Team:Toronto/Results
From 2010.igem.org
| (31 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
| - | + | [[Image:Toronto_logo.png]] | |
| - | + | {|style="color:black;background-color:#FFFFFF;" cellpadding="2" cellspacing="0" border="1" width="100%" align="center" | |
| - | {| style="color: | + | |
!align="center"|[[Team:Toronto|Home]] | !align="center"|[[Team:Toronto|Home]] | ||
| - | !align="center"|[[Team:Toronto/Project|Project | + | !align="center"|[[Team:Toronto/Project|Project]] |
!align="center"|[[Team:Toronto/Design|Design]] | !align="center"|[[Team:Toronto/Design|Design]] | ||
!align="center"|[[Team:Toronto/Protocols|Protocols]] | !align="center"|[[Team:Toronto/Protocols|Protocols]] | ||
| Line 15: | Line 14: | ||
!align="center"|[[Team:Toronto/Team|Team]] | !align="center"|[[Team:Toronto/Team|Team]] | ||
!align="center"|[https://igem.org/Team.cgi?year=2010&team_name=Toronto Official Team Profile] | !align="center"|[https://igem.org/Team.cgi?year=2010&team_name=Toronto Official Team Profile] | ||
| - | |||
| - | |||
| - | |||
| - | |||
|} | |} | ||
<h1>'''Results'''</h1> | <h1>'''Results'''</h1> | ||
| + | |||
| + | <h2>'''''E. coli'' DH5a is sensitive to Catechol exposure'''</h2> | ||
| + | |||
| + | To measure the effect of our pathway manipulations on the viability of ''E.coli'' exposed to catechol we plan to perform a tolerance test. In preliminary experiments we established the baseline sensitivity of ''E.coli'' (DH5a) exposed to 50mM catechol in liquid media over a six hour time course. Aliquots of this mixture were plated on LB media at various dilutions and colonies were counted and compared with untreated controls (see "Protocols"). There was no measurable difference between control and treated cells when bacteria were exposed to catechol in a growth media (data not shown). However, in minimal media (MM2) growth of control cells was moderate while exposed cells exhibited a marked decrease in survival (Figure 1). Although the effects of Catechol exposure in ''E.coli'' and ''P. putida'' have previously been investigated [Park et al. 2001] completion of these baseline experiments will enable us to determine the effect of pathway manipulations on the proposed engineered ''E.coli'' strains containing various configurations of the pathway. | ||
| + | |||
| + | <p> | ||
| + | '''A''' | ||
| + | [[Image:Toronto-LB-Plate_1.jpg|400px]] [[Image:Toronto-LB-Plate_2.jpg|400px]] | ||
| + | </p> | ||
| + | <p> | ||
| + | '''B'''[[Image:Toronto-results-catechol_tolerance.png|center]] | ||
| + | </p> | ||
| + | <blockquote>'''Figure 1:''' A. Growth of ''E.coli'' DH5a on LB plates (1x10<sup>-5</sup> dilution) following exposure to minimal media and minimal media with 50mM catechol in solution. Plates shown are for the last time points plotted. B. Survival curves of ''E. coli'' DH5a exposed to catechol over a six hour time course.</blockquote> | ||
| + | |||
| + | <H2>'''Synthetic construction of a functioning catechol ortho degradation pathway in E.coli is predicted to increase cell growth rate'''</H2> | ||
| + | To predict the effects of adding the catechol ortho-cleavage pathway on ''E.coli'' metabolism we used a previously developed genome-scale reconstruction of E. coli (iJR904) and applied flux balance analysis (FBA) to compare likely growth outcomes for normal and optimized pathways. We found that optimization of this pathway in the presence of glucose and sufficient oxygen leads to a prediction of increased growth rate. This result bodes well for the success of engineered bacteria introduced into mixed culture within the tailings ponds as they might be expected to have a survival advantage over their native counterparts. However, it should be pointed out that a corresponding genome-scale reconstruction of the target organism ''P. putida'' is not available at this time. These methods and results are discussed at length in the Modeling section. | ||
