Team:Kyoto/Project
From 2010.igem.org
(→1. Solution to the Restriction on Promoter's Activity) |
|||
| (168 intermediate revisions not shown) | |||
| Line 1: | Line 1: | ||
{{:Team:Kyoto/Header}} | {{:Team:Kyoto/Header}} | ||
==Project== | ==Project== | ||
| + | [[Image:KyotoTop.png|600px|center]] | ||
| + | ===Abstract=== | ||
| + | <span class="title">'''The Fantastic Lysisbox:'''</span> | ||
| + | Genetically engineered cell death is essential for the application of biotechnology, such as in bioremediation area. In order to control the cell-death, we designed “Lysisbox”, which consists of a pair of modules: “Killer gene” and “Anti-killer gene.” As the Killer gene for ''Escherichia coli'', we noted the lysis cassette [SRRz/Rz1 gene] of lambda phage coding for holin and endolysin. The holin form pores in the inner membrane and the endolysin access and degrade the peptidoglycan passing through the pores, leading the E.coli to death. As the Anti-killer gene, we chose S<sub>ΔTMD1</sub> coding for a dominant-negative holin that inhibits the formation of the fatal pores. The balance of these two genes' expression level has the key to the E.coli’s life or death. In addition, such controllable membrane pores should show critical functions for all living organisms with lipid membranes and a cell wall. “Lysisbox” will contribute a lot to all the future iGEM projects, thus you must say “FANTASTIC!!!" | ||
===Introduction=== | ===Introduction=== | ||
====Background==== | ====Background==== | ||
| - | The cell-death device is required in many fields. In bioremediation, for example, this device can prevent introduced bacteria | + | The cell-death device is required in many fields <sup>[[#RefH001|[1]]] [[#RefH002|[2]]]</sup>. In bioremediation, for example, this device can prevent ecosystem disruption by introduced bacteria <sup>[[#RefH003|[3]]] [[#RefH004|[4]]]</sup>. Also, it can be applied to other ideas such as E.coli capsules which scatter drug or aroma by cell-lysis responding to a certain signal. Therefore, the cell-death devices based on diverse approaches, including previous iGEM projects have been developed <sup>[[#RefH001|[1]]] [[#RefH003|[3]]] [[#RefH005|[5]]] [[#RefH006|[6]]]</sup>. However, present cell-death devices are not useful because they have two problems as follows: |
| + | |||
| + | #'''They are not quantitatively characterized'''<sup>[[#RefH007|[7]]]</sup> | ||
| + | #'''They are not applied to many promoters'''<sup>[[#RefH008|[8]]]</sup>. | ||
| + | |||
| + | Therefore, we attempted to solve these problems and design new BioBricks that are better characterized and more applicable cell-death device to contribute to every future projects. | ||
| + | [[Image:kyoto-intro.png|600px|center]] | ||
====Problems of Present Cell-Death Devices==== | ====Problems of Present Cell-Death Devices==== | ||
| - | |||
| - | =====1. | + | =====1.Need for Quantitative Characterization===== |
| - | In | + | In previous iGEM, many teams focused on λ lysis cassette, which is a killer-gene derived from λ phage and causes cell lysis. However, they have only characterized its lytic activity qualititatively. Quantitative characterization of its lytic activity is necessary in order to design a universal and applicable cell-death device. For example, we wish to analyze when cell lysis occurs after induction of λ lysis cassette, how many cells lyse, whether all of cells lyse or not, and so on. |
| - | |||
| - | =====2. | + | =====2. Solution to the Restriction on Application of Promoters===== |
| - | In | + | In constructing a simple circuit consisting of an inducible promoter and λ lysis cassette, the combination are limited by the promoter's basal expression, strength of killer-function and many other factors. |
| + | We looked for the solution to this problem and came up with an idea; using both the lysis cassette and its antikiller-gene, S<sub>ΔTMD1</sub>, which lacks the code for trans membrane domain 1 of killer-gene. We can use two promoters (a constitutive one and an inducible one) for regulating cell-death balance. Into this circuit, we can apply more promoters to this device by using an appropriate constitutive promoter. | ||
