Team:ETHZ Basel/Achievements
From 2010.igem.org
(Difference between revisions)
(→BioBrick Toolbox) |
|||
| (97 intermediate revisions not shown) | |||
| Line 3: | Line 3: | ||
= Achievements Overview = | = Achievements Overview = | ||
| - | <html><div class="thumb tright"><div class="thumbinner" style="width: | + | <html> |
| - | <iframe title="YouTube video player" class="youtube-player" type="text/html" width=" | + | <div class="thumb tright"><div class="thumbinner" style="width:402px;"> |
| - | <div class="thumbcaption"><div class="magnify"><a href="http://www.youtube.com/watch?v= | + | <iframe title="YouTube video player" class="youtube-player" type="text/html" width="400" height="325" src="http://www.youtube.com/embed/mulRvAVExSc?rel=0&hd=1" frameborder="0"></iframe> |
| - | + | <div class="thumbcaption"><div class="magnify"><a href="http://www.youtube.com/watch?v=mulRvAVExSc&hd=1" class="external" title="Enlarge"><img src="/wiki/skins/common/images/magnify-clip.png" width="15" height="11" alt="" | |
| - | + | /></a></div><b>The E. lemming in action.</b> | |
| + | <br><i>Blue dots</i>: the detected <i>E. coli</i> cells. <i>Yellow dot</i>: the currently selected E. lemming. <i>Yellow cone</i>: the current swimming direction of E. lemming. <i>Blue background</i>: blue light on (inducing directed movement). <i>Gray environment</i>: blue light off (inducing tumbling).</br> | ||
| + | <br> Note how the E. lemming is keeping its direction under the influence of blue light, whereas it is tumbling and quickly changing directions when the blue light is off. </br> | ||
| - | = | + | <br />The unprocessed microscope images are available <a href="https://2010.igem.org/Team:ETHZ_Basel/Achievements/OriginalImages">here</a>.</div></div></div> |
| - | + | </html> | |
| - | + | * '''The E. lemming: '''Watch the '''[https://2010.igem.org/Team:ETHZ_Basel/Achievements/E_lemming smallest remotely controlled robot in action]''' at ETH Zurich! The implementation using the chimeric fusion of the archeal light receptor ([http://partsregistry.org/Part:BBa_K422001 BBa_K422001]) to the bacterial chemotactic transducer enabled us to generate the first E. lemming. You find here recorded videos of live images of the light controlled ''E. coli'' - [https://2010.igem.org/Team:ETHZ_Basel/Achievements/E_lemming '''The E. lemming'''] is alive!<p style="font-size:2px"><br><br></p> | |
| - | + | * '''BioBrick Toolbox: '''Get a glimpse of our collection of [https://2010.igem.org/Team:ETHZ_Basel/Achievements/BioBrick_Toolbox '''BioBricks'''] that shows the proteins we made not only for E. lemming's chemotaxis signaling network and cellular anchoring, but also for its light sensing & fluorescence reporting pathway, including our favorites: the light-sensitive couple PhyB-Pif3 and the Archeal light receptor: an intra-species fusion protein! You also find here the characterization of our parts.<p style="font-size:2px"><br><br></p> | |
| - | + | * '''MATLAB Toolbox (Lemming Toolbox):''' [[Image:lemmingToolbox.jpg|thumb|350px|'''MATLAB Toolbox''' Screenshot of the Simulink blocks available in the toolbox. Please note that these blocks can be visually assembled with the Simulink blocks of other Toolboxes (e.g. blocks for nearly any mathematical function), and that the algorithms we developed can be accessed directly through Matlab code, too.]]We managed to develop all our computer based tools & algorithms as re-usable, publicly available modules and we assembled the entire ''in silico'' setup of the E. Lemming into a [https://2010.igem.org/Team:ETHZ_Basel/Achievements/Matlab_Toolbox '''MATLAB Toolbox''']! Our Toolbox includes 18 Simulink blocks encapsulating the algorithms we developed/implemented. We want to share our work with the iGEM community and interested persons, this is why we made it freely available for [http://sourceforge.net/projects/ethzigem10/files/LemmingToolbox_Setup.zip/download downloading].<p style="font-size:2px"><br><br></p> | |
| - | + | * '''New Technical Standard:''' After weeks of applying the cloning strategy BBF RFC28, we came up with [https://2010.igem.org/Team:ETHZ_Basel/Achievements/Technical_Standard '''our own strategy'''] on how BFF RFC28 could be made compatible with Tom Knight`s original assembly standard.<p style="font-size:2px"><br><br></p> | |