| + | |||
| + | [[image:fba.png|500px|center]] | ||
| + | Figure 1. Predicted growth of E. coli at various uptake rates of glucose and catechol. The points specified on the surface are considered further in simulations. Points are named according to the uptake rate of glucose and catechol (eg. Glc6/cat0 corresponds to a glucose uptake of 6 mmol/gDW/h and catechol uptake of zero). Glc6/cat0 corresponds to a “wildtype” (WT) condition, while glc6/cat2.52 corresponds to the “optimal” (OPT) ratio predicted for by FBA to maximize biomass production. Glc0/cat6 and glc6/cat20 represent additional hypothetical situtions where catechol replaces glucose as an energy source, and where both are used at their maximum rates, respectively. | ||
| + | |||
| + | |||
| + | <h2>'''Colocalization of muconolactone D-isomerase (EC:5.3.3.4) and 3-oxoadipate enol-lactone hydrolase (EC:3.1.1.24) are predicted to lead to increased catechol degradation through metabolic channeling'''</h2> | ||
| + | In order to determine which enzymes would most benefit from metabolic channeling in the catechol degradation pathway, we used a cellular simulation tool, developed in our lab, called Cell++. It allows us to place enzymes of choice in a compartment within a cellular environment and calculates the effect of localization on metabolite concentrations in a user-defined biochemical pathway (Sanford et al. 2006). We gathered kinetic data of the five enzymes in the degradation pathway and investigated the effects of localizing pairs of sequential enzymes (i.e. catechol 1, 2-dioxygenase and muconate cycloisomeras) on metabolite concentrations. Four co-localization simulations were performed in Cell++ along with a control simulation where no enzymes were co-localized. The initial metabolite in the simulation environment, catechol, was added into the simulation environment and the simulation environment was run for 10,000 iterations to observe the effect on catechol degradation in each simulation. The results are displayed as a table indicating the concentration of all metabolites at 1000 equidistant time points during the simulation. | ||
| + | <p>To make use of this information, we extracted the maximum metabolite concentration and the maximum rate of metabolite formation for all metabolites during each simulation. These values are normalized from a scale of 0 to 1, where 0 indicates the lowest value of the concentration/rate of formation (C/RoF) in the simulations and 1 is the highest. These values were further evaluated against the control simulation to the find the percentage change of the C/RoF values of each metabolite in the four co-localization simulations. In order to classify a particular enzyme co-localization as beneficial, we need to see an increase in the maximum concentration and rate of formation of the final metabolite in the pathway (meta6). These increases would represent an increased flux through the pathway. Of the four co-localization simulations, simulation 3 (which co-localizes the enzymes muconolactone D-isomerase and 3-oxodipate-enol lactone hydrolase) was the only simulation that matched the criteria for a beneficial co-localization. From these simulations, we decided to co-localize these two enzymes for our wet lab experiments to see if Cell++ accurately depicted the effects on catechol degradation.</p> | ||
| + | |||
| + | ''' '''[[Image:Cellppheatmap.jpg|center]] | ||
| + | |||
| + | <blockquote>'''Figure 2.''' Simulating the effects of enzyme co-localization on catechol degradation efficiency. | ||