| - | [[# | + | We considered that lysis cassette and S<sub>ΔTMD1</sub> are the most appropriate to killer-gene and anti-killer gene <sup>[[#RefH005|[5]]]</sup>. This was because we found some articles which show the effect of S<sub>ΔTMD1</sub> in the yeast <sup>[[#RefH009|[9]]]</sup> and also because previous iGEM teams already dealt with lysis cassette. |
| - | + | For further description about lysis cassette and S<sub>ΔTMD1</sub>, go [[Team:Kyoto/LearnMore#Lysis Cassette|[learn more]]] | |
| - | ====[[Team:Kyoto/Project/Goal A|Goal A | + | ===Our Goals=== |
| - | [[Image: | + | ====[[Team:Kyoto/Project/Goal A|Goal A: Characterization of R0011, a strong lactose promoter]]==== |
| - | We used R0011, which is one of the most commonly used parts in iGEM, to characterize our genes because the strength of promoter activity is very high. However, in previous iGEM, no teams characterized R0011 with RPU. We tried to determine | + | [[Image:KyotoGoalA.png|450px|left]] |
| + | We used R0011, which is one of the most commonly used parts in iGEM, to characterize our genes because the strength of promoter activity is very high. However, in previous iGEM, no teams characterized R0011 with RPU. We tried to determine how the activity of R0011<sup>[[#RefH010|[10]]]</sup> changes. | ||
| + | [[Image:KyotoGrpA003.png|450px|left]] | ||
| - | = | + | <div style="float:right;">[[Team:Kyoto/Project/Goal A|<html><img src="https://static.igem.org/mediawiki/2010/thumb/1/12/KyotoArrowA.png/200px-KyotoArrowA.png"/></html>]]</div> |
| - | < | + | {{clear}} |
| - | + | ||
| - | + | ||
| - | </div> | + | |
| - | + | ====[[Team:Kyoto/Project/Goal B|Goal B: Quantitative characterization of λ lysis cassette]]==== | |
| - | + | [[Image:KyotoGoalB.png|450px|left]] | |
| + | [[Image:KyotoFigC004.png|450px]]<br/> | ||
| + | We found a new fact that cell lysis caused by λ lysis cassette become an apparent equilibrium state, and the value of OD550 depends on the strength of induction of IPTG. We defiend it as LAV (Lytic Activity Value) as a quantitative parameter of lytic activity, and we established a standard method for lytic activity of λ lysis cassette. We characterized the lytic activity of λ lysis cassette quantitatively as our established method, and linked the result to the strength of induction(RPU). | ||
| - | + | We characterized the lytic ativity of λ lysis cassette with time. | |
| - | [[Image: | + | {{clear}} |
| - | + | [[Image:KyotofigD002.png|450px|left|]] | |
| + | [[Image:KyotoGrp101028-1.png|450px]] | ||
| + | <div style="float:right;">[[Team:Kyoto/Project/Goal B|<html><img src="https://static.igem.org/mediawiki/2010/thumb/a/ac/KyotoArrowB.png/200px-KyotoArrowB.png"/></html>]]</div> | ||
| + | {{clear}} | ||
| - | === | + | ====[[Team:Kyoto/Project/Goal C|Goal C: Characterization of the anti-killer gene, SΔTMD1]]==== |
| - | + | [[Image:KyotoGoalC.png|460px|left]] | |
| - | + | [[Image:KyotoFigC003.png|460px]]<br/> | |
| - | + | To check function of the anti-killer gene, first, we made construct and measured A550 of E.coli transformed with the device in medium with IPTG and without IPTG, and we compared this result with that of E.coli transformed with killer gene, lysis cassette. Then, we were able to make sure that Transmembrane domein1 (TMD1) is essential for , lysis cassette. | |
| + | {{clear}} | ||
| + | [[Image:GoalC fig3-2-2.png|450px]] | ||
| + | <div style="float:right;">[[Team:Kyoto/Project/Goal C|<html><img src="https://static.igem.org/mediawiki/2010/thumb/e/ea/KyotoArrowC.png/200px-KyotoArrowC.png"/></html>]]</div> | ||
| + | {{clear}} | ||
| - | + | [[#top-section|^Top]] | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ===Achievement=== | |
| + | # We measured RPU of R0011 in eight kinds of concentration of IPTG, and made continuous curve '''representing the relation between concentration of IPTG and RPU of R0011''' by simulation fitting. | ||
| + | # We '''characterized the lytic activity''' of λ lysis cassette quantitatively as our established method,and linked the result to the strength of induction (RPU). | ||
| + | # We '''found that a new fact''' that cell lysis caused by λ lysis cassette become an apparent equilibrium state, and the value of OD550 depends on the strength of induction of IPTG. We defined it as LAV(Lytic Activity Value)as a quantitative parameter of lytic activity, and '''we established a standard''' method for lytic activity of λ lysis cassette. | ||