| - | + | * '''Model of light-inducible chemotaxis pathway:''' We worked out a [https://2010.igem.org/Team:ETHZ_Basel/Modeling/Combined '''model to control chemotaxis by light''']. The idea was to instrumentalize light sensitive proteins (LSPs) in order to manipulate the concentrations of the chemotaxis proteins (e.g. CheY), through an intracellular anchoring reaction. As a consequence, we can take control over this pathway and direct the E.lemming's movement!<p style="font-size:2px"><br><br></p> | |
| - | + | * '''Movement Model:''' We made a [https://2010.igem.org/Team:ETHZ_Basel/Modeling/Movement '''stochastic model to realistically simulate the chemotaxis motion'''] of the cell ''in silico''. The bacterium responds to the light input, which is converted into changes in the concentration of the chemotaxis proteins, which further influence the motion of the E. lemming.<p style="font-size:2px"><br><br></p> | |
| - | + | * '''E. lemming 2D - The Game:''' We invented a [https://2010.igem.org/Team:ETHZ_Basel/InformationProcessing/Game '''synthetic biology shooting game'''] featuring our very own E. lemming! Give it a try and play with our E. lemming!<p style="font-size:2px"><br><br></p> | |
| - | * [https://2010.igem.org/Team:ETHZ_Basel/ | + | * '''Controller: ''' We developed [https://2010.igem.org/Team:ETHZ_Basel/InformationProcessing/Controller '''five novel controllers'''] for directing the E. lemming to the desired target. As stimuli for altering the bacterial movement, we use light inputs.<p style="font-size:2px"><br><br></p> |
| - | * | + | * '''Systems Design:''' See how we managed to use our models to support the wetlab and vice versa when we [https://2010.igem.org/Team:ETHZ_Basel/Achievements/Systems_Design '''designed the system''']. |
| - | * | + | |
| - | * | + | |
| - | + | ||
| - | + | ||
| - | * | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
Latest revision as of 00:00, 28 October 2010
Achievements Overview
The E. lemming in action.
Blue dots: the detected E. coli cells. Yellow dot: the currently selected E. lemming. Yellow cone: the current swimming direction of E. lemming. Blue background: blue light on (inducing directed movement). Gray environment: blue light off (inducing tumbling).
Note how the E. lemming is keeping its direction under the influence of blue light, whereas it is tumbling and quickly changing directions when the blue light is off.
The unprocessed microscope images are available here.
Blue dots: the detected E. coli cells. Yellow dot: the currently selected E. lemming. Yellow cone: the current swimming direction of E. lemming. Blue background: blue light on (inducing directed movement). Gray environment: blue light off (inducing tumbling).
Note how the E. lemming is keeping its direction under the influence of blue light, whereas it is tumbling and quickly changing directions when the blue light is off.
The unprocessed microscope images are available here.
- The E. lemming: Watch the smallest remotely controlled robot in action at ETH Zurich! The implementation using the chimeric fusion of the archeal light receptor ([http://partsregistry.org/Part:BBa_K422001 BBa_K422001]) to the bacterial chemotactic transducer enabled us to generate the first E. lemming. You find here recorded videos of live images of the light controlled E. coli - The E. lemming is alive!
- BioBrick Toolbox: Get a glimpse of our collection of BioBricks that shows the proteins we made not only for E. lemming's chemotaxis signaling network and cellular anchoring, but also for its light sensing & fluorescence reporting pathway, including our favorites: the light-sensitive couple PhyB-Pif3 and the Archeal light receptor: an intra-species fusion protein! You also find here the characterization of our parts.
- MATLAB Toolbox (Lemming Toolbox): We managed to develop all our computer based tools & algorithms as re-usable, publicly available modules and we assembled the entire in silico setup of the E. Lemming into a MATLAB Toolbox! Our Toolbox includes 18 Simulink blocks encapsulating the algorithms we developed/implemented. We want to share our work with the iGEM community and interested persons, this is why we made it freely available for [http://sourceforge.net/projects/ethzigem10/files/LemmingToolbox_Setup.zip/download downloading].