| + | Red indicates increase in the value of the kinetic parameter relative to the simulation where no enzymes are co-localized (0.0), while green indicates a decrease in the kinetic parameter. 1.0, 2.0, 3.0 and 4.0 represent pair-wise fusions of the 1st and 2nd, 2nd and 3rd, 3rd and 4th, and 4th and 5th enzymes respectively. “Max[]” and “Vmax” represent the maximum concentration and the maximum rate of formation of a particular metabolite in a particular simulation. The fusion of the 3rd and 4th enzymes (3.0) resulted in an increase of the production of the final pathway product, while the other fusion pairs demonstrate no increase or even reductions in the production of the final metabolite in the simulated pathway.</blockquote> | ||
| + | |||
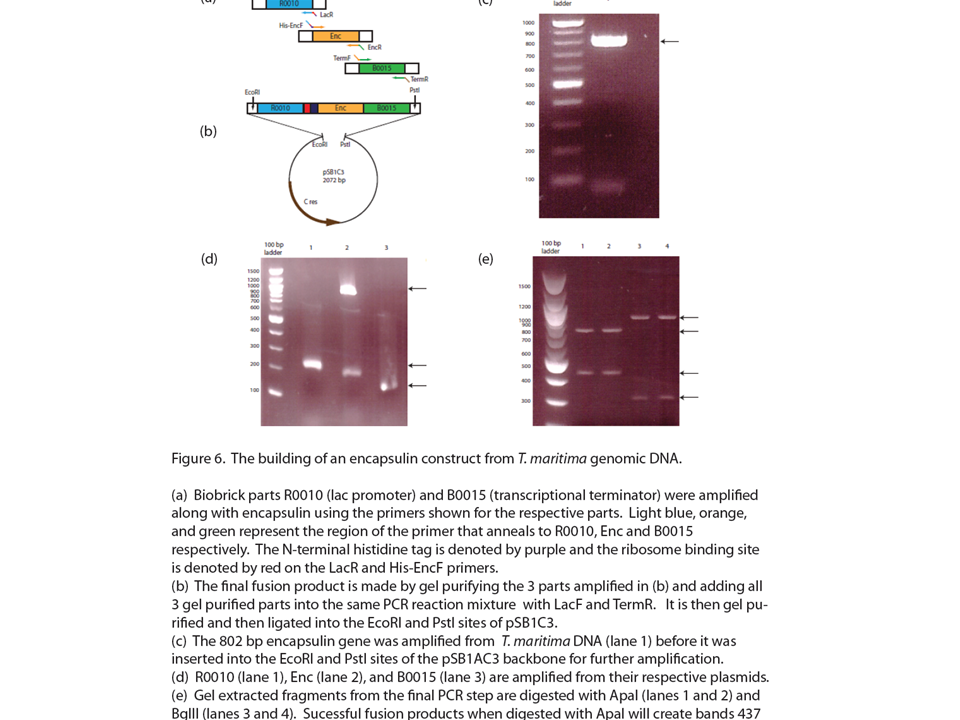
| + | <h2>'''Encapsulin derived from ''T. maritima'' can be stably expressed in ''E.coli'''''</h2> | ||
| + | As part of our 2009 igem project we described a new bacterial microcompartment based on encapsulin which we intend to use for metabolic channeling. We obtained cloned dna from ''T. maritima'' and created a biobrick corresponding to this unique protein. Here we have shown that this part can be stably expressed in ''E.coli''. | ||
| + | |||
| + | [[image:Toronto_EncapsulinConstruction.png]] | ||
| + | [[image:Toronto_EncapsulinExpression.png]] | ||
Latest revision as of 08:22, 27 October 2010
| Home | Project | Design | Protocols | Notebook | Results | Parts Submitted to the Registry | Modeling | Software Used | Human Practices | Safety | Team | Official Team Profile |
|---|
Results
E. coli DH5a is sensitive to Catechol exposure
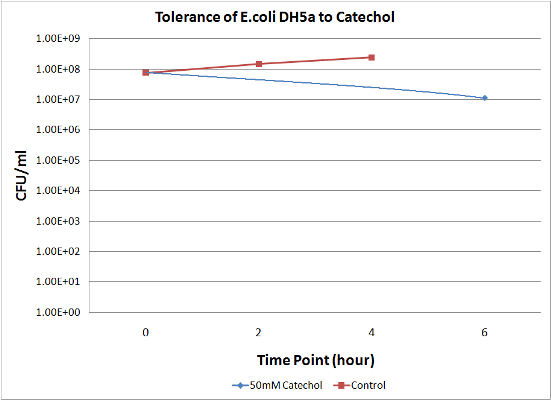
To measure the effect of our pathway manipulations on the viability of E.coli exposed to catechol we plan to perform a tolerance test. In preliminary experiments we established the baseline sensitivity of E.coli (DH5a) exposed to 50mM catechol in liquid media over a six hour time course. Aliquots of this mixture were plated on LB media at various dilutions and colonies were counted and compared with untreated controls (see "Protocols"). There was no measurable difference between control and treated cells when bacteria were exposed to catechol in a growth media (data not shown). However, in minimal media (MM2) growth of control cells was moderate while exposed cells exhibited a marked decrease in survival (Figure 1). Although the effects of Catechol exposure in E.coli and P. putida have previously been investigated [Park et al. 2001] completion of these baseline experiments will enable us to determine the effect of pathway manipulations on the proposed engineered E.coli strains containing various configurations of the pathway.
B
Figure 1: A. Growth of E.coli DH5a on LB plates (1x10-5 dilution) following exposure to minimal media and minimal media with 50mM catechol in solution. Plates shown are for the last time points plotted. B. Survival curves of E. coli DH5a exposed to catechol over a six hour time course.
Synthetic construction of a functioning catechol ortho degradation pathway in E.coli is predicted to increase cell growth rate
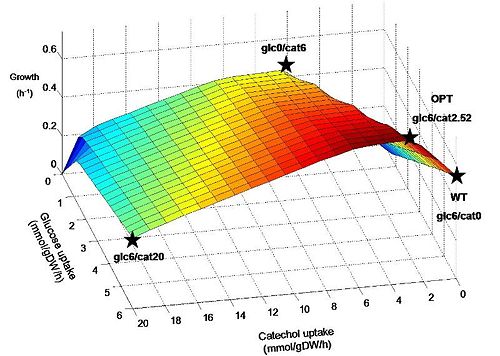
To predict the effects of adding the catechol ortho-cleavage pathway on E.coli metabolism we used a previously developed genome-scale reconstruction of E. coli (iJR904) and applied flux balance analysis (FBA) to compare likely growth outcomes for normal and optimized pathways. We found that optimization of this pathway in the presence of glucose and sufficient oxygen leads to a prediction of increased growth rate. This result bodes well for the success of engineered bacteria introduced into mixed culture within the tailings ponds as they might be expected to have a survival advantage over their native counterparts. However, it should be pointed out that a corresponding genome-scale reconstruction of the target organism P. putida is not available at this time. These methods and results are discussed at length in the Modeling section.