| + | # We '''checked the activity of S<sub>ΔTMD1</sub>''', which can't form pore of holin and cells don't die although endolysin expresses. | ||
| - | + | [[#top-section|^Top]] | |
| - | + | ||
| - | + | ||
| - | # | + | |
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | + | ||
| - | |||
| + | |||
| + | ===Future work=== | ||
| + | 1. Characterization of S<sub>ΔTMD1</sub> quantitatively | ||
| + | |||
| + | We checked the function of S<sub>ΔTMD1</sub>, however, we couldn’t confirm that S<sub>ΔTMD1</sub>can inhibit S gene when S<sub>ΔTMD1</sub> expresses simultaneously. | ||
| + | We think this is because the expression level of S<sub>ΔTMD1</sub> is too low to inhibit the activity of S gene in the experiment. By Jamboree, using more strong promoter to express S<sub>ΔTMD1</sub>, we check the function of S<sub>ΔTMD1</sub>, which prevents cell-lysis. | ||
| + | |||
| + | 2. Change the point of cell lysis and make lysis cassette more useful device. | ||
| + | |||
| + | We used the inducible promoter to express S gene and measured what the promoter activity leads cell to lyse. Such experiment suggests that there is the threshold for cell lysis caused by lysis cassette. By expressing S<sub>ΔTMD1</sub>, we can make cell lysis at higher expression level, and change the threshold of the cell lysis. | ||
| + | |||
| + | If lysis cassette expressed by promoters which cannot exceed this threshold, cell lysis does not occur. | ||
| + | Expression of S<sub>ΔTMD1</sub> by constitutive promoter may inhibit the activity of lysis cassette and increase the threshold. | ||
| + | Therefore, we can regulate cell lysis by controlling expression level of S<sub>ΔTMD1</sub>. | ||
| + | Besides, by characterizing of the relationship between cell lysis and the balance between S gene and S<sub>ΔTMD1</sub> with RPU, we can control cell lysis easily with promoters characterized by RPU in the partsregistry…. | ||
| + | There are many kinds of well- characterized constitutive promoter in the partsregistry. | ||
| + | |||
| + | [[#top-section|^Top]] | ||
| + | |||
| + | ===References=== | ||
| + | # <html><a name="RefH001"></a></html> [http://www.ncbi.nlm.nih.gov/pubmed/20395970 PMID: 20395970] Khalil AS, Collins JJ., Synthetic biology: applications come of age., Nat Rev Genet. 2010 May;11(5):367-79. | ||
| + | # <html><a name="RefH002"></a></html> [http://www.ncbi.nlm.nih.gov/pubmed/15832375 PMID: 15832375] Paul D, Pandey G, Jain RK., Suicidal genetically engineered microorganisms for bioremediation: need and perspectives., Bioessays. 2005 May;27(5):563-73. | ||
| + | # <html><a name="RefH003"></a></html> [http://www.ncbi.nlm.nih.gov/pubmed/15973534 PMID: 15973534] Davison J., Risk mitigation of genetically modified bacteria and plants designed for bioremediation., J Ind Microbiol Biotechnol. 2005 Dec;32(11-12):639-50. Epub 2005 Jun 23. | ||
| + | # <html><a name="RefH004"></a></html>https://2009.igem.org/Team:Michigan | ||
| + | # <html><a name="RefH005"></a></html> [http://www.ncbi.nlm.nih.gov/pubmed/19897658 PMID: 19897658] White R, Tran TA, Dankenbring CA, Deaton J, Young R., The N-terminal transmembrane domain of lambda S is required for holin but not antiholin function., J Bacteriol. 2010 Feb;192(3):725-33. Epub 2009 Nov 6. | ||
| + | # <html><a name="RefH006"></a></html>https://2009.igem.org/Team:UNICAMP-Brazil | ||
| + | # <html><a name="RefH007"></a></html>https://2008.igem.org/Team:UC_Berkeley | ||
| + | # <html><a name="RefH008"></a></html> [http://www.ncbi.nlm.nih.gov/pubmed/18713319 PMID: 18713319] Berry J, Summer EJ, Struck DK, Young R., The final step in the phage infection cycle: the Rz and Rz1 lysis proteins link the inner and outer membranes., Mol Microbiol. 2008 Oct;70(2):341-51. Epub 2008 Aug 18. | ||
| + | # <html><a name="RefH009"></a></html> [http://www.ncbi.nlm.nih.gov/pubmed/2147680 PMID: 2147680] Garrett J, Bruno C, Young R., Lysis protein S of phage lambda functions in Saccharomyces cerevisiae., J Bacteriol. 1990 Dec;172(12):7275-7. | ||
| + | # <html><a name="RefH010"></a></html> [http://www.ncbi.nlm.nih.gov/pubmed/19298678 PMID: 19298678] Kelly JR, Rubin AJ, Davis JH, Ajo-Franklin CM, Cumbers J, Czar MJ, de Mora K, Glieberman AL, Monie DD, Endy D., Measuring the activity of BioBrick promoters using an in vivo reference standard., J Biol Eng. 2009 Mar 20;3:4. | ||
