copy number plasmid used in combination with <partinfo>BBa_F2620</partinfo> in high copy plasmid in M9 medium]] | ||
| + | |} | ||
| + | |||
| + | A strong correlation between the promoter strength, previously measured in RPU, and the O.D.start is depicted by the reported graphs. This is consistent with the expected behaviour of these parts, since the HSL synthesis rate is an increasing function of the upstream promoter's strength. | ||
| + | |||
| + | For RPU values greater or equal to 1, the expected behaviour is not confirmed anymore. This is probably because too high luxI expression levels and/or synthesis rate of HSL are injurious for the cell. | ||
| + | |||
| + | The combination of promoter strength variation and plasmid copy number modulation allows the creation of a library of self-inducible devices able to perform autoinduction at many different O.D.600 values, in any cellular growth phase. | ||
| + | |||
=<partinfo>BBa_F2620</partinfo> - 3OC6HSL -> PoPS Receiver= | =<partinfo>BBa_F2620</partinfo> - 3OC6HSL -> PoPS Receiver= | ||
| + | |||
| + | The measurement system of this part (<partinfo>BBa_T9002</partinfo>) was successfully used as a biosensor to estimate the concentration of HSL present in the growth media of cultures expressing ''luxI'' (<partinfo>BBa_C0061</partinfo>). | ||
| + | |||
| + | Additional information and details are available in [http://partsregistry.org/Part:BBa_K300009:Experience BBa_K300009 Experience page]. | ||
| + | |||
| + | |||
| + | {| align='center' | ||
| + | |[[Image:pv_HCT9002sensor.png|500px|thumb|center|<partinfo>BBa_T9002</partinfo> calibration curve for detection of [HSL] produced by autoinducer generators in high copy vector.]] | ||
| + | |} | ||
| + | |||
| + | |||
| + | {| align='center' | ||
| + | |[[Image:pv_LCT9002sensor.png|500px|thumb|center|<partinfo>BBa_T9002</partinfo> calibration curve for detection of [HSL] produced by the autoinducer generators in low copy vector.]] | ||
| + | |} | ||
| + | |||
=<partinfo>BBa_J61001</partinfo> - R6K Origin of replication= | =<partinfo>BBa_J61001</partinfo> - R6K Origin of replication= | ||
| + | |||
| + | <partinfo>BBa_K300008</partinfo> was used as a validation construct for this conditional replication origin, in order to test its capability to be propagated in pir+ or pir-116 strain and its inability to propagate in the other ''E. coli'' strains. | ||
| + | |||
| + | In particular, <partinfo>BBa_K300008</partinfo> was cut with XbaI-SpeI and the insert was isolated and purified from a 1% agarose gel. Then, it was self-ligated to generate a Cm-resistant R6K plasmid). | ||
| + | |||
| + | BW25141 (<partinfo>BBa_K300984</partinfo>) and BW23474 (<partinfo>BBa_K300985</partinfo>) were chosen as pir+ and pir-116 strains respectively, while DH5alpha (<partinfo>BBa_V1001</partinfo>), MC1061 (<partinfo>BBa_K300078</partinfo>) and MG1655 (<partinfo>BBa_V1000</partinfo>) were chosen as pir- strains. | ||
| + | |||
| + | |||
| + | All these strains were made competent following the commonly used CaCl2 method [Sambrook J, Fritsch EF, and Maniatis T (1989), Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.]. Then, a vial of 100 ul of competent cells was transformed with 2-4 ng of: | ||
| + | *no DNA (negative control); | ||
| + | *a pSB*** series vector (positive control); | ||
| + | *self-ligated <partinfo>BBa_K300008</partinfo>. | ||
| + | |||
| + | and plated on LB+Cm at 34 ug/ml for high-copy plasmids, Cm at 12.5 ug/ml for medium/low copy plasmids and for the negative control strains transformed with the R6K plasmids. | ||
| + | |||
| + | |||
| + | The colonies were counted in each plate and the transformation efficiency was estimated in '''[CFU/ug of DNA]''' as: | ||
| + | |||
| + | <div align=center> | ||
| + | ''efficiency [CFU/ug of DNA]= # CFU * 1000 ng of DNA / amount of transformed DNA [ng]'' | ||
| + | </div> | ||
| + | |||