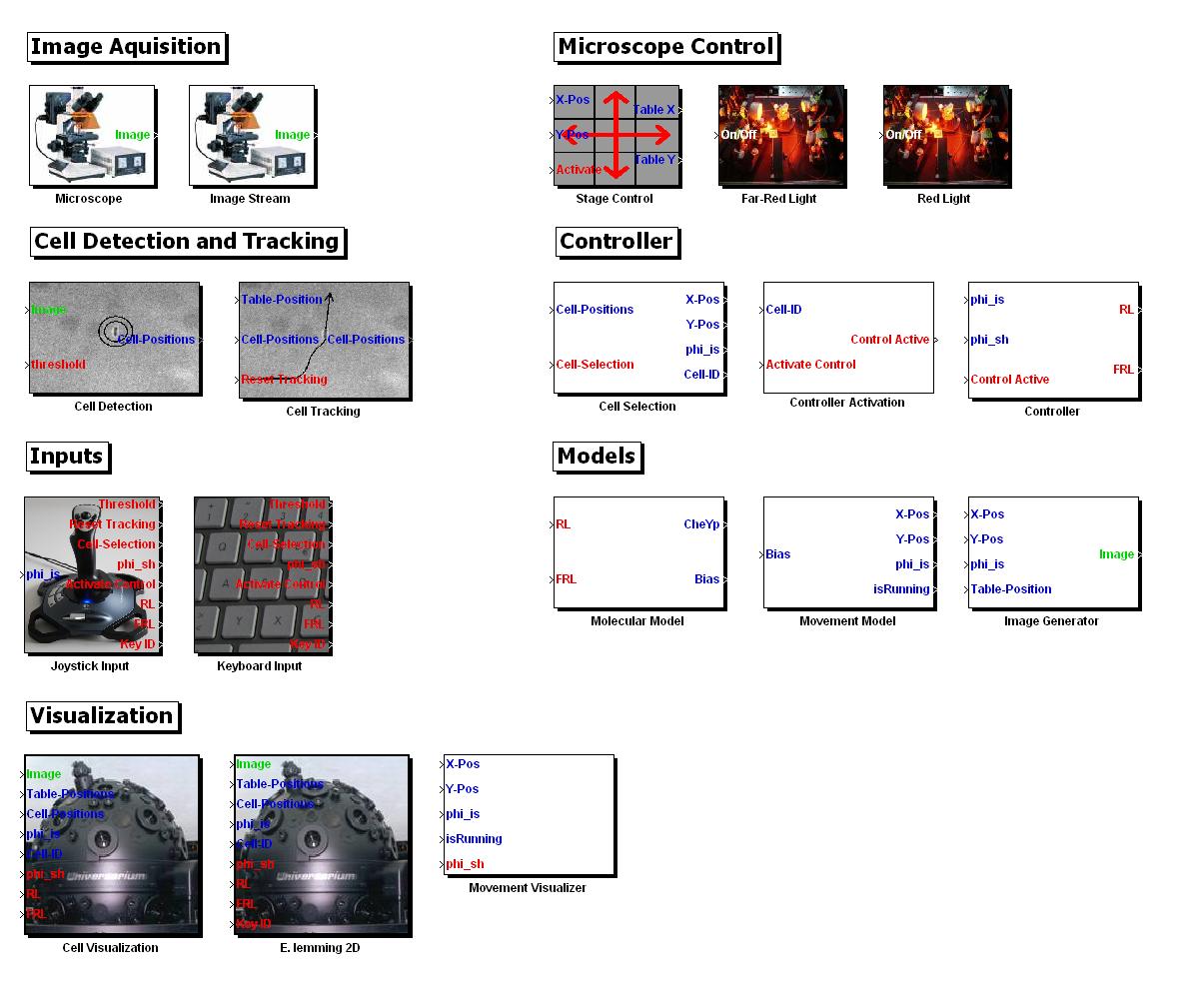 MATLAB Toolbox Screenshot of the Simulink blocks available in the toolbox. Please note that these blocks can be visually assembled with the Simulink blocks of other Toolboxes (e.g. blocks for nearly any mathematical function), and that the algorithms we developed can be accessed directly through Matlab code, too.
MATLAB Toolbox Screenshot of the Simulink blocks available in the toolbox. Please note that these blocks can be visually assembled with the Simulink blocks of other Toolboxes (e.g. blocks for nearly any mathematical function), and that the algorithms we developed can be accessed directly through Matlab code, too. - New Technical Standard: After weeks of applying the cloning strategy BBF RFC28, we came up with our own strategy on how BFF RFC28 could be made compatible with Tom Knight`s original assembly standard.
- Model of light-inducible chemotaxis pathway: We worked out a model to control chemotaxis by light. The idea was to instrumentalize light sensitive proteins (LSPs) in order to manipulate the concentrations of the chemotaxis proteins (e.g. CheY), through an intracellular anchoring reaction. As a consequence, we can take control over this pathway and direct the E.lemming's movement!
- Movement Model: We made a stochastic model to realistically simulate the chemotaxis motion of the cell in silico. The bacterium responds to the light input, which is converted into changes in the concentration of the chemotaxis proteins, which further influence the motion of the E. lemming.
- E. lemming 2D - The Game: We invented a synthetic biology shooting game featuring our very own E. lemming! Give it a try and play with our E. lemming!
- Controller: We developed five novel controllers for directing the E. lemming to the desired target. As stimuli for altering the bacterial movement, we use light inputs.
- Systems Design: See how we managed to use our models to support the wetlab and vice versa when we designed the system.
 "
"