| + | # <html><a name="RefH011"></a></html> [http://www.ncbi.nlm.nih.gov/pubmed/7768829 PMID: 7768829] Chang CY, Nam K, Young R., S gene expression and the timing of lysis by bacteriophage lambda., J Bacteriol. 1995 Jun;177(11):3283-94. | ||
| + | # <html><a name="RefH012"></a></html> [http://www.ncbi.nlm.nih.gov/pubmed/11459934 PMID: 11459934] Gründling A, Manson MD, Young R., Holins kill without warning., Proc Natl Acad Sci U S A. 2001 Jul 31;98(16):9348-52. Epub 2001 Jul 17. | ||
| + | # <html><a name="RefH013"></a></html> [http://www.ncbi.nlm.nih.gov/pubmed/9573208 PMID: 9573208] Smith DL, Struck DK, Scholtz JM, Young R., Purification and biochemical characterization of the lambda holin., J Bacteriol. 1998 May;180(9):2531-40. | ||
| + | |||
| + | [[#top-section|^Top]] | ||
---- | ---- | ||
Latest revision as of 10:52, 18 November 2010
Contents |
Project
Abstract
The Fantastic Lysisbox: Genetically engineered cell death is essential for the application of biotechnology, such as in bioremediation area. In order to control the cell-death, we designed “Lysisbox”, which consists of a pair of modules: “Killer gene” and “Anti-killer gene.” As the Killer gene for Escherichia coli, we noted the lysis cassette [SRRz/Rz1 gene] of lambda phage coding for holin and endolysin. The holin form pores in the inner membrane and the endolysin access and degrade the peptidoglycan passing through the pores, leading the E.coli to death. As the Anti-killer gene, we chose SΔTMD1 coding for a dominant-negative holin that inhibits the formation of the fatal pores. The balance of these two genes' expression level has the key to the E.coli’s life or death. In addition, such controllable membrane pores should show critical functions for all living organisms with lipid membranes and a cell wall. “Lysisbox” will contribute a lot to all the future iGEM projects, thus you must say “FANTASTIC!!!"
Introduction
Background
The cell-death device is required in many fields [1] [2]. In bioremediation, for example, this device can prevent ecosystem disruption by introduced bacteria [3] [4]. Also, it can be applied to other ideas such as E.coli capsules which scatter drug or aroma by cell-lysis responding to a certain signal. Therefore, the cell-death devices based on diverse approaches, including previous iGEM projects have been developed [1] [3] [5] [6]. However, present cell-death devices are not useful because they have two problems as follows:
Therefore, we attempted to solve these problems and design new BioBricks that are better characterized and more applicable cell-death device to contribute to every future projects.
Problems of Present Cell-Death Devices
1.Need for Quantitative Characterization
In previous iGEM, many teams focused on λ lysis cassette, which is a killer-gene derived from λ phage and causes cell lysis. However, they have only characterized its lytic activity qualititatively. Quantitative characterization of its lytic activity is necessary in order to design a universal and applicable cell-death device. For example, we wish to analyze when cell lysis occurs after induction of λ lysis cassette, how many cells lyse, whether all of cells lyse or not, and so on.
2. Solution to the Restriction on Application of Promoters
In constructing a simple circuit consisting of an inducible promoter and λ lysis cassette, the combination are limited by the promoter's basal expression, strength of killer-function and many other factors. We looked for the solution to this problem and came up with an idea; using both the lysis cassette and its antikiller-gene, SΔTMD1, which lacks the code for trans membrane domain 1 of killer-gene. We can use two promoters (a constitutive one and an inducible one) for regulating cell-death balance. Into this circuit, we can apply more promoters to this device by using an appropriate constitutive promoter.
We considered that lysis cassette and SΔTMD1 are the most appropriate to killer-gene and anti-killer gene [5]. This was because we found some articles which show the effect of SΔTMD1 in the yeast [9] and also because previous iGEM teams already dealt with lysis cassette.