Figure 1. Predicted growth of E. coli at various uptake rates of glucose and catechol. The points specified on the surface are considered further in simulations. Points are named according to the uptake rate of glucose and catechol (eg. Glc6/cat0 corresponds to a glucose uptake of 6 mmol/gDW/h and catechol uptake of zero). Glc6/cat0 corresponds to a “wildtype” (WT) condition, while glc6/cat2.52 corresponds to the “optimal” (OPT) ratio predicted for by FBA to maximize biomass production. Glc0/cat6 and glc6/cat20 represent additional hypothetical situtions where catechol replaces glucose as an energy source, and where both are used at their maximum rates, respectively.
Colocalization of muconolactone D-isomerase (EC:5.3.3.4) and 3-oxoadipate enol-lactone hydrolase (EC:3.1.1.24) are predicted to lead to increased catechol degradation through metabolic channeling
In order to determine which enzymes would most benefit from metabolic channeling in the catechol degradation pathway, we used a cellular simulation tool, developed in our lab, called Cell++. It allows us to place enzymes of choice in a compartment within a cellular environment and calculates the effect of localization on metabolite concentrations in a user-defined biochemical pathway (Sanford et al. 2006). We gathered kinetic data of the five enzymes in the degradation pathway and investigated the effects of localizing pairs of sequential enzymes (i.e. catechol 1, 2-dioxygenase and muconate cycloisomeras) on metabolite concentrations. Four co-localization simulations were performed in Cell++ along with a control simulation where no enzymes were co-localized. The initial metabolite in the simulation environment, catechol, was added into the simulation environment and the simulation environment was run for 10,000 iterations to observe the effect on catechol degradation in each simulation. The results are displayed as a table indicating the concentration of all metabolites at 1000 equidistant time points during the simulation.
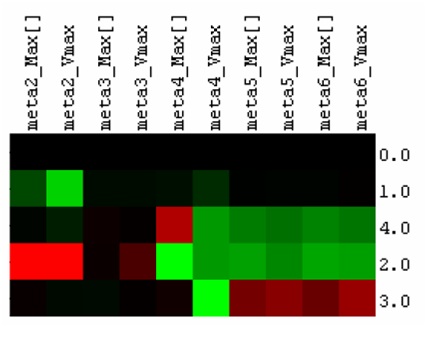
To make use of this information, we extracted the maximum metabolite concentration and the maximum rate of metabolite formation for all metabolites during each simulation. These values are normalized from a scale of 0 to 1, where 0 indicates the lowest value of the concentration/rate of formation (C/RoF) in the simulations and 1 is the highest. These values were further evaluated against the control simulation to the find the percentage change of the C/RoF values of each metabolite in the four co-localization simulations. In order to classify a particular enzyme co-localization as beneficial, we need to see an increase in the maximum concentration and rate of formation of the final metabolite in the pathway (meta6). These increases would represent an increased flux through the pathway. Of the four co-localization simulations, simulation 3 (which co-localizes the enzymes muconolactone D-isomerase and 3-oxodipate-enol lactone hydrolase) was the only simulation that matched the criteria for a beneficial co-localization. From these simulations, we decided to co-localize these two enzymes for our wet lab experiments to see if Cell++ accurately depicted the effects on catechol degradation.
Figure 2. Simulating the effects of enzyme co-localization on catechol degradation efficiency. Red indicates increase in the value of the kinetic parameter relative to the simulation where no enzymes are co-localized (0.0), while green indicates a decrease in the kinetic parameter. 1.0, 2.0, 3.0 and 4.0 represent pair-wise fusions of the 1st and 2nd, 2nd and 3rd, 3rd and 4th, and 4th and 5th enzymes respectively. “Max[]” and “Vmax” represent the maximum concentration and the maximum rate of formation of a particular metabolite in a particular simulation. The fusion of the 3rd and 4th enzymes (3.0) resulted in an increase of the production of the final pathway product, while the other fusion pairs demonstrate no increase or even reductions in the production of the final metabolite in the simulated pathway.
Encapsulin derived from T. maritima can be stably expressed in E.coli
As part of our 2009 igem project we described a new bacterial microcompartment based on encapsulin which we intend to use for metabolic channeling. We obtained cloned dna from T. maritima and created a biobrick corresponding to this unique protein. Here we have shown that this part can be stably expressed in E.coli.
 "
"