| + | The results are shown here: | ||
| + | {|border=1 | ||
| + | |'''Strain''' | ||
| + | |'''Efficiency with no DNA''' | ||
| + | |'''Efficiency with pSB*** (positive control)''' | ||
| + | |'''Efficiency with the self-ligated <partinfo>BBa_K300008</partinfo> (R6K plasmid)''' | ||
| + | |- | ||
| + | |<partinfo>BBa_K300084</partinfo> | ||
| + | |0 | ||
| + | |10^5 | ||
| + | |10^5 | ||
| + | |- | ||
| + | |<partinfo>BBa_K300085</partinfo> | ||
| + | |0 | ||
| + | |10^6 | ||
| + | |10^6 | ||
| + | |- | ||
| + | |<partinfo>BBa_V1001</partinfo> | ||
| + | |0 | ||
| + | |10^8 | ||
| + | |0 | ||
| + | |- | ||
| + | |<partinfo>BBa_K300078</partinfo> | ||
| + | |0 | ||
| + | |10^6 | ||
| + | |0 | ||
| + | |- | ||
| + | |<partinfo>BBa_V1000</partinfo> | ||
| + | |0 | ||
| + | |10^5 | ||
| + | |0 | ||
| + | |} | ||
| + | |||
| + | These results show that <partinfo>BBa_J61001</partinfo> replication origin can be only propagated in pir+ and pir-116 strains (<partinfo>BBa_K300084</partinfo> and <partinfo>BBa_K300085</partinfo>), while the transformation of other strains with the R6K plasmid yielded no colonies after transformation. | ||
| + | |||
| + | Moreover, these results show that the R6K plasmid in pir+ and pir-116 strains was transformed with the same efficiency as the pSB*** positive control plasmid, demonstrating that the R6K origin doesn't give any handicap in plasmid transformation. | ||
=<partinfo>BBa_J23100</partinfo>, <partinfo>BBa_J23101</partinfo>, <partinfo>BBa_J23105</partinfo>, <partinfo>BBa_J23106</partinfo>, <partinfo>BBa_J23110</partinfo>, <partinfo>BBa_J23114</partinfo>, <partinfo>BBa_J23116</partinfo>, <partinfo>BBa_J23118</partinfo> - constitutive promoters from Anderson's collection= | =<partinfo>BBa_J23100</partinfo>, <partinfo>BBa_J23101</partinfo>, <partinfo>BBa_J23105</partinfo>, <partinfo>BBa_J23106</partinfo>, <partinfo>BBa_J23110</partinfo>, <partinfo>BBa_J23114</partinfo>, <partinfo>BBa_J23116</partinfo>, <partinfo>BBa_J23118</partinfo> - constitutive promoters from Anderson's collection= | ||
| Line 232: | Line 611: | ||
=<partinfo>BBa_P1004</partinfo> - chloramphenicol resistance cassette= | =<partinfo>BBa_P1004</partinfo> - chloramphenicol resistance cassette= | ||
| + | |||
| + | <partinfo>BBa_P1004</partinfo> has been successfully used in the assembly of <partinfo>BBa_K300000</partinfo> integrative base vector for ''E. coli''. All the intermediate parts which contained <partinfo>BBa_P1004</partinfo> showed Chloramphenicol resistance (tested up to 34 ug/ml in LB media) when transformed in TOP10 (<partinfo>BBa_V1009</partinfo>), DH5alpha (<partinfo>BBa_V1001</partinfo>), MC1061 (<partinfo>BBa_K300078</partinfo>), MG1655 (<partinfo>BBa_V1000</partinfo>), BW23474 (<partinfo>BBa_K300985</partinfo>) and DB3.1 (<partinfo>BBa_V1005</partinfo>) strains in a high-copy plasmid. | ||
| + | |||
=<partinfo>BBa_K125500</partinfo> - GFP fusion brick= | =<partinfo>BBa_K125500</partinfo> - GFP fusion brick= | ||
| Line 273: | Line 655: | ||
=<partinfo>BBa_J72008</partinfo> - phi80 integration helper plasmid pInt80-649= | =<partinfo>BBa_J72008</partinfo> - phi80 integration helper plasmid pInt80-649= | ||
| + | |||
| + | <partinfo>BBa_J72008</partinfo> has been successfully used in the integration protocol of both MC1061 (<partinfo>BBa_K300078</partinfo>) and MG1655 (<partinfo>BBa_V1000</partinfo>) ''E. coli'' strains. See [http://partsregistry.org/Part:BBa_K300000:Experience BBa_K300000 Experience page] for details about how this plasmid was used. | ||
| + | |||
| + | It can be actually cured at 37-42°C, while it can be propagated at 30°C. | ||
| + | |||
| + | It actually enables the propagation of R6K (conditional replication origin) plasmids, thanks to its pir-116 gene. | ||
<tr><td><br> | <tr><td><br> | ||
Latest revision as of 03:42, 28 October 2010
|
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
 "
"