For further description about lysis cassette and SΔTMD1, go [learn more]
Our Goals
Goal A: Characterization of R0011, a strong lactose promoter
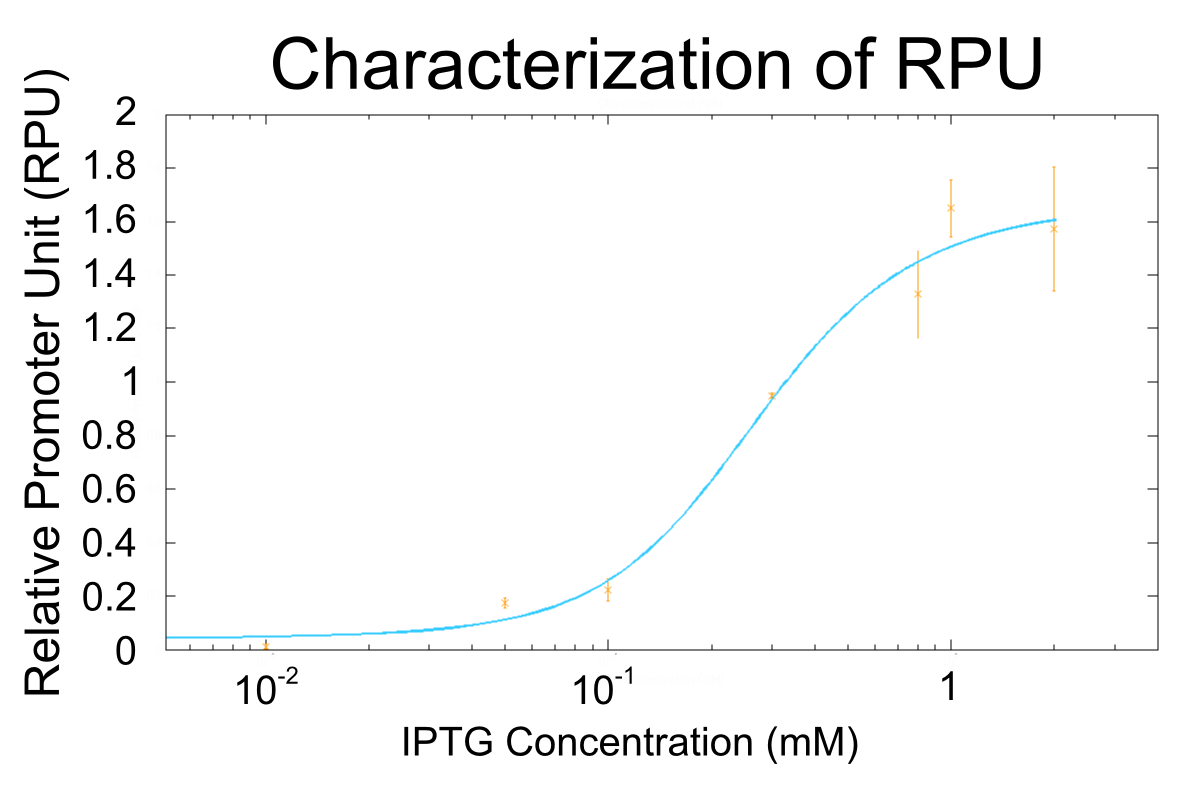
We used R0011, which is one of the most commonly used parts in iGEM, to characterize our genes because the strength of promoter activity is very high. However, in previous iGEM, no teams characterized R0011 with RPU. We tried to determine how the activity of R0011[10] changes.
Goal B: Quantitative characterization of λ lysis cassette
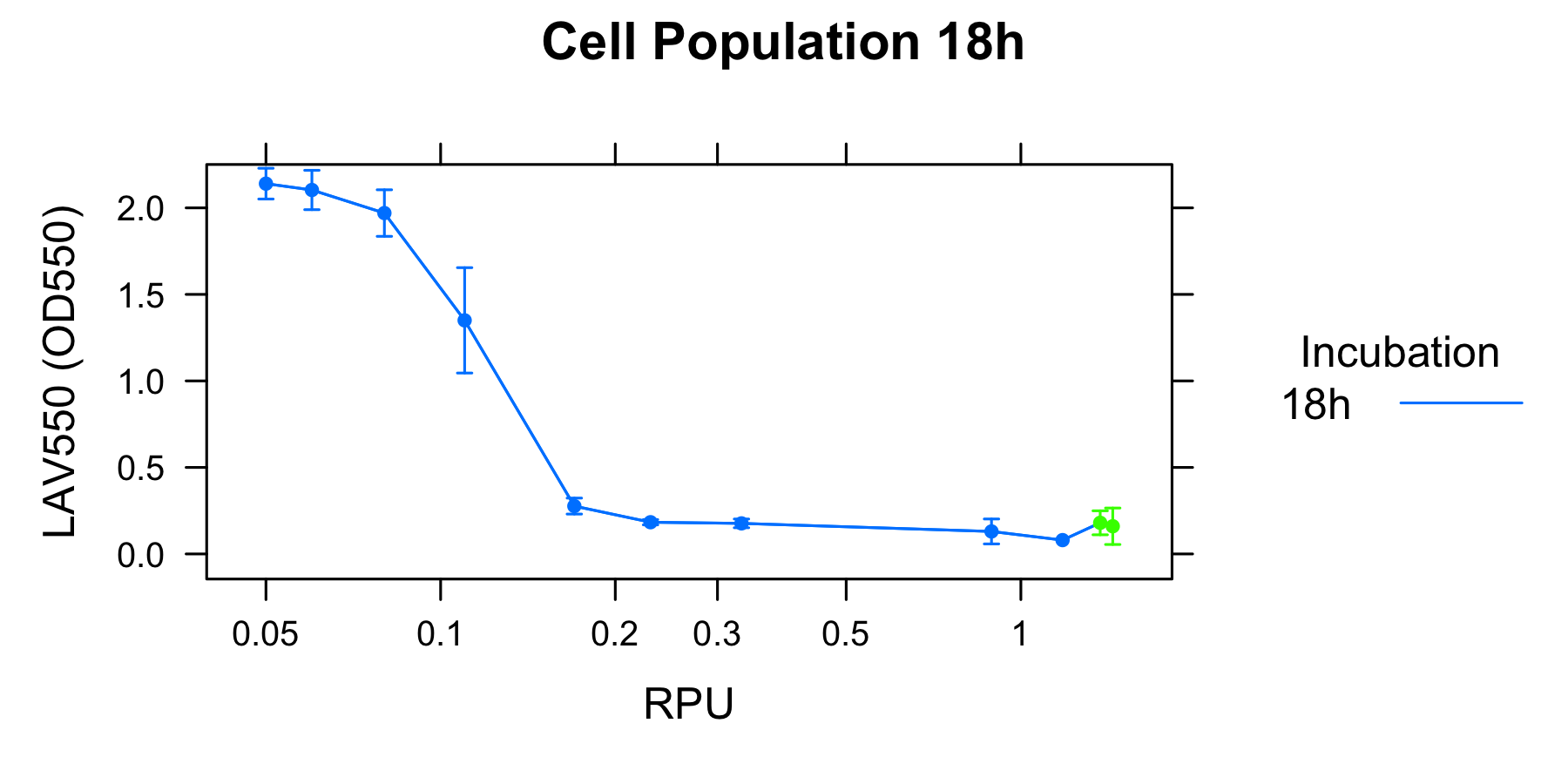
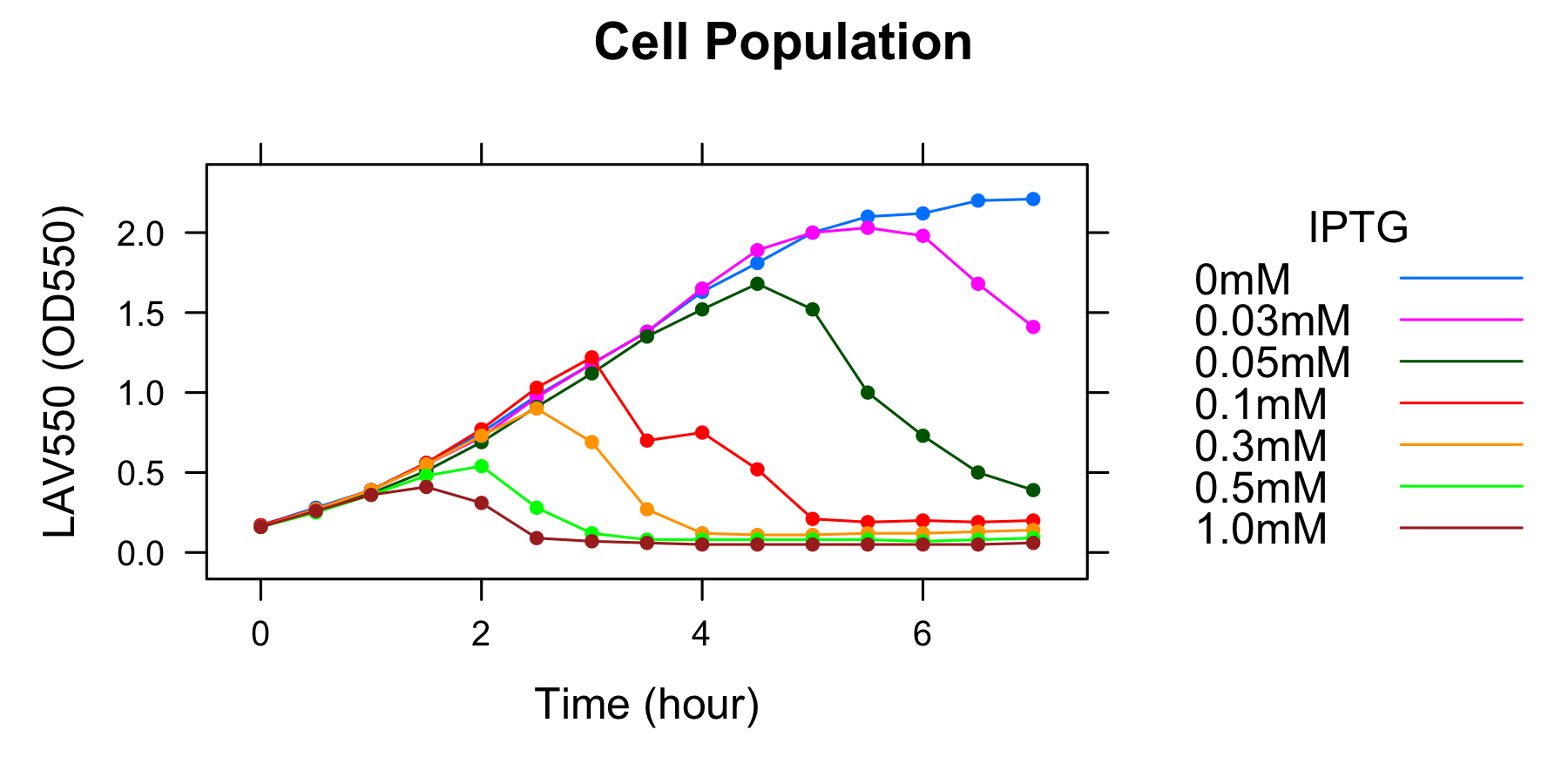
We found a new fact that cell lysis caused by λ lysis cassette become an apparent equilibrium state, and the value of OD550 depends on the strength of induction of IPTG. We defiend it as LAV (Lytic Activity Value) as a quantitative parameter of lytic activity, and we established a standard method for lytic activity of λ lysis cassette. We characterized the lytic activity of λ lysis cassette quantitatively as our established method, and linked the result to the strength of induction(RPU).
We characterized the lytic ativity of λ lysis cassette with time.
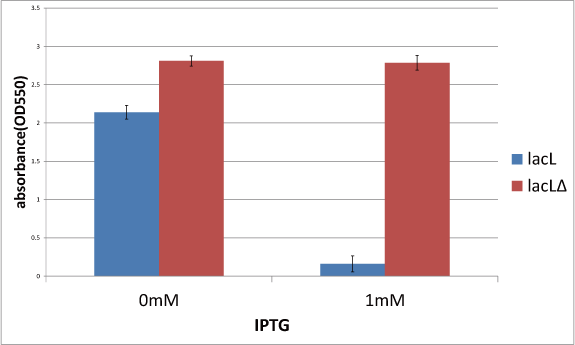
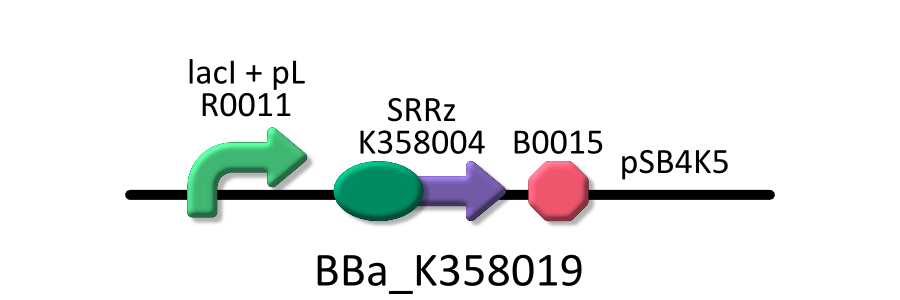
Goal C: Characterization of the anti-killer gene, SΔTMD1
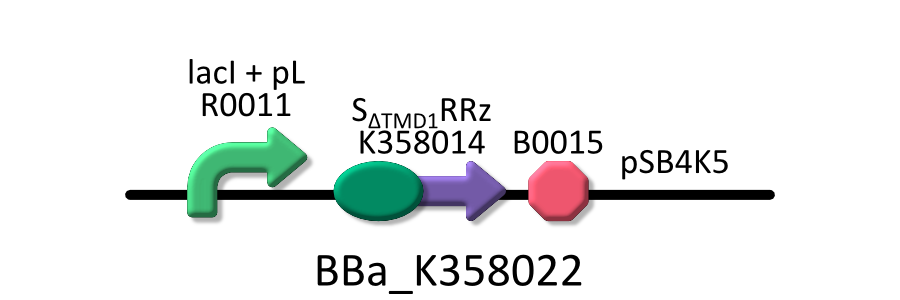
To check function of the anti-killer gene, first, we made construct and measured A550 of E.coli transformed with the device in medium with IPTG and without IPTG, and we compared this result with that of E.coli transformed with killer gene, lysis cassette. Then, we were able to make sure that Transmembrane domein1 (TMD1) is essential for , lysis cassette.
Achievement
- We measured RPU of R0011 in eight kinds of concentration of IPTG, and made continuous curve representing the relation between concentration of IPTG and RPU of R0011 by simulation fitting.
- We characterized the lytic activity of λ lysis cassette quantitatively as our established method,and linked the result to the strength of induction (RPU).
- We found that a new fact that cell lysis caused by λ lysis cassette become an apparent equilibrium state, and the value of OD550 depends on the strength of induction of IPTG. We defined it as LAV(Lytic Activity Value)as a quantitative parameter of lytic activity, and we established a standard method for lytic activity of λ lysis cassette.
- We checked the activity of SΔTMD1, which can't form pore of holin and cells don't die although endolysin expresses.
Future work
1. Characterization of SΔTMD1 quantitatively
We checked the function of SΔTMD1, however, we couldn’t confirm that SΔTMD1can inhibit S gene when SΔTMD1 expresses simultaneously. We think this is because the expression level of SΔTMD1 is too low to inhibit the activity of S gene in the experiment. By Jamboree, using more strong promoter to express SΔTMD1, we check the function of SΔTMD1, which prevents cell-lysis.
2. Change the point of cell lysis and make lysis cassette more useful device.
We used the inducible promoter to express S gene and measured what the promoter activity leads cell to lyse. Such experiment suggests that there is the threshold for cell lysis caused by lysis cassette. By expressing SΔTMD1, we can make cell lysis at higher expression level, and change the threshold of the cell lysis.
If lysis cassette expressed by promoters which cannot exceed this threshold, cell lysis does not occur. Expression of SΔTMD1 by constitutive promoter may inhibit the activity of lysis cassette and increase the threshold. Therefore, we can regulate cell lysis by controlling expression level of SΔTMD1. Besides, by characterizing of the relationship between cell lysis and the balance between S gene and SΔTMD1 with RPU, we can control cell lysis easily with promoters characterized by RPU in the partsregistry…. There are many kinds of well- characterized constitutive promoter in the partsregistry.
References
- [http://www.ncbi.nlm.nih.gov/pubmed/20395970 PMID: 20395970] Khalil AS, Collins JJ., Synthetic biology: applications come of age., Nat Rev Genet. 2010 May;11(5):367-79.
- [http://www.ncbi.nlm.nih.gov/pubmed/15832375 PMID: 15832375] Paul D, Pandey G, Jain RK., Suicidal genetically engineered microorganisms for bioremediation: need and perspectives., Bioessays. 2005 May;27(5):563-73.
- [http://www.ncbi.nlm.nih.gov/pubmed/15973534 PMID: 15973534] Davison J., Risk mitigation of genetically modified bacteria and plants designed for bioremediation., J Ind Microbiol Biotechnol. 2005 Dec;32(11-12):639-50. Epub 2005 Jun 23.
- https://2009.igem.org/Team:Michigan
- [http://www.ncbi.nlm.nih.gov/pubmed/19897658 PMID: 19897658] White R, Tran TA, Dankenbring CA, Deaton J, Young R., The N-terminal transmembrane domain of lambda S is required for holin but not antiholin function., J Bacteriol. 2010 Feb;192(3):725-33. Epub 2009 Nov 6.
- https://2009.igem.org/Team:UNICAMP-Brazil
- https://2008.igem.org/Team:UC_Berkeley
- [http://www.ncbi.nlm.nih.gov/pubmed/18713319 PMID: 18713319] Berry J, Summer EJ, Struck DK, Young R., The final step in the phage infection cycle: the Rz and Rz1 lysis proteins link the inner and outer membranes., Mol Microbiol. 2008 Oct;70(2):341-51. Epub 2008 Aug 18.
- [http://www.ncbi.nlm.nih.gov/pubmed/2147680 PMID: 2147680] Garrett J, Bruno C, Young R., Lysis protein S of phage lambda functions in Saccharomyces cerevisiae., J Bacteriol. 1990 Dec;172(12):7275-7.
- [http://www.ncbi.nlm.nih.gov/pubmed/19298678 PMID: 19298678] Kelly JR, Rubin AJ, Davis JH, Ajo-Franklin CM, Cumbers J, Czar MJ, de Mora K, Glieberman AL, Monie DD, Endy D., Measuring the activity of BioBrick promoters using an in vivo reference standard., J Biol Eng. 2009 Mar 20;3:4.
- [http://www.ncbi.nlm.nih.gov/pubmed/7768829 PMID: 7768829] Chang CY, Nam K, Young R., S gene expression and the timing of lysis by bacteriophage lambda., J Bacteriol. 1995 Jun;177(11):3283-94.
- [http://www.ncbi.nlm.nih.gov/pubmed/11459934 PMID: 11459934] Gründling A, Manson MD, Young R., Holins kill without warning., Proc Natl Acad Sci U S A. 2001 Jul 31;98(16):9348-52. Epub 2001 Jul 17.
- [http://www.ncbi.nlm.nih.gov/pubmed/9573208 PMID: 9573208] Smith DL, Struck DK, Scholtz JM, Young R., Purification and biochemical characterization of the lambda holin., J Bacteriol. 1998 May;180(9):2531-40.
 "
"